Kinetworks™ Applications Examples
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
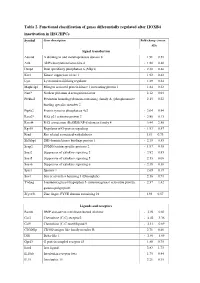
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Allosteric Activation of Functionally Asymmetric RAF Kinase Dimers
Allosteric Activation of Functionally Asymmetric RAF Kinase Dimers Jiancheng Hu,1,2 Edward C. Stites,1 Haiyang Yu,1 Elizabeth A. Germino,1 Hiruy S. Meharena,3,4 Philip J.S. Stork,6 Alexandr P. Kornev,3,4,5 Susan S. Taylor,3,4,5,* and Andrey S. Shaw1,2,* 1Department of Pathology and Immunology 2Howard Hughes Medical Institute Washington University School of Medicine, 660 South Euclid, Box 8118, St. Louis, MO 63110, USA 3Department of Chemistry/Biochemistry 4Department of Pharmacology 5Howard Hughes Medical Institute University of California San Diego, 9500 Gilman Drive, Mail code 0654, La Jolla, CA 92093, USA 6Vollum Institute, Oregon Health & Science University, 3181 SW Sam Jackson Park Road, Portland, OR 97239, USA *Correspondence: [email protected] (S.S.T.), [email protected] (A.S.S.) http://dx.doi.org/10.1016/j.cell.2013.07.046 SUMMARY There are three isoforms of RAF: A-RAF, B-RAF, and C-RAF. Each appears to have a distinct mechanism of activation (Baljuls Although RAF kinases are critical for controlling et al., 2007; Galabova-Kovacs et al., 2006; Wimmer and Baccar- cell growth, their mechanism of activation is incom- ini, 2010). BRAF is considered to be more active and have a pletely understood. Recently, dimerization was simpler mechanism of activation in comparison to CRAF and shown to be important for activation. Here we show ARAF (Chong et al., 2001; Rebocho and Marais, 2013; Zhang that the dimer is functionally asymmetric with one and Guan, 2000). This difference can potentially explain why kinase functioning as an activator to stimulate activ- BRAF mutations are so common in cancer, in contrast to CRAF and ARAF, where mutations are rarely found (Davies ity of the partner, receiver kinase. -

KSR1- and ERK-Dependent Translational Regulation of 2 the Epithelial-To-Mesenchymal Transition
bioRxiv preprint doi: https://doi.org/10.1101/2021.01.18.427224; this version posted January 21, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 KSR1- and ERK-dependent Translational Regulation of 2 the Epithelial-to-Mesenchymal Transition 3 Chaitra Rao1, Danielle E. Frodyma1, Siddesh Southekal2, Robert A. Svoboda3, Adrian R. Black1, 4 Chittibabu Guda2, Tomohiro Mizutani5, Hans Clevers5, Keith R. Johnson1,4, Kurt W. Fisher3, and 5 Robert E. Lewis1* 6 1 Eppley Institute, Fred & Pamela Buffett Cancer Center, University of Nebraska Medical Center, Omaha, NE 68198, USA. 7 2 Department of Genetics, Cell Biology and Anatomy, University of Nebraska Medical Center, Omaha, NE, University of 8 Nebraska Medical Center, Omaha, NE 68198, USA. 9 3 Department of Pathology and Microbiology, University of Nebraska Medical Center, Omaha, NE 68198, USA. 10 4 Department of Oral Biology, University of Nebraska Medical Center, Omaha, NE 68198, USA 11 5 Hubrecht Institute, Royal Netherlands Academy of Arts and Sciences (KNAW) and UMC Utrecht, 3584CT Utrecht, The 12 Netherlands 13 *Correspondence: [email protected] 14 Abstract: 15 The epithelial-to-mesenchymal transition (EMT) is considered a transcriptional process 16 that induces a switch in cells from a polarized state to a migratory phenotype. Here we show that 17 KSR1 and ERK promote EMT through the preferential translation of Epithelial-Stromal 18 Interaction 1 (EPSTI1), which is required to induce the switch from E- to N-cadherin and 19 coordinate migratory and invasive behavior. -

Ciulli Group Journal Club Targeted Protein Degradation and Other
Ciulli Group Journal Club Targeted protein degradation and Other literature highlights August 2020 Edition Ciulli Group August 2020 Journal Club: edited by Sarath Ramachandran and Angus Cowan Ciulli Journal Club Contents Targeted Protein Degradation 1 - 12 Other highlighted publications 13 - 14 Targeted Protein Degradation Contributor: Angus Structural Insights into PROTAC-Mediated Degradation of Bcl-xL Chun-wa Chung§*, ….., Andrew B. Benowitz ACS Chem. Biol. 2020, 13, 103. Bcl-2 family proteins regulate the intrinsic apoptotic pathway. Pro-survival Bcl-2 family members antagonise the activity of pro-apoptotic Bcl-2 proteins to prevent permeabilization of the mitochondrial outer membrane and subsequent release of apoptogenic factors from the intermembrane space. In cancer, pro-survival Bcl-2 proteins such as Bcl-2 and Bcl-XL are often upregulated, preventing cancer cell death. Structure-based drug design has played an essential role in the development of protein-protein interaction (PPI) inhibitors of pro-survival Bcl-2 family members, with the Bcl-2-selective inhibitor Venclexta recently receiving FDA approval for treatment of chronic lymphocytic leukaemia and small lymphocytic leukaemia. Bcl-XL is overexpressed in many solid tumours and haematological malignancies and is correlated with poor prognosis and resistance to chemotherapy. Although PPI inhibitors of Bcl- XL have been developed, they suffer from on-target platelet toxicity due to platelet dependence on Bcl-XL for survival. Recently, a VHL-based Bcl-XL-degrading PROTAC (DT2216) was demonstrated to circumvent platelet toxicity due to low expression of VHL in platelets, thereby maintaining Bcl-XL levels in platelets and degrading it on other cell types. In addition, while the parent Bcl-XL compound also bound to Bcl-2, PROTAC-induced degradation was Bcl-XL- specific, highlighting the importance of productive ternary complex formation between the target and the E3 that leads to target ubiquitination. -

1 Kinobead Profiling Reveals Reprogramming of B-Cell Receptor Signaling in Response to Therapy Within Primary CLL Cells. Linley
bioRxiv preprint doi: https://doi.org/10.1101/841312; this version posted November 14, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Kinobead profiling reveals reprogramming of B-cell receptor signaling in response to therapy within primary CLL cells. Linley AJ1, Griffin R2, Cicconi S2, D’Avola A3$, MacEwan DJ4, Pettit AR1, Kalakonda N1, Packham G3, Prior IA5, Slupsky JR1. 1. Department of Molecular and Clinical Cancer Medicine, Institute of Translational Medicine, University of Liverpool, Sherrington Building, Ashton Street, Liverpool, UK. 2. CRUK Clinical Trials Unit, University of Liverpool, Waterhouse Building, Ashton Street, Liverpool. 3. Southampton Cancer Research UK Centre, Cancer Sciences Unit, Faculty of Medicine, University of Southampton, Southampton, UK. 4. Department of Molecular and Clinical Pharmacology, Institute of Translational Medicine, University of Liverpool, Sherrington Building, Ashton Street, Liverpool, UK. 5. Department of Cellular and Molecular Physiology, Institute of Translational Medicine, University of Liverpool, Nuffield Wing, Crown Street, Liverpool, UK. $ Current Address: The Francis Crick Institute, 1 Midland Road, London, UK. Corresponding author: Dr Adam J Linley, Department of Molecular and Clinical Cancer Medicine, Institute of Translational Medicine, University of Liverpool, Sherrington Building, Ashton Street, Liverpool, UK; [email protected]; +44(0)151 794 5310 Running head: Therapy brings about BCR signal changes. Key points 1. sIgM signaling patterns alter following in vivo therapy using either chemoimmunotherapy or ibrutinib. 2. Kinobeads provide a novel method for high-resolution investigation of signaling in primary CLL cells. 1 bioRxiv preprint doi: https://doi.org/10.1101/841312; this version posted November 14, 2019. -

Dependent Translational Regulation of the Epithelial
RESEARCH ARTICLE KSR1- and ERK- dependent translational regulation of the epithelial- to- mesenchymal transition Chaitra Rao1, Danielle E Frodyma1, Siddesh Southekal2, Robert A Svoboda3, Adrian R Black1, Chittibabu Guda2, Tomohiro Mizutani4, Hans Clevers4, Keith R Johnson1,5, Kurt W Fisher3, Robert E Lewis1* 1Eppley Institute, University of Nebraska Medical Center, Omaha, United States; 2Department of Genetics, Cell Biology and Anatomy, University of Nebraska Medical Center, Omaha, United States; 3Department of Pathology and Microbiology, University of Nebraska Medical Center, Omaha, United States; 4Hubrecht Institute, Royal Netherlands Academy of Arts and Sciences (KNAW) and UMC Utrecht, Utrecht, Netherlands; 5Department of Oral Biology, University of Nebraska Medical Center, Omaha, United States Abstract: The epithelial- to- mesenchymal transition (EMT) is considered a transcriptional process that induces a switch in cells from a polarized state to a migratory phenotype. Here, we show that KSR1 and ERK promote EMT- like phenotype through the preferential translation of Epithelial- Stromal Interaction 1 (EPSTI1), which is required to induce the switch from E- to N- cadherin and coordinate migratory and invasive behavior. EPSTI1 is overexpressed in human colorectal cancer (CRC) cells. Disruption of KSR1 or EPSTI1 significantly impairs cell migration and invasion in vitro, and reverses EMT- like phenotype, in part, by decreasing the expression of N- cadherin and the transcriptional repressors of E- cadherin expression, ZEB1 and Slug. In CRC cells lacking KSR1, ectopic EPSTI1 expression restored the E- to N- cadherin switch, migration, invasion, and anchorage- independent growth. KSR1- dependent induction of EMT- like phenotype via selective translation of *For correspondence: mRNAs reveals its underappreciated role in remodeling the translational landscape of CRC cells to rlewis@ unmc. -

P16 and ARF: Activation of Teenage Proteins in Old Age
p16 and ARF: activation of teenage proteins in old age Ande Satyanarayana, K. Lenhard Rudolph J Clin Invest. 2004;114(9):1237-1240. https://doi.org/10.1172/JCI23437. Commentary Cellular senescence induced by different stresses and telomere shortening appears to play an important role in the aging process. The products of the INK4a/ARF locus — p16INK4a and ARF — arrest cell proliferation at the senescence stage by exerting their effects on retinoblastoma protein– and p53-mediated responsive pathways. A study in this issue of the JCI provides experimental evidence of a specific upregulation of these cell cycle inhibitors in a variety of organs during mammalian aging. Find the latest version: https://jci.me/23437/pdf commentaries 21. Matheny, S.A., et al. 2004. Ras regulates assembly by ceramide-activated protein kinase. Nature. 42. Volle, D.J., et al. 1999. Phosphorylation of the of mitogenic signalling complexes through the 378:307–310. kinase suppressor of ras by associated kinases. Bio- effector protein IMP. Nature. 427:256–260. 32. Mathias, S., Dressler, K.A., and Kolesnick, R.N. chemistry. 38:5130–5137. 22. Wang, X., and Studzinski, G.P. 2001. Phos- 1991. Characterization of a ceramide-activated pro- 43. Zhou, M., Horita, D.A., Waugh, D.S., Byrd, R.A., phorylation of raf-1 by kinase suppressor of ras is tein kinase: stimulation by tumor necrosis factor and Morrison, D.K. 2002. Solution structure and inhibited by “MEK-specific” inhibitors PD 098059 alpha. Proc. Natl. Acad. Sci. U. S. A. 88:10009–10013. functional analysis of the cysteine-rich C1 domain and U0126 in differentiating HL60 cells. -

KSR1 Is Coordinately Regulated with Notch Signaling and Oxidative Phosphorylation in Thyroid Cancer
J LEE and others The role of KSR1 in 54:2 115–124 Research thyroid cancer KSR1 is coordinately regulated with Notch signaling and oxidative phosphorylation in thyroid cancer Jandee Lee1, Mi-Youn Seol1, Seonhyang Jeong2, Hyeong Ju Kwon3, Cho Rok Lee1, 2 1 1 2 Cheol Ryong Ku , Sang-Wook Kang , Jong Ju Jeong , Dong Yeob Shin , Correspondence Kee-Hyun Nam1, Eun Jig Lee2, Woong Youn Chung1 and Young Suk Jo2 should be addressed to Y S Jo Departments of 1Surgery, 2Internal Medicine, and 3Pathology, Yonsei Cancer Center, Severance Hospital, Email Yonsei University College of Medicine, 120-752 Seoul, Korea [email protected] Abstract Kinase suppressor of RAS1 (KSR1) is a scaffold protein implicated in RAS-mediated RAF Key Words activation. However, the molecular function of KSR in papillary thyroid cancer (PTC) is " kinase suppressor of RAS unknown. Thus, this study aimed to characterize the role of KSR1 in patients with PTC. (KSR) qRT-PCR and immunohistochemistry (IHC) revealed inter-tumor heterogeneities in the " connector enhancer of KSR (CNKSR) expression of KSR1 in PTC tissues. Interestingly, BRAFV600E-positive PTC showed higher " thyroid cancer KSR1 mRNA expression than BRAFV600E-negative PTC (P!0.001). Gene Set Enrichment " notch signaling Analysis (GSEA) using public repositories showed that high KSR1 expression coordinately " oxidative phosphorylation upregulated Notch signaling (nominal PZ0.019, false discovery rate (FDR) q-valueZ0.165); this finding was supported by GeneNetwork analysis, indicating that KSR1 expression is positively correlated with NOTCH1 expression (rZ0.677, PZ6.15!10K9). siRNA against KSR1 Journal of Molecular Endocrinology (siKSR1) significantly decreased ERK phosphorylation induced by BRAFV600E, resulting in reduced expression of NOTCH1 and HES1, targets of Notch signaling. -

Functional Signature Ontology-Based Identification and Validation of Novel Therapeutic Targets and Natural Products for the Treatment of Cancer
University of Nebraska Medical Center DigitalCommons@UNMC Theses & Dissertations Graduate Studies Spring 5-5-2018 Functional Signature Ontology-Based Identification and alidationV of Novel Therapeutic Targets and Natural Products for the Treatment of Cancer Beth Neilsen University of Nebraska Medical Center Follow this and additional works at: https://digitalcommons.unmc.edu/etd Part of the Bioinformatics Commons, Cancer Biology Commons, and the Oncology Commons Recommended Citation Neilsen, Beth, "Functional Signature Ontology-Based Identification and alidationV of Novel Therapeutic Targets and Natural Products for the Treatment of Cancer" (2018). Theses & Dissertations. 268. https://digitalcommons.unmc.edu/etd/268 This Dissertation is brought to you for free and open access by the Graduate Studies at DigitalCommons@UNMC. It has been accepted for inclusion in Theses & Dissertations by an authorized administrator of DigitalCommons@UNMC. For more information, please contact [email protected]. Functional Signature Ontology-Based Identification and Validation of Novel Therapeutic Targets and Natural Products for the Treatment of Cancer By Beth K. Neilsen A DISSERTATION Presented to the Faculty of the University of Nebraska Graduate College in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy Cancer Research Graduate Program Under the Supervision of Professor Robert E. Lewis University of Nebraska Medical Center Omaha, Nebraska May 2018 Supervisory Committee: Jennifer Black, Ph.D. Jing (Jenny) Wang, Ph.D. Allison Cushman-Vokoun, M.D./Ph.D. Juan Cui, Ph.D. ii To my parents, Dr. and Mrs. Mitchell and Rebecca Neilsen, For always encouraging me to think critically, pushing me to be the best version of me, for giving me the space to think for myself, teaching me compassion, and most importantly, for molding me into the person I am today. -

Identification of KSR1 As a Novel Target and Decoding Tyrosine Kinase Proteome in Breast Cancer
Identification of KSR1 as a novel target and decoding tyrosine kinase proteome in breast cancer By Hua Zhang A thesis submitted for the degree of Doctor of Philosophy at Imperial College London Faculty of Medicine, Department of Surgery and Cancer, 1st floor ICTEM Imperial College London Hammersmith Hospital, Du Cane Road London, W12 0NN Supervisor: Prof Justin Stebbing Co-supervisor: Dr Georgios Giamas Dedicated to my family and everyone who loves me to the 1,278 days to the stars in the night and the beautiful eyes in my heart 1 Abstract Kinase suppressor of Ras-1 (KSR1), originally identified as a novel protein kinase in the Ras- Raf cascade, plays a role in activation of mitogen-activated protein kinases (MAPKs). Although efforts have been devoted to study the role of KSR1 in certain tumour types, its biological functions in breast cancer have remained largely undefined. A SILAC-based proteomic analysis was conducted to identify the KSR1-regulated phospho- proteins in breast cancer. Our results revealed that KSR1 overexpression decreases deleted in breast cancer 1 (DBC1) phosphorylation. We then demonstrated that KSR1 decreases transcriptional activity of p53 by reducing phosphorylation of DBC1, which leads to a reduced interaction of DBC1 with sirtuin 1 (SIRT1); this in turn enables SIRT1 to deacetylate p53. We further examined the correlation between KSR1 expression and clinical outcome in breast cancer. Our results showed that patients with breast cancer with high KSR1 in our cohort (n>1000) had better disease free- and overall survival. Moreover, in KSR1-transfected stable cells, fewer and smaller size colonies were formed in comparison to parental cells, while an in vivo study demonstrated that the growth of xenograft tumours overexpressing KSR1 was inhibited. -

Mutations and Deregulation of Ras/Raf/MEK/ERK and PI3K/ PTEN/Akt/Mtor Cascades Which Alter Therapy Response
www.impactjournals.com/oncotarget/ Oncotarget, September, Vol.3, No 9 Mutations and Deregulation of Ras/Raf/MEK/ERK and PI3K/ PTEN/Akt/mTOR Cascades Which Alter Therapy Response. James A. McCubrey1, Linda S. Steelman1, William H. Chappell1, Stephen L. Abrams1, Giuseppe Montalto2, Melchiorre Cervello3, Ferdinando Nicoletti4, Paolo Fagone4, Grazia Malaponte4, Maria C. Mazzarino4, Saverio Candido4, Massimo Libra4, Jörg Bäsecke5, Sanja Mijatovic6, Danijela Maksimovic-Ivanic6 Michele Milella7, Agostino Tafuri8 , Lucio Cocco9, Camilla Evangelisti10, Francesca Chiarini10, Alberto M. Martelli9,10 1 Department of Microbiology and Immunology, Brody School of Medicine at East Carolina University, Greenville, NC, USA 2 Department of Internal Medicine and Specialties, University of Palermo, Palermo, Italy 3 Consiglio Nazionale delle Ricerche, Istituto di Biomedicina e Immunologia Molecolare “Alberto Monroy”, Palermo, Italy 4 Department of Biomedical Sciences, University of Catania, Catania, Italy 5 Department of Medicine, University of Göttingen, Göttingen, Germany 6 Department of Immunology, Instititue for Biological Research “Sinisa Stankovic”, University of Belgrade, Belgrade, Serbia 7 Regina Elena National Cancer Institute, Rome, Italy 8 Sapienza, University of Rome, Department of Cellular Biotechnology and Hematology, Rome, Italy 9 Department of Biomedical and Neuromotor Sciences, Università di Bologna, Bologna, Italy 10 Institute of Molecular Genetics, National Research Council-Rizzoli Orthopedic Institute, Bologna, Italy Correspondence to: James A. McCubrey, email: [email protected] Keywords: Targeted Therapy, Therapy Resistance, Mutations, Raf, Akt, PI3K, mTOR Received: August 12, 2012, Accepted: September 17, 2012, Published: September 20, 2012 Copyright: © McCubrey et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. -

KSR1 Antibody A
Revision 1 C 0 2 - t KSR1 Antibody a e r o t S Orders: 877-616-CELL (2355) [email protected] Support: 877-678-TECH (8324) 0 4 Web: [email protected] 6 www.cellsignal.com 4 # 3 Trask Lane Danvers Massachusetts 01923 USA For Research Use Only. Not For Use In Diagnostic Procedures. Applications: Reactivity: Sensitivity: MW (kDa): Source: UniProt ID: Entrez-Gene Id: WB H M R Mk Endogenous 100 Rabbit Q8IVT5 8844 Product Usage Information 4. Michaud, N. R. et al. (1997) Proc. Natl. Acad. Sci. USA 94, 12792-12796. 5. Stewart, S. et al. (1999) Mol. Cell. Biol. 19, 5523-5534. Application Dilution 6. Muller, J. et al. (2001) Mol. Cell 8, 983-993. 7. Cacace, A. M. et al. (1999) Mol. Cell. Biol. 19, 229-240. Western Blotting 1:1000 8. Ory, S. et al. (2003) Curr. Biol. 13, 1356-1364. 9. Matheny, S. A. et al. (2004) Nature 427, 256-260. Storage 10. Kortum, R.L. and Lewis, R.E. (2004) Mol Cell Biol 24, 4407-16. Supplied in 10 mM sodium HEPES (pH 7.5), 150 mM NaCl, 100 µg/ml BSA and 50% glycerol. Store at –20°C. Do not aliquot the antibody. Specificity / Sensitivity KSR1 Antibody detects endogenous levels of total KSR1 protein. Species Reactivity: Human, Mouse, Rat, Monkey Source / Purification Polyclonal antibodies are produced by immunizing animals with a synthetic peptide corresponding to the region surrounding Gly910 of human KSR1. Antibodies are purified by protein A and peptide affinity chromatography. Background KSR1 (kinase supressor of Ras) was identified from a genetic screen in Drosophila and C.