Zeb2 Chip-Seq in Stem Cells Zeb2 DNA-Binding Sites in ES Cell
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Experimental Chronic Jet Lag Promotes Growth and Lung Metastasis of Lewis Lung Carcinoma in C57BL/6 Mice
ONCOLOGY REPORTS 27: 1417-1428, 2012 Experimental chronic jet lag promotes growth and lung metastasis of Lewis lung carcinoma in C57BL/6 mice 1,2,6 1,3 1,4 1,2 1,5 MINGWEI WU , JING ZENG , YANFENG CHEN , ZHAOLEI ZENG , JINXIN ZHANG , YUCHEN CAI1,2, YANLI YE1,2, LIWU FU1,2, LIJIAN XIAN1,2 and ZHONGPING CHEN1,6 1 2 3 4 State Key Laboratory of Oncology in South China; Departments of Research, Pathology, and Head and Neck Cancer, Cancer Center, Sun Yat-Sen University; 5Department of Medical Statistics and Epidemiology, Sun Yat-Sen University; 6Department of Neurosurgery, Cancer Center, Sun Yat-Sen University, Guangzhou, Guangdong, P.R. China Received December 8, 2011; Accepted January 17, 2012 DOI: 10.3892/or.2012.1688 Abstract. Circadian rhythm has been linked to cancer genesis are governed by a biological clock. The mammalian circadian and development, but the detailed mechanism by which circa- clock contains three components: input pathways, a central dian disruption accelerates tumor growth remains unclear. The pacemaker and output pathways. The mammalian central purpose of this study was to investigate the effect of circadian pacemaker is located in the suprachiasmatic nuclei (SCN) disruption on tumor growth and metastasis in male C57BL/6 of the anterior hypothalamus and controls the activity of the mice, using an experimental chronic jet lag model. Lewis lung peripheral clocks through the neuroendocrine and autonomic carcinoma cells were inoculated into both flanks of the mice nervous systems (1,2). Circadian rhythms govern the rhythmic following 10 days of exposure to experimental chronic jet lag changes in the behavior and/or physiology of mammals, such or control conditions. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Transcriptome Dynamics and Potential Roles of Sox6 in the Postnatal Heart
RESEARCH ARTICLE Transcriptome Dynamics and Potential Roles of Sox6 in the Postnatal Heart Chung-Il An1☯*, Yasunori Ichihashi2☯¤a¤b*, Jie Peng3, Neelima R. Sinha2, Nobuko Hagiwara1* 1 Division of Cardiovascular Medicine, Department of Internal Medicine, University of California Davis, Davis, California, United States of America, 2 Department of Plant Biology, University of California Davis, Davis, California, United States of America, 3 Department of Statistics, University of California Davis, Davis, California, United States of America ☯ These authors contributed equally to this work. a11111 ¤a Current address: RIKEN Center for Sustainable Resource Science, Yokohama, Kanagawa, Japan ¤b Current address: JST, PRESTO, Kawaguchi, Saitama, Japan * [email protected] (CA); [email protected] (YI); [email protected] (NH) Abstract OPEN ACCESS The postnatal heart undergoes highly coordinated developmental processes culminating in Citation: An C-I, Ichihashi Y, Peng J, Sinha NR, the complex physiologic properties of the adult heart. The molecular mechanisms of postna- Hagiwara N (2016) Transcriptome Dynamics and tal heart development remain largely unexplored despite their important clinical implications. Potential Roles of Sox6 in the Postnatal Heart. To gain an integrated view of the dynamic changes in gene expression during postnatal PLoS ONE 11(11): e0166574. doi:10.1371/journal. heart development at the organ level, time-series transcriptome analyses of the postnatal pone.0166574 hearts of neonatal through adult mice (P1, P7, P14, P30, and P60) were performed using a Editor: Katherine Yutzey, Cincinnati Children's newly developed bioinformatics pipeline. We identified functional gene clusters by principal Hospital Medical Center, UNITED STATES component analysis with self-organizing map clustering which revealed organized, discrete Received: July 16, 2016 gene expression patterns corresponding to biological functions associated with the neona- Accepted: October 31, 2016 tal, juvenile and adult stages of postnatal heart development. -

Prox1regulates the Subtype-Specific Development of Caudal Ganglionic
The Journal of Neuroscience, September 16, 2015 • 35(37):12869–12889 • 12869 Development/Plasticity/Repair Prox1 Regulates the Subtype-Specific Development of Caudal Ganglionic Eminence-Derived GABAergic Cortical Interneurons X Goichi Miyoshi,1 Allison Young,1 Timothy Petros,1 Theofanis Karayannis,1 Melissa McKenzie Chang,1 Alfonso Lavado,2 Tomohiko Iwano,3 Miho Nakajima,4 Hiroki Taniguchi,5 Z. Josh Huang,5 XNathaniel Heintz,4 Guillermo Oliver,2 Fumio Matsuzaki,3 Robert P. Machold,1 and Gord Fishell1 1Department of Neuroscience and Physiology, NYU Neuroscience Institute, Smilow Research Center, New York University School of Medicine, New York, New York 10016, 2Department of Genetics & Tumor Cell Biology, St. Jude Children’s Research Hospital, Memphis, Tennessee 38105, 3Laboratory for Cell Asymmetry, RIKEN Center for Developmental Biology, Kobe 650-0047, Japan, 4Laboratory of Molecular Biology, Howard Hughes Medical Institute, GENSAT Project, The Rockefeller University, New York, New York 10065, and 5Cold Spring Harbor Laboratory, Cold Spring Harbor, New York 11724 Neurogliaform (RELNϩ) and bipolar (VIPϩ) GABAergic interneurons of the mammalian cerebral cortex provide critical inhibition locally within the superficial layers. While these subtypes are known to originate from the embryonic caudal ganglionic eminence (CGE), the specific genetic programs that direct their positioning, maturation, and integration into the cortical network have not been eluci- dated. Here, we report that in mice expression of the transcription factor Prox1 is selectively maintained in postmitotic CGE-derived cortical interneuron precursors and that loss of Prox1 impairs the integration of these cells into superficial layers. Moreover, Prox1 differentially regulates the postnatal maturation of each specific subtype originating from the CGE (RELN, Calb2/VIP, and VIP). -
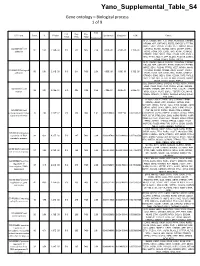
Supplemental Tables4.Pdf
Yano_Supplemental_Table_S4 Gene ontology – Biological process 1 of 9 Fold List Pop Pop GO Term Count % PValue Bonferroni Benjamini FDR Genes Total Hits Total Enrichment DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0007155~cell CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DLC1, CADM1, NELL2, CLSTN1, PCDHGA8, CTNNB1, NRCAM, APP, CNTNAP2, FERT2, RAPGEF1, PTPRM, MPDZ, SDK1, PCDH9, PTPRS, VEZT, NRXN1, MYH9, GO:0022610~biological CTNNA2, NCAM1, NCAM2, DDR1, LSAMP, CNTN1, 50 5.61 2.14E-08 510 311 7436 2.34 4.50E-05 4.50E-05 3.70E-05 adhesion ROR2, VCAN, DST, LIMS1, TNC, ASTN1, CTNND2, CTNND1, CDH2, NEO1, CDH4, CD24A, FAT3, PVRL3, TRO, TTYH1, MLLT4, LPP, NLGN1, PCDH19, LAMA1, ITGA9, CDH13, CDON, PSPC1 DCC, ENAH, PLXNA2, CAPZA2, ATP5B, ASTN1, PAX6, ZEB2, CDH2, CDH4, GLI3, CD24A, EPHB1, NRCAM, GO:0006928~cell CTTNBP2, EDNRB, APP, PTK2, ETV1, CLASP2, STRBP, 36 4.04 3.46E-07 510 205 7436 2.56 7.28E-04 3.64E-04 5.98E-04 motion NRG1, DCLK1, PLAT, SGPL1, TGFBR1, EVL, MYH9, YWHAE, NCKAP1, CTNNA2, SEMA6A, EPHA4, NDEL1, FYN, LRP6 PLXNA2, ADCY5, PAX6, GLI3, CTNNB1, LPHN2, EDNRB, LPHN3, APP, CSNK2A1, GPR45, NRG1, RAPGEF1, WWOX, SGPL1, TLE4, SPEN, NCAM1, DDR1, GRB10, GRM3, GNAQ, HIPK1, GNB1, HIPK2, PYGO1, GO:0007166~cell RNF138, ROR2, CNTN1, -

UNIVERSITY of CALIFORNIA, IRVINE Combinatorial Regulation By
UNIVERSITY OF CALIFORNIA, IRVINE Combinatorial regulation by maternal transcription factors during activation of the endoderm gene regulatory network DISSERTATION submitted in partial satisfaction of the requirements for the degree of DOCTOR OF PHILOSOPHY in Biological Sciences by Kitt D. Paraiso Dissertation Committee: Professor Ken W.Y. Cho, Chair Associate Professor Olivier Cinquin Professor Thomas Schilling 2018 Chapter 4 © 2017 Elsevier Ltd. © 2018 Kitt D. Paraiso DEDICATION To the incredibly intelligent and talented people, who in one way or another, helped complete this thesis. ii TABLE OF CONTENTS Page LIST OF FIGURES vii LIST OF TABLES ix LIST OF ABBREVIATIONS X ACKNOWLEDGEMENTS xi CURRICULUM VITAE xii ABSTRACT OF THE DISSERTATION xiv CHAPTER 1: Maternal transcription factors during early endoderm formation in 1 Xenopus Transcription factors co-regulate in a cell type-specific manner 2 Otx1 is expressed in a variety of cell lineages 4 Maternal otx1 in the endodermal conteXt 5 Establishment of enhancers by maternal transcription factors 9 Uncovering the endodermal gene regulatory network 12 Zygotic genome activation and temporal control of gene eXpression 14 The role of maternal transcription factors in early development 18 References 19 CHAPTER 2: Assembly of maternal transcription factors initiates the emergence 26 of tissue-specific zygotic cis-regulatory regions Introduction 28 Identification of maternal vegetally-localized transcription factors 31 Vegt and OtX1 combinatorially regulate the endodermal 33 transcriptome iii -

Accompanies CD8 T Cell Effector Function Global DNA Methylation
Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer, Benjamin G. Barwick, Benjamin A. Youngblood, Rafi Ahmed and Jeremy M. Boss This information is current as of October 1, 2021. J Immunol 2013; 191:3419-3429; Prepublished online 16 August 2013; doi: 10.4049/jimmunol.1301395 http://www.jimmunol.org/content/191/6/3419 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130139 Material 5.DC1 References This article cites 81 articles, 25 of which you can access for free at: http://www.jimmunol.org/content/191/6/3419.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists by guest on October 1, 2021 • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer,* Benjamin G. Barwick,* Benjamin A. Youngblood,*,† Rafi Ahmed,*,† and Jeremy M. -

The Many Roles of MITF in Melanoma
e Cell Bio gl lo n g i y S Vachtenheim, Single Cell Biol 2017, 6:2 Single-Cell Biology DOI: 10.4172/2168-9431.1000162 ISSN: 2168-9431 Mini Review Open Access The Many Roles of MITF in Melanoma Jiri Vachtenheim* Department of Transcription and Cell Signaling, Institute of Medical Biochemistry and Laboratory Diagnostics, First Faculty of Medicine, Charles University and General University Hospital Prague, Czech Republic Abstract Microphthalmia-associated transcription factor (MITF) plays pivotal role in the maintenance of the melanocyte lineage, differentiation of normal and malignant melanocytes and the survival of melanoma cells. MITF regulates expression of many genes with critical functions in cell differentiation, proliferation, and pro-survival properties. Melanoma is an extremely resilient tumor for which no effective therapy exists when the tumor progresses into metastasis. Melanoma is a heterogenous tumor in which the microheterogeneity arises already in the first stages of the tumor development. Because the dependence of the melanocyte lineage on MITF is critical, MITF is regarded as the paradigmatic lineage-addiction oncogene and its gene is amplified in a smaller subset of melanomas. The level of MITF protein greatly differs among the tumor cells. Intriguingly, low MITF level cells are slowly proliferating but constitute an invasive subpopulation of tumor cells. In this minireview, I briefly discuss the many roles and activities of MITF in melanoma cells and the future prospects for melanoma therapy. Keywords: MITF; Melanoma; Phenotype switching; Melanoma was shown to be the necessary epigenetic transcriptional coactivator of proliferation; Differentiation; Invasion; Apoptosis MITF [21] and some of MITF targets [22]. Introduction Importance of MITF for Melanoma Differentiation and Malignant melanoma is a highly aggressive skin cancer, the Proliferation incidence of which is steadily on the rise. -

Supplemental Materials and Methods
Supplemental Material for: Transcriptional silencing of γ-globin by BCL11A involves long-range interactions and cooperation with SOX6 Jian Xu, Vijay G. Sankaran, Min Ni, Tobias F. Menne, Rishi V. Puram, Woojin Kim, Stuart H. Orkin* *To whom correspondence should be addressed. E-mail: [email protected] Supplemental Materials and Methods Flow cytometry Cells at various stages of differentiation were analyzed by flow cytometry using FACSCalibur (BD Biosciences, San Jose, CA). Live cells were identified and gated by exclusion of 7-amino-actinomycin D (7-AAD; BD Pharmingen). The cells were analyzed for expression of cell surface receptors with antibodies specific for CD34, CD45, CD71, CD235, and CD36 conjugated to phycoerythrin (PE), fluorescein isothiocyanate (FITC), or allophycocyanin (APC; BD Pharmingen). Data were analyzed using FlowJo software (Ashland, OR). Cytology Cytocentrifuge preparations were stained with May-Grunwald-Giemsa as previously described (Sankaran et al. 2008). Real-time RT-PCR Real-time quantitative RT-PCR was performed using the iQ SYBR Green Supermix (Bio- Rad). The following primers were used for real-time RT-PCR: human and mouse BCL11A-XL (forward, 5’-ATGCGAGCTGTGCAACTATG-3’; reverse, 5’- GTAAACGTCCTTCCCCACCT-3’), human and mouse BCL11A-L (forward, 5’- CAGCTCAAAAGAGGGCAGAC-3’; reverse, 5’-GAGCTTCCATCCGAAAACTG-3’), and human BCL11A exon 1 and 2 (common between all known isoforms; forward, 5’- AACCCCAGCACTTAAGCAAA-3’; reverse, 5’-GGAGGTCATGATCCCCTTCT-3’). Supplemental Figure Legends Supplemental Figure 1. Expression of BCL11A isoforms in human and mouse erythroid cells. (A) Schematic diagram of human BCL11A isoforms (Liu et al. 2006). The antibodies used for ChIP experiments and their corresponding epitopes are indicated. Locations of primers used for RT-PCR analysis of all BCL11A isoforms (forward and reverse primers indicated by arrowheads), XL and L isoforms are indicated. -

KRAS Drives Immune Evasion in a Genetic Model of Pancreatic Cancer
ARTICLE https://doi.org/10.1038/s41467-021-21736-w OPEN KRAS drives immune evasion in a genetic model of pancreatic cancer Irene Ischenko1, Stephen D’Amico1, Manisha Rao2, Jinyu Li2, Michael J. Hayman1, Scott Powers 2, ✉ ✉ Oleksi Petrenko 1,3 & Nancy C. Reich 1,3 Immune evasion is a hallmark of KRAS-driven cancers, but the underlying causes remain unresolved. Here, we use a mouse model of pancreatic ductal adenocarcinoma to inactivate 1234567890():,; KRAS by CRISPR-mediated genome editing. We demonstrate that at an advanced tumor stage, dependence on KRAS for tumor growth is reduced and is manifested in the sup- pression of antitumor immunity. KRAS-deficient cells retain the ability to form tumors in immunodeficient mice. However, they fail to evade the host immune system in syngeneic wild-type mice, triggering strong antitumor response. We uncover changes both in tumor cells and host immune cells attributable to oncogenic KRAS expression. We identify BRAF and MYC as key mediators of KRAS-driven tumor immune suppression and show that loss of BRAF effectively blocks tumor growth in mice. Applying our results to human PDAC we show that lowering KRAS activity is likewise associated with a more vigorous immune environment. 1 Department of Molecular Genetics and Microbiology, Stony Brook University, Stony Brook, NY, USA. 2 Department of Pathology, Stony Brook University, ✉ Stony Brook, NY, USA. 3These authors jointly supervised this work: Oleksi Petrenko, Nancy C. Reich. email: [email protected]; [email protected] NATURE COMMUNICATIONS | (2021) 12:1482 | https://doi.org/10.1038/s41467-021-21736-w | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | https://doi.org/10.1038/s41467-021-21736-w RAS is frequently associated with some of the deadliest and characterization of KRASG12D p53KO mouse cell lines forms of cancer. -

The Global Gene Expression Profile of the Secondary Transition During Pancreatic Development
ÔØ ÅÒÙ×Ö ÔØ The global gene expression profile of the secondary transition during pancre- atic development Stefanie J. Willmann, Nikola S. Mueller, Silvia Engert, Michael Sterr, Ingo Burtscher, Aurelia Raducanu, Martin Irmler, Johannes Beckers, Steffen Sass, Fabian J. Theis, Heiko Lickert PII: S0925-4773(15)30037-X DOI: doi: 10.1016/j.mod.2015.11.004 Reference: MOD 3386 To appear in: Received date: 19 June 2015 Revised date: 26 November 2015 Accepted date: 27 November 2015 Please cite this article as: Willmann, Stefanie J., Mueller, Nikola S., Engert, Sil- via, Sterr, Michael, Burtscher, Ingo, Raducanu, Aurelia, Irmler, Martin, Beckers, Johannes, Sass, Steffen, Theis, Fabian J., Lickert, Heiko, The global gene expres- sion profile of the secondary transition during pancreatic development, (2015), doi: 10.1016/j.mod.2015.11.004 This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain. ACCEPTED MANUSCRIPT The global gene expression profile of the secondary transition during pancreatic development Stefanie J. Willmann*1,5, Nikola S. Mueller*2, Silvia Engert1, Michael Sterr1, Ingo Burtscher1, Aurelia Raducanu1, Martin Irmler3, Johannes Beckers3,4,5, -

Transcriptional Silencing of G-Globin by BCL11A Involves Long-Range Interactions and Cooperation with SOX6
Downloaded from genesdev.cshlp.org on September 25, 2021 - Published by Cold Spring Harbor Laboratory Press Transcriptional silencing of g-globin by BCL11A involves long-range interactions and cooperation with SOX6 Jian Xu,1,2,3 Vijay G. Sankaran,1,2,4 Min Ni,5 Tobias F. Menne,1,2 Rishi V. Puram,4 Woojin Kim,1,2 and Stuart H. Orkin1,2,3,6 1Division of Hematology/Oncology, Children’s Hospital Boston, Boston, Massachusetts 02115, USA; 2Department of Pediatric Oncology, Dana-Farber Cancer Institute, Harvard Stem Cell Institute, Harvard Medical School, Boston, Massachusetts 02115, USA; 3Howard Hughes Medical Institute, Boston, Massachusetts 02115, USA; 4Harvard-MIT Division of Health Sciences and Technology, Boston, Massachusetts 02115, USA; 5Division of Molecular and Cellular Oncology, Department of Medical Oncology, Dana-Farber Cancer Institute, Harvard Medical School, Boston, Massachusetts 02115, USA The developmental switch from human fetal (g) to adult (b) hemoglobin represents a clinically important example of developmental gene regulation. The transcription factor BCL11A is a central mediator of g-globin silencing and hemoglobin switching. Here we determine chromatin occupancy of BCL11A at the human b-globin locus and other genomic regions in vivo by high-resolution chromatin immunoprecipitation (ChIP)–chip analysis. BCL11A binds the upstream locus control region (LCR), e-globin, and the intergenic regions between g-globin and d-globin genes. A chromosome conformation capture (3C) assay shows that BCL11A reconfigures the b-globin cluster by modulating chromosomal loop formation. We also show that BCL11A and the HMG-box-containing transcription factor SOX6 interact physically and functionally during erythroid maturation. BCL11A and SOX6 co-occupy the human b-globin cluster along with GATA1, and cooperate in silencing g-globin transcription in adult human erythroid progenitors.