Transcriptome Dynamics and Potential Roles of Sox6 in the Postnatal Heart
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Prox1regulates the Subtype-Specific Development of Caudal Ganglionic
The Journal of Neuroscience, September 16, 2015 • 35(37):12869–12889 • 12869 Development/Plasticity/Repair Prox1 Regulates the Subtype-Specific Development of Caudal Ganglionic Eminence-Derived GABAergic Cortical Interneurons X Goichi Miyoshi,1 Allison Young,1 Timothy Petros,1 Theofanis Karayannis,1 Melissa McKenzie Chang,1 Alfonso Lavado,2 Tomohiko Iwano,3 Miho Nakajima,4 Hiroki Taniguchi,5 Z. Josh Huang,5 XNathaniel Heintz,4 Guillermo Oliver,2 Fumio Matsuzaki,3 Robert P. Machold,1 and Gord Fishell1 1Department of Neuroscience and Physiology, NYU Neuroscience Institute, Smilow Research Center, New York University School of Medicine, New York, New York 10016, 2Department of Genetics & Tumor Cell Biology, St. Jude Children’s Research Hospital, Memphis, Tennessee 38105, 3Laboratory for Cell Asymmetry, RIKEN Center for Developmental Biology, Kobe 650-0047, Japan, 4Laboratory of Molecular Biology, Howard Hughes Medical Institute, GENSAT Project, The Rockefeller University, New York, New York 10065, and 5Cold Spring Harbor Laboratory, Cold Spring Harbor, New York 11724 Neurogliaform (RELNϩ) and bipolar (VIPϩ) GABAergic interneurons of the mammalian cerebral cortex provide critical inhibition locally within the superficial layers. While these subtypes are known to originate from the embryonic caudal ganglionic eminence (CGE), the specific genetic programs that direct their positioning, maturation, and integration into the cortical network have not been eluci- dated. Here, we report that in mice expression of the transcription factor Prox1 is selectively maintained in postmitotic CGE-derived cortical interneuron precursors and that loss of Prox1 impairs the integration of these cells into superficial layers. Moreover, Prox1 differentially regulates the postnatal maturation of each specific subtype originating from the CGE (RELN, Calb2/VIP, and VIP). -
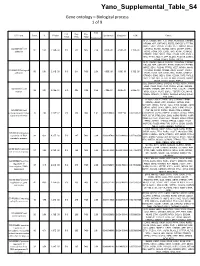
Supplemental Tables4.Pdf
Yano_Supplemental_Table_S4 Gene ontology – Biological process 1 of 9 Fold List Pop Pop GO Term Count % PValue Bonferroni Benjamini FDR Genes Total Hits Total Enrichment DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0007155~cell CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0022610~biological CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DCC, ENAH, PLXNA2, CAPZA2, ATP5B, ASTN1, PAX6, ZEB2, CDH2, CDH4, GLI3, CD24A, EPHB1, NRCAM, GO:0006928~cell CTTNBP2, EDNRB, APP, PTK2, ETV1, CLASP2, STRBP, 36 4.04 3.46E-07 510 205 7436 2.56 7.28E-04 3.64E-04 5.98E-04 motion NRG1, DCLK1, PLAT, SGPL1, TGFBR1, EVL, MYH9, YWHAE, NCKAP1, CTNNA2, SEMA6A, EPHA4, NDEL1, FYN, LRP6 PLXNA2, ADCY5, PAX6, GLI3, CTNNB1, LPHN2, EDNRB, LPHN3, APP, CSNK2A1, GPR45, NRG1, RAPGEF1, WWOX, SGPL1, TLE4, SPEN, NCAM1, DDR1, GRB10, GRM3, GNAQ, HIPK1, GNB1, HIPK2, PYGO1, GO:0007166~cell RNF138, ROR2, CNTN1, -

Accompanies CD8 T Cell Effector Function Global DNA Methylation
Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer, Benjamin G. Barwick, Benjamin A. Youngblood, Rafi Ahmed and Jeremy M. Boss This information is current as of October 1, 2021. J Immunol 2013; 191:3419-3429; Prepublished online 16 August 2013; doi: 10.4049/jimmunol.1301395 http://www.jimmunol.org/content/191/6/3419 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130139 Material 5.DC1 References This article cites 81 articles, 25 of which you can access for free at: http://www.jimmunol.org/content/191/6/3419.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists by guest on October 1, 2021 • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer,* Benjamin G. Barwick,* Benjamin A. Youngblood,*,† Rafi Ahmed,*,† and Jeremy M. -

Supplemental Materials and Methods
Supplemental Material for: Transcriptional silencing of γ-globin by BCL11A involves long-range interactions and cooperation with SOX6 Jian Xu, Vijay G. Sankaran, Min Ni, Tobias F. Menne, Rishi V. Puram, Woojin Kim, Stuart H. Orkin* *To whom correspondence should be addressed. E-mail: [email protected] Supplemental Materials and Methods Flow cytometry Cells at various stages of differentiation were analyzed by flow cytometry using FACSCalibur (BD Biosciences, San Jose, CA). Live cells were identified and gated by exclusion of 7-amino-actinomycin D (7-AAD; BD Pharmingen). The cells were analyzed for expression of cell surface receptors with antibodies specific for CD34, CD45, CD71, CD235, and CD36 conjugated to phycoerythrin (PE), fluorescein isothiocyanate (FITC), or allophycocyanin (APC; BD Pharmingen). Data were analyzed using FlowJo software (Ashland, OR). Cytology Cytocentrifuge preparations were stained with May-Grunwald-Giemsa as previously described (Sankaran et al. 2008). Real-time RT-PCR Real-time quantitative RT-PCR was performed using the iQ SYBR Green Supermix (Bio- Rad). The following primers were used for real-time RT-PCR: human and mouse BCL11A-XL (forward, 5’-ATGCGAGCTGTGCAACTATG-3’; reverse, 5’- GTAAACGTCCTTCCCCACCT-3’), human and mouse BCL11A-L (forward, 5’- CAGCTCAAAAGAGGGCAGAC-3’; reverse, 5’-GAGCTTCCATCCGAAAACTG-3’), and human BCL11A exon 1 and 2 (common between all known isoforms; forward, 5’- AACCCCAGCACTTAAGCAAA-3’; reverse, 5’-GGAGGTCATGATCCCCTTCT-3’). Supplemental Figure Legends Supplemental Figure 1. Expression of BCL11A isoforms in human and mouse erythroid cells. (A) Schematic diagram of human BCL11A isoforms (Liu et al. 2006). The antibodies used for ChIP experiments and their corresponding epitopes are indicated. Locations of primers used for RT-PCR analysis of all BCL11A isoforms (forward and reverse primers indicated by arrowheads), XL and L isoforms are indicated. -

Transcriptional Silencing of G-Globin by BCL11A Involves Long-Range Interactions and Cooperation with SOX6
Downloaded from genesdev.cshlp.org on September 25, 2021 - Published by Cold Spring Harbor Laboratory Press Transcriptional silencing of g-globin by BCL11A involves long-range interactions and cooperation with SOX6 Jian Xu,1,2,3 Vijay G. Sankaran,1,2,4 Min Ni,5 Tobias F. Menne,1,2 Rishi V. Puram,4 Woojin Kim,1,2 and Stuart H. Orkin1,2,3,6 1Division of Hematology/Oncology, Children’s Hospital Boston, Boston, Massachusetts 02115, USA; 2Department of Pediatric Oncology, Dana-Farber Cancer Institute, Harvard Stem Cell Institute, Harvard Medical School, Boston, Massachusetts 02115, USA; 3Howard Hughes Medical Institute, Boston, Massachusetts 02115, USA; 4Harvard-MIT Division of Health Sciences and Technology, Boston, Massachusetts 02115, USA; 5Division of Molecular and Cellular Oncology, Department of Medical Oncology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, Massachusetts 02115, USA The developmental switch from human fetal (g) to adult (b) hemoglobin represents a clinically important example of developmental gene regulation. The transcription factor BCL11A is a central mediator of g-globin silencing and hemoglobin switching. Here we determine chromatin occupancy of BCL11A at the human b-globin locus and other genomic regions in vivo by high-resolution chromatin immunoprecipitation (ChIP)–chip analysis. BCL11A binds the upstream locus control region (LCR), e-globin, and the intergenic regions between g-globin and d-globin genes. A chromosome conformation capture (3C) assay shows that BCL11A reconfigures the b-globin cluster by modulating chromosomal loop formation. We also show that BCL11A and the HMG-box-containing transcription factor SOX6 interact physically and functionally during erythroid maturation. BCL11A and SOX6 co-occupy the human b-globin cluster along with GATA1, and cooperate in silencing g-globin transcription in adult human erythroid progenitors. -

Molecular Regulation in Dopaminergic Neuron Development
International Journal of Molecular Sciences Review Molecular Regulation in Dopaminergic Neuron Development. Cues to Unveil Molecular Pathogenesis and Pharmacological Targets of Neurodegeneration Floriana Volpicelli 1 , Carla Perrone-Capano 1,2 , Gian Carlo Bellenchi 2,3, Luca Colucci-D’Amato 4,* and Umberto di Porzio 2 1 Department of Pharmacy, University of Naples Federico II, 80131 Naples, Italy; fl[email protected] (F.V.); [email protected] (C.P.C.) 2 Institute of Genetics and Biophysics “Adriano Buzzati Traverso”, CNR, 80131 Rome, Italy; [email protected] (G.C.B.); [email protected] (U.d.P.) 3 Department of Systems Medicine, University of Rome Tor Vergata, 00133 Rome, Italy 4 Department of Environmental, Biological and Pharmaceutical Sciences and Technologies, University of Campania “Luigi Vanvitelli”, 81100 Caserta, Italy * Correspondence: [email protected]; Tel.: +39-0823-274577 Received: 28 April 2020; Accepted: 1 June 2020; Published: 3 June 2020 Abstract: The relatively few dopaminergic neurons in the mammalian brain are mostly located in the midbrain and regulate many important neural functions, including motor integration, cognition, emotive behaviors and reward. Therefore, alteration of their function or degeneration leads to severe neurological and neuropsychiatric diseases. Unraveling the mechanisms of midbrain dopaminergic (mDA) phenotype induction and maturation and elucidating the role of the gene network involved in the development and maintenance of these neurons is of pivotal importance to rescue or substitute these cells in order to restore dopaminergic functions. Recently, in addition to morphogens and transcription factors, microRNAs have been identified as critical players to confer mDA identity. The elucidation of the gene network involved in mDA neuron development and function will be crucial to identify early changes of mDA neurons that occur in pre-symptomatic pathological conditions, such as Parkinson’s disease. -

SOX6 and PDCD4 Enhance Cardiomyocyte Apoptosis Through LPS-Induced Mir-499 Inhibition
Apoptosis (2016) 21:174–183 DOI 10.1007/s10495-015-1201-6 ORIGINAL PAPER SOX6 and PDCD4 enhance cardiomyocyte apoptosis through LPS-induced miR-499 inhibition 1 2 1 3 1 Zhuqing Jia • Jiaji Wang • Qiong Shi • Siyu Liu • Weiping Wang • 1 1 1 1 1 Yuyao Tian • Qin Lu • Ping Chen • Kangtao Ma • Chunyan Zhou Published online: 10 December 2015 Ó The Author(s) 2015. This article is published with open access at Springerlink.com Abstract Sepsis-induced cardiac apoptosis is one of the the cardiomyocytes against LPS-induced apoptosis. In major pathogenic factors in myocardial dysfunction. As it brief, our results demonstrate the existence of a miR-499- enhances numerous proinflammatory factors, lipopolysac- SOX6/PDCD4-BCL-2 family pathway in cardiomyocytes charide (LPS) is considered the principal mediator in this in response to LPS stimulation. pathological process. However, the detailed mechanisms involved are unclear. In this study, we attempted to explore Keywords SOX6 Á PDCD4 Á LPS Á miR-499 Á the mechanisms involved in LPS-induced cardiomyocyte Cardiomyocyte Á Apoptosis apoptosis. We found that LPS stimulation inhibited microRNA (miR)-499 expression and thereby upregulated the expression of SOX6 and PDCD4 in neonatal rat car- Introduction diomyocytes. We demonstrate that SOX6 and PDCD4 are target genes of miR-499, and they enhance LPS-induced Sepsis-induced myocardial functional disorder is one of the cardiomyocyte apoptosis by activating the BCL-2 family main predictors of morbidity and mortality of sepsis [1]; pathway. The apoptosis process enhanced by overexpres- apoptosis is one of the major contributors to the patho- sion of SOX6 or PDCD4, was rescued by the cardiac- physiology of sepsis [2]. -

Zeb2 Chip-Seq in Stem Cells Zeb2 DNA-Binding Sites in ES Cell
bioRxiv preprint doi: https://doi.org/10.1101/2021.07.06.451350; this version posted July 6, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Zeb2 ChIP-seq in stem cells Zeb2 DNA-binding sites in ES cell derived neuroprogenitor cells reveal autoregulation and align with neurodevelopmental knockout mouse and disease phenotypes. Judith C. Birkhoff1, Anne L. Korporaal1, Rutger W.W. Brouwer2, Claudia Milazzo1#, Lidia Mouratidou1, Mirjam C.G.N. van den Hout2, Wilfred F.J. van IJcken1,2, Danny Huylebroeck1,3*, $, Andrea Conidi1*, $ 1 Dept. Cell Biology, Erasmus University Medical Center, Rotterdam, 3015 CN, The Netherlands; 2 Center for Biomics-Genomics, Erasmus University Medical Center, Rotterdam 3015 CN, The Netherlands; 3 Dept. Development and Regeneration, KU Leuven, Leuven 3000, Belgium * shared senior authors # Present address: Dept. Clinical Genetics, Erasmus University Medical Center, Rotterdam, 3015 CN, The Netherlands $ Corresponding author: Andrea Conidi, PhD, Dept. Cell Biology, Erasmus MC, Building Ee - room Ee-1040, Wytemaweg 80, 3015 CN Rotterdam, The Netherlands; Tel: +31-10-7043169; Fax: +31-10-7044743; E-mail: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.07.06.451350; this version posted July 6, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Author contributions -

Genome-Wide Mapping of Sox6 Binding Sites in Skeletal Muscle Reveals Both Direct and Indirect Regulation of Muscle Terminal Differentiation by Sox6
UC Davis UC Davis Previously Published Works Title Genome-wide mapping of Sox6 binding sites in skeletal muscle reveals both direct and indirect regulation of muscle terminal differentiation by Sox6 Permalink https://escholarship.org/uc/item/7fm1r9gq Journal BMC Developmental Biology, 11(1) ISSN 1471-213X Authors An, Chung-Il Dong, Yao Hagiwara, Nobuko Publication Date 2011-10-10 DOI http://dx.doi.org/10.1186/1471-213X-11-59 Supplemental Material https://escholarship.org/uc/item/7fm1r9gq#supplemental Peer reviewed eScholarship.org Powered by the California Digital Library University of California An et al. BMC Developmental Biology 2011, 11:59 http://www.biomedcentral.com/1471-213X/11/59 RESEARCHARTICLE Open Access Genome-wide mapping of Sox6 binding sites in skeletal muscle reveals both direct and indirect regulation of muscle terminal differentiation by Sox6 Chung-Il An, Yao Dong and Nobuko Hagiwara* Abstract Background: Sox6 is a multi-faceted transcription factor involved in the terminal differentiation of many different cell types in vertebrates. It has been suggested that in mice as well as in zebrafish Sox6 plays a role in the terminal differentiation of skeletal muscle by suppressing transcription of slow fiber specific genes. In order to understand how Sox6 coordinately regulates the transcription of multiple fiber type specific genes during muscle development, we have performed ChIP-seq analyses to identify Sox6 target genes in mouse fetal myotubes and generated muscle-specific Sox6 knockout (KO) mice to determine the Sox6 null muscle phenotype in adult mice. Results: We have identified 1,066 Sox6 binding sites using mouse fetal myotubes. The Sox6 binding sites were found to be associated with slow fiber-specific, cardiac, and embryonic isoform genes that are expressed in the sarcomere as well as transcription factor genes known to play roles in muscle development. -
![RT² Profiler PCR Array (96-Well Format and 384-Well [4 X 96] Format)](https://docslib.b-cdn.net/cover/3255/rt%C2%B2-profiler-pcr-array-96-well-format-and-384-well-4-x-96-format-3773255.webp)
RT² Profiler PCR Array (96-Well Format and 384-Well [4 X 96] Format)
RT² Profiler PCR Array (96-Well Format and 384-Well [4 x 96] Format) Human Stem Cell Transcription Factors Cat. no. 330231 PAHS-501ZA For pathway expression analysis Format For use with the following real-time cyclers RT² Profiler PCR Array, Applied Biosystems® models 5700, 7000, 7300, 7500, Format A 7700, 7900HT, ViiA™ 7 (96-well block); Bio-Rad® models iCycler®, iQ™5, MyiQ™, MyiQ2; Bio-Rad/MJ Research Chromo4™; Eppendorf® Mastercycler® ep realplex models 2, 2s, 4, 4s; Stratagene® models Mx3005P®, Mx3000P®; Takara TP-800 RT² Profiler PCR Array, Applied Biosystems models 7500 (Fast block), 7900HT (Fast Format C block), StepOnePlus™, ViiA 7 (Fast block) RT² Profiler PCR Array, Bio-Rad CFX96™; Bio-Rad/MJ Research models DNA Format D Engine Opticon®, DNA Engine Opticon 2; Stratagene Mx4000® RT² Profiler PCR Array, Applied Biosystems models 7900HT (384-well block), ViiA 7 Format E (384-well block); Bio-Rad CFX384™ RT² Profiler PCR Array, Roche® LightCycler® 480 (96-well block) Format F RT² Profiler PCR Array, Roche LightCycler 480 (384-well block) Format G RT² Profiler PCR Array, Fluidigm® BioMark™ Format H Sample & Assay Technologies Description The Human Stem Cell Transcription Factors RT² Profiler PCR Array profiles the expression of 84 key genes associated with stem cell differentiation and development. During development, a few key transcription factors (NANOG, POU5F1, and SOX2) drive embryonic stem cell maintenance. These genes activate or repress other key transcription factors represented by this array that are necessary for development and differentiation, forming a transcriptional regulatory network. For example, embryonic stem cell maintenance requires POU5F1 (OCT4), whereas trophectoderm development requires CDX2. -

Autocrine IFN Signaling Inducing Profibrotic Fibroblast Responses By
Downloaded from http://www.jimmunol.org/ by guest on September 23, 2021 Inducing is online at: average * The Journal of Immunology , 11 of which you can access for free at: 2013; 191:2956-2966; Prepublished online 16 from submission to initial decision 4 weeks from acceptance to publication August 2013; doi: 10.4049/jimmunol.1300376 http://www.jimmunol.org/content/191/6/2956 A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Autocrine IFN Signaling Feng Fang, Kohtaro Ooka, Xiaoyong Sun, Ruchi Shah, Swati Bhattacharyya, Jun Wei and John Varga J Immunol cites 49 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130037 6.DC1 This article http://www.jimmunol.org/content/191/6/2956.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 23, 2021. The Journal of Immunology A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Inducing Autocrine IFN Signaling Feng Fang,* Kohtaro Ooka,* Xiaoyong Sun,† Ruchi Shah,* Swati Bhattacharyya,* Jun Wei,* and John Varga* Activation of TLR3 by exogenous microbial ligands or endogenous injury-associated ligands leads to production of type I IFN. -

SUPPLEMENTARY DATA Retinoic Acid Mediates Visceral-Specific
SUPPLEMENTARY DATA Retinoic Acid Mediates Visceral-specific Adipogenic Defects of Human Adipose-derived Stem Cells Kosuke Takeda, Sandhya Sriram, Xin Hui Derryn Chan, Wee Kiat Ong, Chia Rou Yeo, Betty Tan, Seung-Ah Lee, Kien Voon Kong, Shawn Hoon, Hongfeng Jiang, Jason J. Yuen, Jayakumar Perumal, Madhur Agrawal, Candida Vaz, Jimmy So, Asim Shabbir, William S. Blaner, Malini Olivo, Weiping Han, Vivek Tanavde, Sue-Anne Toh, and Shigeki Sugii Supplementary Experimental Procedures RNA-Seq experiment and analysis Total RNA was extracted from SC and VS fat depots using Qiagen RNeasy plus kit. Poly-A mRNA was then enriched from ~5 µg of total RNA with oligo dT beads (Life Technologies). Approximately 100 ng of poly-A mRNA recovered was used to construct multiplexed strand-specific RNA-seq libraries as per manufacturer’s instruction (NEXTflexTM Rapid Directional RNA-SEQ Kit (dUTP-Based) v2). Individual library quality was assessed with an Agilent 2100 Bioanalyzer and quantified with a QuBit 2.0 fluorometer before pooling for sequencing on Illumina HiSeq 2000 (1x101 bp read). The pooled libraries were quantified using the KAPA quantification kit (KAPA Biosystems) prior to cluster formation. Fastq formatted reads were processed with Trimmomatic to remove adapter sequences and trim low quality bases (LEADING:3 TRAILING:3 SLIDINGWINDOW:4:15 MINLEN:36). Reads were aligned to the human genome (hg19) using Tophat version 2 (settings --no-coverage-search --library- type=fr-firststrand). Feature read counts were generated using htseq-count (Python package HTSeq default union-counting mode, strand=reverse). Differential Expression analysis was performed using the edgeR package in both ‘classic’ and generalized linear model (glm) modes to contrast SC and VS adipose tissues from 8 non-diabetic patients.