Mouse Fcrl1 Conditional Knockout Project (CRISPR/Cas9)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Rabbit Anti-FCRL1 Antibody-SL16064R
SunLong Biotech Co.,LTD Tel: 0086-571- 56623320 Fax:0086-571- 56623318 E-mail:[email protected] www.sunlongbiotech.com Rabbit Anti-FCRL1 antibody SL16064R Product Name: FCRL1 Chinese Name: CD307a抗体 CD307a; Fc receptor homolog 1; Fc receptor like protein 1; Fc receptor-like protein 1; FcR like protein 1; FcR-like protein 1; FCRH 1; FcRH1; FCRL 1; FcRL1; FCRL1_HUMAN; hIFGP 1; hIFGP1; IFGP 1; IFGP family protein 1; IFGP1; Immune Alias: receptor translocation associated protein 5; Immune receptor translocation-associated protein 5; immunoglobulin superfamily Fc receptor, gp42; IRTA 5; IRTA5; RP11 367J7.7. Organism Species: Rabbit Clonality: Polyclonal React Species: Human, WB=1:500-2000ELISA=1:500-1000IHC-P=1:400-800IHC-F=1:400-800ICC=1:100- 500IF=1:100-500(Paraffin sections need antigen repair) Applications: not yet tested in other applications. optimal dilutions/concentrations should be determined by the end user. Molecular weight: 45kDa Cellular localization: The cell membrane Form: Lyophilizedwww.sunlongbiotech.com or Liquid Concentration: 1mg/ml KLH conjugated synthetic peptide derived from human FCRL1:251- immunogen: 350/429<Extracellular> Lsotype: IgG Purification: affinity purified by Protein A Storage Buffer: 0.01M TBS(pH7.4) with 1% BSA, 0.03% Proclin300 and 50% Glycerol. Store at -20 °C for one year. Avoid repeated freeze/thaw cycles. The lyophilized antibody is stable at room temperature for at least one month and for greater than a year Storage: when kept at -20°C. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 °C. -
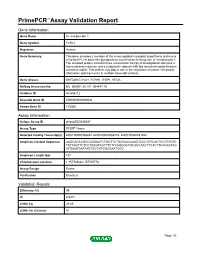
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name Fc receptor-like 1 Gene Symbol FCRL1 Organism Human Gene Summary This gene encodes a member of the immunoglobulin receptor superfamily and is one of several Fc receptor-like glycoproteins clustered on the long arm of chromosome 1. The encoded protein contains three extracellular C2-like immunoglobulin domains a transmembrane domain and a cytoplasmic domain with two immunoreceptor-tyrosine activation motifs. This protein may play a role in the regulation of cancer cell growth. Alternative splicing results in multiple transcript variants. Gene Aliases DKFZp667O1421, FCRH1, IFGP1, IRTA5 RefSeq Accession No. NC_000001.10, NT_004487.19 UniGene ID Hs.656112 Ensembl Gene ID ENSG00000163534 Entrez Gene ID 115350 Assay Information Unique Assay ID qHsaCED0046097 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000358292, ENST00000368176, ENST00000491942 Amplicon Context Sequence AAGGAGCCGGCAGGAATCTGGTTCTGATAACAAAGTCCCTGTCACTCCTGTGTC TATTAGTTCTCCTAGGTAGTTTCTTCAGGGCTGCGCCAACTTCACTTCACAGTAG ATGAAGGAATAGTGCCATGGAGAATGGC Amplicon Length (bp) 107 Chromosome Location 1:157765642-157765778 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 99 R2 0.9971 cDNA Cq 29.05 cDNA Tm (Celsius) 81 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq 25.04 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report FCRL1, Human Amplification Plot Amplification of cDNA generated from 25 ng -

Fc Receptors for Immunoglobulins and Their Appearance During Vertebrate Evolution
Fc Receptors for Immunoglobulins and Their Appearance during Vertebrate Evolution Srinivas Akula, Sayran Mohammadamin¤, Lars Hellman* Department of Cell and Molecular Biology, Uppsala University, The Biomedical Center, Uppsala, Sweden Abstract Receptors interacting with the constant domain of immunoglobulins (Igs) have a number of important functions in vertebrates. They facilitate phagocytosis by opsonization, are key components in antibody-dependent cellular cytotoxicity as well as activating cells to release granules. In mammals, four major types of classical Fc receptors (FcRs) for IgG have been identified, one high-affinity receptor for IgE, one for both IgM and IgA, one for IgM and one for IgA. All of these receptors are related in structure and all of them, except the IgA receptor, are found in primates on chromosome 1, indicating that they originate from a common ancestor by successive gene duplications. The number of Ig isotypes has increased gradually during vertebrate evolution and this increase has likely been accompanied by a similar increase in isotype-specific receptors. To test this hypothesis we have performed a detailed bioinformatics analysis of a panel of vertebrate genomes. The first components to appear are the poly-Ig receptors (PIGRs), receptors similar to the classic FcRs in mammals, so called FcRL receptors, and the FcR c chain. These molecules are not found in cartilagous fish and may first appear within bony fishes, indicating a major step in Fc receptor evolution at the appearance of bony fish. In contrast, the receptor for IgA is only found in placental mammals, indicating a relatively late appearance. The IgM and IgA/M receptors are first observed in the monotremes, exemplified by the platypus, indicating an appearance during early mammalian evolution. -

Mouse Fcrl1 Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Fcrl1 Knockout Project (CRISPR/Cas9) Objective: To create a Fcrl1 knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Fcrl1 gene (NCBI Reference Sequence: NM_153090 ; Ensembl: ENSMUSG00000059994 ) is located on Mouse chromosome 3. 11 exons are identified, with the ATG start codon in exon 1 and the TAG stop codon in exon 11 (Transcript: ENSMUST00000163661). Exon 2~10 will be selected as target site. Cas9 and gRNA will be co-injected into fertilized eggs for KO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Exon 2 starts from about 2.94% of the coding region. Exon 2~10 covers 90.27% of the coding region. The size of effective KO region: ~8937 bp. The KO region does not have any other known gene. Page 1 of 9 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele 5' gRNA region gRNA region 3' 1 2 3 4 5 6 7 8 9 10 11 Legends Exon of mouse Fcrl1 Knockout region Page 2 of 9 https://www.alphaknockout.com Overview of the Dot Plot (up) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 2000 bp section upstream of Exon 2 is aligned with itself to determine if there are tandem repeats. Tandem repeats are found in the dot plot matrix. The gRNA site is selected outside of these tandem repeats. Overview of the Dot Plot (down) Window size: 15 bp Forward Reverse Complement Sequence 12 Note: The 952 bp section downstream of Exon 10 is aligned with itself to determine if there are tandem repeats. -

Table S1. 103 Ferroptosis-Related Genes Retrieved from the Genecards
Table S1. 103 ferroptosis-related genes retrieved from the GeneCards. Gene Symbol Description Category GPX4 Glutathione Peroxidase 4 Protein Coding AIFM2 Apoptosis Inducing Factor Mitochondria Associated 2 Protein Coding TP53 Tumor Protein P53 Protein Coding ACSL4 Acyl-CoA Synthetase Long Chain Family Member 4 Protein Coding SLC7A11 Solute Carrier Family 7 Member 11 Protein Coding VDAC2 Voltage Dependent Anion Channel 2 Protein Coding VDAC3 Voltage Dependent Anion Channel 3 Protein Coding ATG5 Autophagy Related 5 Protein Coding ATG7 Autophagy Related 7 Protein Coding NCOA4 Nuclear Receptor Coactivator 4 Protein Coding HMOX1 Heme Oxygenase 1 Protein Coding SLC3A2 Solute Carrier Family 3 Member 2 Protein Coding ALOX15 Arachidonate 15-Lipoxygenase Protein Coding BECN1 Beclin 1 Protein Coding PRKAA1 Protein Kinase AMP-Activated Catalytic Subunit Alpha 1 Protein Coding SAT1 Spermidine/Spermine N1-Acetyltransferase 1 Protein Coding NF2 Neurofibromin 2 Protein Coding YAP1 Yes1 Associated Transcriptional Regulator Protein Coding FTH1 Ferritin Heavy Chain 1 Protein Coding TF Transferrin Protein Coding TFRC Transferrin Receptor Protein Coding FTL Ferritin Light Chain Protein Coding CYBB Cytochrome B-245 Beta Chain Protein Coding GSS Glutathione Synthetase Protein Coding CP Ceruloplasmin Protein Coding PRNP Prion Protein Protein Coding SLC11A2 Solute Carrier Family 11 Member 2 Protein Coding SLC40A1 Solute Carrier Family 40 Member 1 Protein Coding STEAP3 STEAP3 Metalloreductase Protein Coding ACSL1 Acyl-CoA Synthetase Long Chain Family Member 1 Protein -

Annotation of Functional Variation Within Non-MHC MS Susceptibility Loci Through Bioinformatics Analysis
Genes and Immunity (2014) 15, 466–476 & 2014 Macmillan Publishers Limited All rights reserved 1466-4879/14 www.nature.com/gene ORIGINAL ARTICLE Annotation of functional variation within non-MHC MS susceptibility loci through bioinformatics analysis FBS Briggs, LJ Leung and LF Barcellos There is a strong and complex genetic component to multiple sclerosis (MS). In addition to variation in the major histocompatibility complex (MHC) region on chromosome 6p21.3, 110 non-MHC susceptibility variants have been identified in Northern Europeans, thus far. The majority of the MS-associated genes are immune related; however, similar to most other complex genetic diseases, the causal variants and biological processes underlying pathogenesis remain largely unknown. We created a comprehensive catalog of putative functional variants that reside within linkage disequilibrium regions of the MS-associated genic variants to guide future studies. Bioinformatics analyses were also conducted using publicly available resources to identify plausible pathological processes relevant to MS and functional hypotheses for established MS-associated variants. Genes and Immunity (2014) 15, 466–476; doi:10.1038/gene.2014.37; published online 17 July 2014 INTRODUCTION protein structure through alternative splicing within IL7R7 and 8 Multiple sclerosis (MS) is a clinically heterogeneous autoimmune TNFRSF1A. However, the causal variants and the pathological disease of the central nervous system with a complex etiology, biological processes mediated by the remaining 103 loci -
A Gene–Environment Interaction Between Smoking and Gene Polymorphisms Provides a High Risk of Two Subgroups of Sarcoidosis Natalia V
www.nature.com/scientificreports OPEN A Gene–Environment Interaction Between Smoking and Gene polymorphisms Provides a High Risk of Two Subgroups of Sarcoidosis Natalia V. Rivera 1,2*, Karina Patasova1, Susanna Kullberg1, Lina Marcela Diaz-Gallo2, Tomoko Iseda1, Camilla Bengtsson3, Lars Alfredsson3, Anders Eklund1, Ingrid Kockum4, Johan Grunewald1,5 & Leonid Padyukov 2,5 The infuence and efect of cigarette smoking in sarcoidosis is unclear. Here, we evaluated gene- environment interaction between multiple genetic variants including HLA genes and smoking in sarcoidosis defned by two clinical phenotypes, Löfgren’s syndrome (LS) and patients without Löfgren’s syndrome (non-LS). To quantify smoking efects in sarcoidosis, we performed a gene-environment interaction study in a Swedish population-based case-control study consisting of 3,713 individuals. Cases and controls were classifed according to their cigarette smoking status and genotypes by Immunochip platform. Gene-smoking interactions were quantifed by an additive interaction model using a logistic regression adjusted by sex, age and frst two principal components. The estimated attributable proportion (AP) was used to quantify the interaction efect. Assessment of smoking efects with inclusion of genetic information revealed 53 (in LS) and 34 (in non-LS) SNP-smoking additive interactions at false discovery rate (FDR) below 5%. The lead signals interacting with smoking were rs12132140 (AP = 0.56, 95% CI = 0.22–0.90), p = 1.28e-03) in FCRL1 for LS and rs61780312 (AP = 0.62, 95% CI = 0.28–0.90), p = 3e-04) in IL23R for non-LS. We further identifed 16 genomic loci (in LS) and 13 (in non-LS) that interact with cigarette smoking. -

Mitochondrial Dynamics in Hematopoietic Stem Cells
Mitochondrial dynamics in hematopoietic stem cells Mariana Justino Lage de Almeida Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2019 © 2019 Mariana Justino Lage de Almeida All rights reserved ABSTRACT Mitochondrial dynamics in hematopoietic stem cells Mariana Justino Lage de Almeida Hematopoietic stem cells (HSCs) take on the extraordinary role of sustaining life-long production of blood cells. Despite their indisputable therapeutic potential, HSC biology is poorly understood, and the field remains limited by the inability to maintain, expand, or generate HSCs in vitro. The aim of this study was to elucidate a particular gap in our understanding of the organellar cell biology of HSCs, specifically the role and function of the mitochondria. Several signaling pathways and biological processes converge onto the mitochondria, yet these organelles were found to be largely dispensable in HSCs on the basis of their predominantly glycolytic metabolism and reports of low mitochondrial content. Our studies show that MitoTracker Green (MTG), a frequently used fluorescent dye to measure mitochondrial mass in hematopoietic populations, is effluxed by HSCs resulting in their systematic and deceptive enrichment in the subset of cells with the lowest MTG fluorescence. Using dye-independent methods we discovered that HSCs have elevated mitochondrial content despite their reliance on glycolysis for ATP production. Moreover, mechanisms of mitochondrial quality control and clearance by autophagy appear to be comparatively lower in HSCs than in any other hematopoietic population we analyzed, suggesting HSCs maintain their mitochondria over time. To investigate the function of mitochondria in HSCs we generated mice with disruption of mitofusins (MFN) 1 and 2. -

1 Imipramine Treatment and Resiliency Exhibit Similar
Imipramine Treatment and Resiliency Exhibit Similar Chromatin Regulation in the Mouse Nucleus Accumbens in Depression Models Wilkinson et al. Supplemental Material 1. Supplemental Methods 2. Supplemental References for Tables 3. Supplemental Tables S1 – S24 SUPPLEMENTAL TABLE S1: Genes Demonstrating Increased Repressive DimethylK9/K27-H3 Methylation in the Social Defeat Model (p<0.001) SUPPLEMENTAL TABLE S2: Genes Demonstrating Decreased Repressive DimethylK9/K27-H3 Methylation in the Social Defeat Model (p<0.001) SUPPLEMENTAL TABLE S3: Genes Demonstrating Increased Repressive DimethylK9/K27-H3 Methylation in the Social Isolation Model (p<0.001) SUPPLEMENTAL TABLE S4: Genes Demonstrating Decreased Repressive DimethylK9/K27-H3 Methylation in the Social Isolation Model (p<0.001) SUPPLEMENTAL TABLE S5: Genes Demonstrating Common Altered Repressive DimethylK9/K27-H3 Methylation in the Social Defeat and Social Isolation Models (p<0.001) SUPPLEMENTAL TABLE S6: Genes Demonstrating Increased Repressive DimethylK9/K27-H3 Methylation in the Social Defeat and Social Isolation Models (p<0.001) SUPPLEMENTAL TABLE S7: Genes Demonstrating Decreased Repressive DimethylK9/K27-H3 Methylation in the Social Defeat and Social Isolation Models (p<0.001) SUPPLEMENTAL TABLE S8: Genes Demonstrating Increased Phospho-CREB Binding in the Social Defeat Model (p<0.001) SUPPLEMENTAL TABLE S9: Genes Demonstrating Decreased Phospho-CREB Binding in the Social Defeat Model (p<0.001) SUPPLEMENTAL TABLE S10: Genes Demonstrating Increased Phospho-CREB Binding in the Social