Dependent Translational Regulation of the Epithelial
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
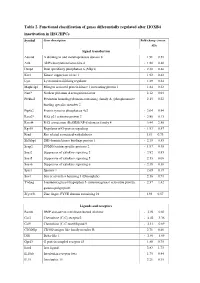
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Allosteric Activation of Functionally Asymmetric RAF Kinase Dimers
Allosteric Activation of Functionally Asymmetric RAF Kinase Dimers Jiancheng Hu,1,2 Edward C. Stites,1 Haiyang Yu,1 Elizabeth A. Germino,1 Hiruy S. Meharena,3,4 Philip J.S. Stork,6 Alexandr P. Kornev,3,4,5 Susan S. Taylor,3,4,5,* and Andrey S. Shaw1,2,* 1Department of Pathology and Immunology 2Howard Hughes Medical Institute Washington University School of Medicine, 660 South Euclid, Box 8118, St. Louis, MO 63110, USA 3Department of Chemistry/Biochemistry 4Department of Pharmacology 5Howard Hughes Medical Institute University of California San Diego, 9500 Gilman Drive, Mail code 0654, La Jolla, CA 92093, USA 6Vollum Institute, Oregon Health & Science University, 3181 SW Sam Jackson Park Road, Portland, OR 97239, USA *Correspondence: [email protected] (S.S.T.), [email protected] (A.S.S.) http://dx.doi.org/10.1016/j.cell.2013.07.046 SUMMARY There are three isoforms of RAF: A-RAF, B-RAF, and C-RAF. Each appears to have a distinct mechanism of activation (Baljuls Although RAF kinases are critical for controlling et al., 2007; Galabova-Kovacs et al., 2006; Wimmer and Baccar- cell growth, their mechanism of activation is incom- ini, 2010). BRAF is considered to be more active and have a pletely understood. Recently, dimerization was simpler mechanism of activation in comparison to CRAF and shown to be important for activation. Here we show ARAF (Chong et al., 2001; Rebocho and Marais, 2013; Zhang that the dimer is functionally asymmetric with one and Guan, 2000). This difference can potentially explain why kinase functioning as an activator to stimulate activ- BRAF mutations are so common in cancer, in contrast to CRAF and ARAF, where mutations are rarely found (Davies ity of the partner, receiver kinase. -

Experimental Chronic Jet Lag Promotes Growth and Lung Metastasis of Lewis Lung Carcinoma in C57BL/6 Mice
ONCOLOGY REPORTS 27: 1417-1428, 2012 Experimental chronic jet lag promotes growth and lung metastasis of Lewis lung carcinoma in C57BL/6 mice 1,2,6 1,3 1,4 1,2 1,5 MINGWEI WU , JING ZENG , YANFENG CHEN , ZHAOLEI ZENG , JINXIN ZHANG , YUCHEN CAI1,2, YANLI YE1,2, LIWU FU1,2, LIJIAN XIAN1,2 and ZHONGPING CHEN1,6 1 2 3 4 State Key Laboratory of Oncology in South China; Departments of Research, Pathology, and Head and Neck Cancer, Cancer Center, Sun Yat-Sen University; 5Department of Medical Statistics and Epidemiology, Sun Yat-Sen University; 6Department of Neurosurgery, Cancer Center, Sun Yat-Sen University, Guangzhou, Guangdong, P.R. China Received December 8, 2011; Accepted January 17, 2012 DOI: 10.3892/or.2012.1688 Abstract. Circadian rhythm has been linked to cancer genesis are governed by a biological clock. The mammalian circadian and development, but the detailed mechanism by which circa- clock contains three components: input pathways, a central dian disruption accelerates tumor growth remains unclear. The pacemaker and output pathways. The mammalian central purpose of this study was to investigate the effect of circadian pacemaker is located in the suprachiasmatic nuclei (SCN) disruption on tumor growth and metastasis in male C57BL/6 of the anterior hypothalamus and controls the activity of the mice, using an experimental chronic jet lag model. Lewis lung peripheral clocks through the neuroendocrine and autonomic carcinoma cells were inoculated into both flanks of the mice nervous systems (1,2). Circadian rhythms govern the rhythmic following 10 days of exposure to experimental chronic jet lag changes in the behavior and/or physiology of mammals, such or control conditions. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

KSR1- and ERK-Dependent Translational Regulation of 2 the Epithelial-To-Mesenchymal Transition
bioRxiv preprint doi: https://doi.org/10.1101/2021.01.18.427224; this version posted January 21, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 KSR1- and ERK-dependent Translational Regulation of 2 the Epithelial-to-Mesenchymal Transition 3 Chaitra Rao1, Danielle E. Frodyma1, Siddesh Southekal2, Robert A. Svoboda3, Adrian R. Black1, 4 Chittibabu Guda2, Tomohiro Mizutani5, Hans Clevers5, Keith R. Johnson1,4, Kurt W. Fisher3, and 5 Robert E. Lewis1* 6 1 Eppley Institute, Fred & Pamela Buffett Cancer Center, University of Nebraska Medical Center, Omaha, NE 68198, USA. 7 2 Department of Genetics, Cell Biology and Anatomy, University of Nebraska Medical Center, Omaha, NE, University of 8 Nebraska Medical Center, Omaha, NE 68198, USA. 9 3 Department of Pathology and Microbiology, University of Nebraska Medical Center, Omaha, NE 68198, USA. 10 4 Department of Oral Biology, University of Nebraska Medical Center, Omaha, NE 68198, USA 11 5 Hubrecht Institute, Royal Netherlands Academy of Arts and Sciences (KNAW) and UMC Utrecht, 3584CT Utrecht, The 12 Netherlands 13 *Correspondence: [email protected] 14 Abstract: 15 The epithelial-to-mesenchymal transition (EMT) is considered a transcriptional process 16 that induces a switch in cells from a polarized state to a migratory phenotype. Here we show that 17 KSR1 and ERK promote EMT through the preferential translation of Epithelial-Stromal 18 Interaction 1 (EPSTI1), which is required to induce the switch from E- to N-cadherin and 19 coordinate migratory and invasive behavior. -

UNIVERSITY of CALIFORNIA, IRVINE Combinatorial Regulation By
UNIVERSITY OF CALIFORNIA, IRVINE Combinatorial regulation by maternal transcription factors during activation of the endoderm gene regulatory network DISSERTATION submitted in partial satisfaction of the requirements for the degree of DOCTOR OF PHILOSOPHY in Biological Sciences by Kitt D. Paraiso Dissertation Committee: Professor Ken W.Y. Cho, Chair Associate Professor Olivier Cinquin Professor Thomas Schilling 2018 Chapter 4 © 2017 Elsevier Ltd. © 2018 Kitt D. Paraiso DEDICATION To the incredibly intelligent and talented people, who in one way or another, helped complete this thesis. ii TABLE OF CONTENTS Page LIST OF FIGURES vii LIST OF TABLES ix LIST OF ABBREVIATIONS X ACKNOWLEDGEMENTS xi CURRICULUM VITAE xii ABSTRACT OF THE DISSERTATION xiv CHAPTER 1: Maternal transcription factors during early endoderm formation in 1 Xenopus Transcription factors co-regulate in a cell type-specific manner 2 Otx1 is expressed in a variety of cell lineages 4 Maternal otx1 in the endodermal conteXt 5 Establishment of enhancers by maternal transcription factors 9 Uncovering the endodermal gene regulatory network 12 Zygotic genome activation and temporal control of gene eXpression 14 The role of maternal transcription factors in early development 18 References 19 CHAPTER 2: Assembly of maternal transcription factors initiates the emergence 26 of tissue-specific zygotic cis-regulatory regions Introduction 28 Identification of maternal vegetally-localized transcription factors 31 Vegt and OtX1 combinatorially regulate the endodermal 33 transcriptome iii -

Kinetworks™ Applications Examples
EXAMPLES OF APPLICATIONS 1 This presentation demonstrated the power of the Kinetworks™ approach to uncover significant research results in a cost effective and efficient manner. 1 Drug Target Discovery A major bottleneck in drug discovery is the identification and validation of drug targets Human genome project has identified over 5,000 cell signalling proteins as potential drug targets Less than 500 proteins have been investigated as drug targets by pharmaceutical industry in its entire history $5 billion spent annually screening non-ideal targets Kinexus is dedicated to identifying and validating drug targets and leads for our clients through offer of our unique Kinetworks™ proteomics services 2 Drug discovery is a very risky and expensive proposition. Only one in a hundred thousand compounds may become a successful drug, after the investment of over US $600 million on average and frequently more than 10 years in development. To discover a drug, the right target must be identified first. This target can be used as the bait to fish for small molecule compounds that are typically inhibitors of the chosen target by high throughput screening. Lead compounds are put sequentially through cell-based, then animal- based and finally human trials. Each subsequent step of the drug discovery and validation process is associated with dramatic increases in cost. Only about 1 in 10 lead compounds that look promising at the end of the animal trials make it successfully through Phase III human trials and receive FDA approval. It is this high rate of failure at the late stages of clinical trials that has made drug discovery so expensive. -

The Many Roles of MITF in Melanoma
e Cell Bio gl lo n g i y S Vachtenheim, Single Cell Biol 2017, 6:2 Single-Cell Biology DOI: 10.4172/2168-9431.1000162 ISSN: 2168-9431 Mini Review Open Access The Many Roles of MITF in Melanoma Jiri Vachtenheim* Department of Transcription and Cell Signaling, Institute of Medical Biochemistry and Laboratory Diagnostics, First Faculty of Medicine, Charles University and General University Hospital Prague, Czech Republic Abstract Microphthalmia-associated transcription factor (MITF) plays pivotal role in the maintenance of the melanocyte lineage, differentiation of normal and malignant melanocytes and the survival of melanoma cells. MITF regulates expression of many genes with critical functions in cell differentiation, proliferation, and pro-survival properties. Melanoma is an extremely resilient tumor for which no effective therapy exists when the tumor progresses into metastasis. Melanoma is a heterogenous tumor in which the microheterogeneity arises already in the first stages of the tumor development. Because the dependence of the melanocyte lineage on MITF is critical, MITF is regarded as the paradigmatic lineage-addiction oncogene and its gene is amplified in a smaller subset of melanomas. The level of MITF protein greatly differs among the tumor cells. Intriguingly, low MITF level cells are slowly proliferating but constitute an invasive subpopulation of tumor cells. In this minireview, I briefly discuss the many roles and activities of MITF in melanoma cells and the future prospects for melanoma therapy. Keywords: MITF; Melanoma; Phenotype switching; Melanoma was shown to be the necessary epigenetic transcriptional coactivator of proliferation; Differentiation; Invasion; Apoptosis MITF [21] and some of MITF targets [22]. Introduction Importance of MITF for Melanoma Differentiation and Malignant melanoma is a highly aggressive skin cancer, the Proliferation incidence of which is steadily on the rise. -

KRAS Drives Immune Evasion in a Genetic Model of Pancreatic Cancer
ARTICLE https://doi.org/10.1038/s41467-021-21736-w OPEN KRAS drives immune evasion in a genetic model of pancreatic cancer Irene Ischenko1, Stephen D’Amico1, Manisha Rao2, Jinyu Li2, Michael J. Hayman1, Scott Powers 2, ✉ ✉ Oleksi Petrenko 1,3 & Nancy C. Reich 1,3 Immune evasion is a hallmark of KRAS-driven cancers, but the underlying causes remain unresolved. Here, we use a mouse model of pancreatic ductal adenocarcinoma to inactivate 1234567890():,; KRAS by CRISPR-mediated genome editing. We demonstrate that at an advanced tumor stage, dependence on KRAS for tumor growth is reduced and is manifested in the sup- pression of antitumor immunity. KRAS-deficient cells retain the ability to form tumors in immunodeficient mice. However, they fail to evade the host immune system in syngeneic wild-type mice, triggering strong antitumor response. We uncover changes both in tumor cells and host immune cells attributable to oncogenic KRAS expression. We identify BRAF and MYC as key mediators of KRAS-driven tumor immune suppression and show that loss of BRAF effectively blocks tumor growth in mice. Applying our results to human PDAC we show that lowering KRAS activity is likewise associated with a more vigorous immune environment. 1 Department of Molecular Genetics and Microbiology, Stony Brook University, Stony Brook, NY, USA. 2 Department of Pathology, Stony Brook University, ✉ Stony Brook, NY, USA. 3These authors jointly supervised this work: Oleksi Petrenko, Nancy C. Reich. email: [email protected]; [email protected] NATURE COMMUNICATIONS | (2021) 12:1482 | https://doi.org/10.1038/s41467-021-21736-w | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | https://doi.org/10.1038/s41467-021-21736-w RAS is frequently associated with some of the deadliest and characterization of KRASG12D p53KO mouse cell lines forms of cancer. -

Ciulli Group Journal Club Targeted Protein Degradation and Other
Ciulli Group Journal Club Targeted protein degradation and Other literature highlights August 2020 Edition Ciulli Group August 2020 Journal Club: edited by Sarath Ramachandran and Angus Cowan Ciulli Journal Club Contents Targeted Protein Degradation 1 - 12 Other highlighted publications 13 - 14 Targeted Protein Degradation Contributor: Angus Structural Insights into PROTAC-Mediated Degradation of Bcl-xL Chun-wa Chung§*, ….., Andrew B. Benowitz ACS Chem. Biol. 2020, 13, 103. Bcl-2 family proteins regulate the intrinsic apoptotic pathway. Pro-survival Bcl-2 family members antagonise the activity of pro-apoptotic Bcl-2 proteins to prevent permeabilization of the mitochondrial outer membrane and subsequent release of apoptogenic factors from the intermembrane space. In cancer, pro-survival Bcl-2 proteins such as Bcl-2 and Bcl-XL are often upregulated, preventing cancer cell death. Structure-based drug design has played an essential role in the development of protein-protein interaction (PPI) inhibitors of pro-survival Bcl-2 family members, with the Bcl-2-selective inhibitor Venclexta recently receiving FDA approval for treatment of chronic lymphocytic leukaemia and small lymphocytic leukaemia. Bcl-XL is overexpressed in many solid tumours and haematological malignancies and is correlated with poor prognosis and resistance to chemotherapy. Although PPI inhibitors of Bcl- XL have been developed, they suffer from on-target platelet toxicity due to platelet dependence on Bcl-XL for survival. Recently, a VHL-based Bcl-XL-degrading PROTAC (DT2216) was demonstrated to circumvent platelet toxicity due to low expression of VHL in platelets, thereby maintaining Bcl-XL levels in platelets and degrading it on other cell types. In addition, while the parent Bcl-XL compound also bound to Bcl-2, PROTAC-induced degradation was Bcl-XL- specific, highlighting the importance of productive ternary complex formation between the target and the E3 that leads to target ubiquitination. -

The Global Gene Expression Profile of the Secondary Transition During Pancreatic Development
ÔØ ÅÒÙ×Ö ÔØ The global gene expression profile of the secondary transition during pancre- atic development Stefanie J. Willmann, Nikola S. Mueller, Silvia Engert, Michael Sterr, Ingo Burtscher, Aurelia Raducanu, Martin Irmler, Johannes Beckers, Steffen Sass, Fabian J. Theis, Heiko Lickert PII: S0925-4773(15)30037-X DOI: doi: 10.1016/j.mod.2015.11.004 Reference: MOD 3386 To appear in: Received date: 19 June 2015 Revised date: 26 November 2015 Accepted date: 27 November 2015 Please cite this article as: Willmann, Stefanie J., Mueller, Nikola S., Engert, Sil- via, Sterr, Michael, Burtscher, Ingo, Raducanu, Aurelia, Irmler, Martin, Beckers, Johannes, Sass, Steffen, Theis, Fabian J., Lickert, Heiko, The global gene expres- sion profile of the secondary transition during pancreatic development, (2015), doi: 10.1016/j.mod.2015.11.004 This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain. ACCEPTED MANUSCRIPT The global gene expression profile of the secondary transition during pancreatic development Stefanie J. Willmann*1,5, Nikola S. Mueller*2, Silvia Engert1, Michael Sterr1, Ingo Burtscher1, Aurelia Raducanu1, Martin Irmler3, Johannes Beckers3,4,5, -

Role of Heat Shock Transcription Factor 1 in Ovarian Cancer
University of South Florida Scholar Commons Graduate Theses and Dissertations Graduate School November 2017 Role of Heat Shock Transcription Factor 1 in Ovarian Cancer Epithelial-Mesenchymal Transition and Drug Sensitivity Chase David Powell University of South Florida, [email protected] Follow this and additional works at: http://scholarcommons.usf.edu/etd Part of the Cell Biology Commons, Molecular Biology Commons, and the Oncology Commons Scholar Commons Citation Powell, Chase David, "Role of Heat Shock Transcription Factor 1 in Ovarian Cancer Epithelial-Mesenchymal Transition and Drug Sensitivity" (2017). Graduate Theses and Dissertations. http://scholarcommons.usf.edu/etd/7079 This Dissertation is brought to you for free and open access by the Graduate School at Scholar Commons. It has been accepted for inclusion in Graduate Theses and Dissertations by an authorized administrator of Scholar Commons. For more information, please contact [email protected]. Role of Heat Shock Transcription Factor 1 in Ovarian Cancer Epithelial-Mesenchymal Transition and Drug Sensitivity by Chase David Powell A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy Department of cell Biology, Microbiology, and Molecular Biology College of Arts and Sciences University of South Florida Major Professor: Sandy D. Westerheide, Ph.D. Brant R. Burkhardt, Ph.D. Younghoon Kee, Ph.D Meera Nanjundan, Ph.D Date of Approval: November 3, 2017 Keywords: Heat Shock Factor 1, Ovarian Cancer, Epithelial to Mesenchymal Transition, Transforming Growth Factor β, HSP90 Inhibitors, Spheroid Culture, Intrinsic Disorder Copyright © 2017, Chase D. Powell DEDICATION I would like to dedicate this work to my wife, Anne T Powell, for all of her support, patience, and love.