Identification of KSR1 As a Novel Target and Decoding Tyrosine Kinase Proteome in Breast Cancer
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -
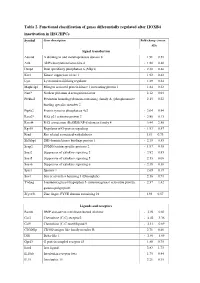
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Viewed Under 23 (B) Or 203 (C) fi M M Male Cko Mice, and Largely Unaffected Magni Cation; Scale Bars, 500 M (B) and 50 M (C)
BRIEF COMMUNICATION www.jasn.org Renal Fanconi Syndrome and Hypophosphatemic Rickets in the Absence of Xenotropic and Polytropic Retroviral Receptor in the Nephron Camille Ansermet,* Matthias B. Moor,* Gabriel Centeno,* Muriel Auberson,* † † ‡ Dorothy Zhang Hu, Roland Baron, Svetlana Nikolaeva,* Barbara Haenzi,* | Natalya Katanaeva,* Ivan Gautschi,* Vladimir Katanaev,*§ Samuel Rotman, Robert Koesters,¶ †† Laurent Schild,* Sylvain Pradervand,** Olivier Bonny,* and Dmitri Firsov* BRIEF COMMUNICATION *Department of Pharmacology and Toxicology and **Genomic Technologies Facility, University of Lausanne, Lausanne, Switzerland; †Department of Oral Medicine, Infection, and Immunity, Harvard School of Dental Medicine, Boston, Massachusetts; ‡Institute of Evolutionary Physiology and Biochemistry, St. Petersburg, Russia; §School of Biomedicine, Far Eastern Federal University, Vladivostok, Russia; |Services of Pathology and ††Nephrology, Department of Medicine, University Hospital of Lausanne, Lausanne, Switzerland; and ¶Université Pierre et Marie Curie, Paris, France ABSTRACT Tight control of extracellular and intracellular inorganic phosphate (Pi) levels is crit- leaves.4 Most recently, Legati et al. have ical to most biochemical and physiologic processes. Urinary Pi is freely filtered at the shown an association between genetic kidney glomerulus and is reabsorbed in the renal tubule by the action of the apical polymorphisms in Xpr1 and primary fa- sodium-dependent phosphate transporters, NaPi-IIa/NaPi-IIc/Pit2. However, the milial brain calcification disorder.5 How- molecular identity of the protein(s) participating in the basolateral Pi efflux remains ever, the role of XPR1 in the maintenance unknown. Evidence has suggested that xenotropic and polytropic retroviral recep- of Pi homeostasis remains unknown. Here, tor 1 (XPR1) might be involved in this process. Here, we show that conditional in- we addressed this issue in mice deficient for activation of Xpr1 in the renal tubule in mice resulted in impaired renal Pi Xpr1 in the nephron. -

Two Locus Inheritance of Non-Syndromic Midline Craniosynostosis Via Rare SMAD6 and 4 Common BMP2 Alleles 5 6 Andrew T
1 2 3 Two locus inheritance of non-syndromic midline craniosynostosis via rare SMAD6 and 4 common BMP2 alleles 5 6 Andrew T. Timberlake1-3, Jungmin Choi1,2, Samir Zaidi1,2, Qiongshi Lu4, Carol Nelson- 7 Williams1,2, Eric D. Brooks3, Kaya Bilguvar1,5, Irina Tikhonova5, Shrikant Mane1,5, Jenny F. 8 Yang3, Rajendra Sawh-Martinez3, Sarah Persing3, Elizabeth G. Zellner3, Erin Loring1,2,5, Carolyn 9 Chuang3, Amy Galm6, Peter W. Hashim3, Derek M. Steinbacher3, Michael L. DiLuna7, Charles 10 C. Duncan7, Kevin A. Pelphrey8, Hongyu Zhao4, John A. Persing3, Richard P. Lifton1,2,5,9 11 12 1Department of Genetics, Yale University School of Medicine, New Haven, CT, USA 13 2Howard Hughes Medical Institute, Yale University School of Medicine, New Haven, CT, USA 14 3Section of Plastic and Reconstructive Surgery, Department of Surgery, Yale University School of Medicine, New Haven, CT, USA 15 4Department of Biostatistics, Yale University School of Medicine, New Haven, CT, USA 16 5Yale Center for Genome Analysis, New Haven, CT, USA 17 6Craniosynostosis and Positional Plagiocephaly Support, New York, NY, USA 18 7Department of Neurosurgery, Yale University School of Medicine, New Haven, CT, USA 19 8Child Study Center, Yale University School of Medicine, New Haven, CT, USA 20 9The Rockefeller University, New York, NY, USA 21 22 ABSTRACT 23 Premature fusion of the cranial sutures (craniosynostosis), affecting 1 in 2,000 24 newborns, is treated surgically in infancy to prevent adverse neurologic outcomes. To 25 identify mutations contributing to common non-syndromic midline (sagittal and metopic) 26 craniosynostosis, we performed exome sequencing of 132 parent-offspring trios and 59 27 additional probands. -

Allosteric Activation of Functionally Asymmetric RAF Kinase Dimers
Allosteric Activation of Functionally Asymmetric RAF Kinase Dimers Jiancheng Hu,1,2 Edward C. Stites,1 Haiyang Yu,1 Elizabeth A. Germino,1 Hiruy S. Meharena,3,4 Philip J.S. Stork,6 Alexandr P. Kornev,3,4,5 Susan S. Taylor,3,4,5,* and Andrey S. Shaw1,2,* 1Department of Pathology and Immunology 2Howard Hughes Medical Institute Washington University School of Medicine, 660 South Euclid, Box 8118, St. Louis, MO 63110, USA 3Department of Chemistry/Biochemistry 4Department of Pharmacology 5Howard Hughes Medical Institute University of California San Diego, 9500 Gilman Drive, Mail code 0654, La Jolla, CA 92093, USA 6Vollum Institute, Oregon Health & Science University, 3181 SW Sam Jackson Park Road, Portland, OR 97239, USA *Correspondence: [email protected] (S.S.T.), [email protected] (A.S.S.) http://dx.doi.org/10.1016/j.cell.2013.07.046 SUMMARY There are three isoforms of RAF: A-RAF, B-RAF, and C-RAF. Each appears to have a distinct mechanism of activation (Baljuls Although RAF kinases are critical for controlling et al., 2007; Galabova-Kovacs et al., 2006; Wimmer and Baccar- cell growth, their mechanism of activation is incom- ini, 2010). BRAF is considered to be more active and have a pletely understood. Recently, dimerization was simpler mechanism of activation in comparison to CRAF and shown to be important for activation. Here we show ARAF (Chong et al., 2001; Rebocho and Marais, 2013; Zhang that the dimer is functionally asymmetric with one and Guan, 2000). This difference can potentially explain why kinase functioning as an activator to stimulate activ- BRAF mutations are so common in cancer, in contrast to CRAF and ARAF, where mutations are rarely found (Davies ity of the partner, receiver kinase. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

KSR1- and ERK-Dependent Translational Regulation of 2 the Epithelial-To-Mesenchymal Transition
bioRxiv preprint doi: https://doi.org/10.1101/2021.01.18.427224; this version posted January 21, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 KSR1- and ERK-dependent Translational Regulation of 2 the Epithelial-to-Mesenchymal Transition 3 Chaitra Rao1, Danielle E. Frodyma1, Siddesh Southekal2, Robert A. Svoboda3, Adrian R. Black1, 4 Chittibabu Guda2, Tomohiro Mizutani5, Hans Clevers5, Keith R. Johnson1,4, Kurt W. Fisher3, and 5 Robert E. Lewis1* 6 1 Eppley Institute, Fred & Pamela Buffett Cancer Center, University of Nebraska Medical Center, Omaha, NE 68198, USA. 7 2 Department of Genetics, Cell Biology and Anatomy, University of Nebraska Medical Center, Omaha, NE, University of 8 Nebraska Medical Center, Omaha, NE 68198, USA. 9 3 Department of Pathology and Microbiology, University of Nebraska Medical Center, Omaha, NE 68198, USA. 10 4 Department of Oral Biology, University of Nebraska Medical Center, Omaha, NE 68198, USA 11 5 Hubrecht Institute, Royal Netherlands Academy of Arts and Sciences (KNAW) and UMC Utrecht, 3584CT Utrecht, The 12 Netherlands 13 *Correspondence: [email protected] 14 Abstract: 15 The epithelial-to-mesenchymal transition (EMT) is considered a transcriptional process 16 that induces a switch in cells from a polarized state to a migratory phenotype. Here we show that 17 KSR1 and ERK promote EMT through the preferential translation of Epithelial-Stromal 18 Interaction 1 (EPSTI1), which is required to induce the switch from E- to N-cadherin and 19 coordinate migratory and invasive behavior. -

Kinetworks™ Applications Examples
EXAMPLES OF APPLICATIONS 1 This presentation demonstrated the power of the Kinetworks™ approach to uncover significant research results in a cost effective and efficient manner. 1 Drug Target Discovery A major bottleneck in drug discovery is the identification and validation of drug targets Human genome project has identified over 5,000 cell signalling proteins as potential drug targets Less than 500 proteins have been investigated as drug targets by pharmaceutical industry in its entire history $5 billion spent annually screening non-ideal targets Kinexus is dedicated to identifying and validating drug targets and leads for our clients through offer of our unique Kinetworks™ proteomics services 2 Drug discovery is a very risky and expensive proposition. Only one in a hundred thousand compounds may become a successful drug, after the investment of over US $600 million on average and frequently more than 10 years in development. To discover a drug, the right target must be identified first. This target can be used as the bait to fish for small molecule compounds that are typically inhibitors of the chosen target by high throughput screening. Lead compounds are put sequentially through cell-based, then animal- based and finally human trials. Each subsequent step of the drug discovery and validation process is associated with dramatic increases in cost. Only about 1 in 10 lead compounds that look promising at the end of the animal trials make it successfully through Phase III human trials and receive FDA approval. It is this high rate of failure at the late stages of clinical trials that has made drug discovery so expensive. -

Genome-Wide DNA Methylation Analysis of KRAS Mutant Cell Lines Ben Yi Tew1,5, Joel K
www.nature.com/scientificreports OPEN Genome-wide DNA methylation analysis of KRAS mutant cell lines Ben Yi Tew1,5, Joel K. Durand2,5, Kirsten L. Bryant2, Tikvah K. Hayes2, Sen Peng3, Nhan L. Tran4, Gerald C. Gooden1, David N. Buckley1, Channing J. Der2, Albert S. Baldwin2 ✉ & Bodour Salhia1 ✉ Oncogenic RAS mutations are associated with DNA methylation changes that alter gene expression to drive cancer. Recent studies suggest that DNA methylation changes may be stochastic in nature, while other groups propose distinct signaling pathways responsible for aberrant methylation. Better understanding of DNA methylation events associated with oncogenic KRAS expression could enhance therapeutic approaches. Here we analyzed the basal CpG methylation of 11 KRAS-mutant and dependent pancreatic cancer cell lines and observed strikingly similar methylation patterns. KRAS knockdown resulted in unique methylation changes with limited overlap between each cell line. In KRAS-mutant Pa16C pancreatic cancer cells, while KRAS knockdown resulted in over 8,000 diferentially methylated (DM) CpGs, treatment with the ERK1/2-selective inhibitor SCH772984 showed less than 40 DM CpGs, suggesting that ERK is not a broadly active driver of KRAS-associated DNA methylation. KRAS G12V overexpression in an isogenic lung model reveals >50,600 DM CpGs compared to non-transformed controls. In lung and pancreatic cells, gene ontology analyses of DM promoters show an enrichment for genes involved in diferentiation and development. Taken all together, KRAS-mediated DNA methylation are stochastic and independent of canonical downstream efector signaling. These epigenetically altered genes associated with KRAS expression could represent potential therapeutic targets in KRAS-driven cancer. Activating KRAS mutations can be found in nearly 25 percent of all cancers1. -

Inhibition of Mitochondrial Complex II in Neuronal Cells Triggers Unique
www.nature.com/scientificreports OPEN Inhibition of mitochondrial complex II in neuronal cells triggers unique pathways culminating in autophagy with implications for neurodegeneration Sathyanarayanan Ranganayaki1, Neema Jamshidi2, Mohamad Aiyaz3, Santhosh‑Kumar Rashmi4, Narayanappa Gayathri4, Pulleri Kandi Harsha5, Balasundaram Padmanabhan6 & Muchukunte Mukunda Srinivas Bharath7* Mitochondrial dysfunction and neurodegeneration underlie movement disorders such as Parkinson’s disease, Huntington’s disease and Manganism among others. As a corollary, inhibition of mitochondrial complex I (CI) and complex II (CII) by toxins 1‑methyl‑4‑phenylpyridinium (MPP+) and 3‑nitropropionic acid (3‑NPA) respectively, induced degenerative changes noted in such neurodegenerative diseases. We aimed to unravel the down‑stream pathways associated with CII inhibition and compared with CI inhibition and the Manganese (Mn) neurotoxicity. Genome‑wide transcriptomics of N27 neuronal cells exposed to 3‑NPA, compared with MPP+ and Mn revealed varied transcriptomic profle. Along with mitochondrial and synaptic pathways, Autophagy was the predominant pathway diferentially regulated in the 3‑NPA model with implications for neuronal survival. This pathway was unique to 3‑NPA, as substantiated by in silico modelling of the three toxins. Morphological and biochemical validation of autophagy markers in the cell model of 3‑NPA revealed incomplete autophagy mediated by mechanistic Target of Rapamycin Complex 2 (mTORC2) pathway. Interestingly, Brain Derived Neurotrophic Factor -

A Raf-Induced Allosteric Transition of KSR Stimulates Phosphorylation of MEK
LETTER doi:10.1038/nature09860 A Raf-induced allosteric transition of KSR stimulates phosphorylation of MEK Damian F. Brennan1*,ArvinC.Dar2*, Nicholas T. Hertz2, William C. H. Chao1, Alma L. Burlingame3, Kevan M. Shokat2 &DavidBarford1 In metazoans, the Ras–Raf–MEK (mitogen-activated protein- (Supplementary Fig. 3), indicates that MEK inhibitors engage a physio- kinase kinase)–ERK (extracellular signal-regulated kinase) signal- logically relevant conformation of the kinase. ling pathway relays extracellular stimuli to elicit changes in cellular MEK and KSR form constitutive complexes14,15 stable to Raf phos- function and gene expression. Aberrant activation of this pathway phorylation16,17, even though Ser 218M and Ser 222M phosphorylation through oncogenic mutations is responsible for a large proportion would alter the conformation of the MEK1 activation segment. The of human cancer. Kinase suppressor of Ras (KSR)1–3 functions as integrity of the complex is probably conferred by the more extensive an essential scaffolding protein to coordinate the assembly of Raf– interface created by engagement of their respective aG helices (Fig. 1a, MEK–ERK complexes4,5. Here we integrate structural and bio- c). This is consistent with studies showing that mutation of MEK chemical studies to understand how KSR promotes stimulatory within a conserved hydrophobic motif (Met 308M to Ile 310M), con- Raf phosphorylation of MEK (refs 6, 7). We show, from the crystal tiguous with the aG helix, disrupts MEK–KSR1 interactions17. In the structure of the kinase domain of human KSR2 (KSR2(KD)) in complex with rabbit MEK1, that interactions between KSR2(KD) a and MEK1 are mediated by their respective activation segments G683E/V and C-lobe aG helices. -

Ciulli Group Journal Club Targeted Protein Degradation and Other
Ciulli Group Journal Club Targeted protein degradation and Other literature highlights August 2020 Edition Ciulli Group August 2020 Journal Club: edited by Sarath Ramachandran and Angus Cowan Ciulli Journal Club Contents Targeted Protein Degradation 1 - 12 Other highlighted publications 13 - 14 Targeted Protein Degradation Contributor: Angus Structural Insights into PROTAC-Mediated Degradation of Bcl-xL Chun-wa Chung§*, ….., Andrew B. Benowitz ACS Chem. Biol. 2020, 13, 103. Bcl-2 family proteins regulate the intrinsic apoptotic pathway. Pro-survival Bcl-2 family members antagonise the activity of pro-apoptotic Bcl-2 proteins to prevent permeabilization of the mitochondrial outer membrane and subsequent release of apoptogenic factors from the intermembrane space. In cancer, pro-survival Bcl-2 proteins such as Bcl-2 and Bcl-XL are often upregulated, preventing cancer cell death. Structure-based drug design has played an essential role in the development of protein-protein interaction (PPI) inhibitors of pro-survival Bcl-2 family members, with the Bcl-2-selective inhibitor Venclexta recently receiving FDA approval for treatment of chronic lymphocytic leukaemia and small lymphocytic leukaemia. Bcl-XL is overexpressed in many solid tumours and haematological malignancies and is correlated with poor prognosis and resistance to chemotherapy. Although PPI inhibitors of Bcl- XL have been developed, they suffer from on-target platelet toxicity due to platelet dependence on Bcl-XL for survival. Recently, a VHL-based Bcl-XL-degrading PROTAC (DT2216) was demonstrated to circumvent platelet toxicity due to low expression of VHL in platelets, thereby maintaining Bcl-XL levels in platelets and degrading it on other cell types. In addition, while the parent Bcl-XL compound also bound to Bcl-2, PROTAC-induced degradation was Bcl-XL- specific, highlighting the importance of productive ternary complex formation between the target and the E3 that leads to target ubiquitination.