Matlabbridge Extension
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Management of Large Sets of Image Data Capture, Databases, Image Processing, Storage, Visualization Karol Kozak
Management of large sets of image data Capture, Databases, Image Processing, Storage, Visualization Karol Kozak Download free books at Karol Kozak Management of large sets of image data Capture, Databases, Image Processing, Storage, Visualization Download free eBooks at bookboon.com 2 Management of large sets of image data: Capture, Databases, Image Processing, Storage, Visualization 1st edition © 2014 Karol Kozak & bookboon.com ISBN 978-87-403-0726-9 Download free eBooks at bookboon.com 3 Management of large sets of image data Contents Contents 1 Digital image 6 2 History of digital imaging 10 3 Amount of produced images – is it danger? 18 4 Digital image and privacy 20 5 Digital cameras 27 5.1 Methods of image capture 31 6 Image formats 33 7 Image Metadata – data about data 39 8 Interactive visualization (IV) 44 9 Basic of image processing 49 Download free eBooks at bookboon.com 4 Click on the ad to read more Management of large sets of image data Contents 10 Image Processing software 62 11 Image management and image databases 79 12 Operating system (os) and images 97 13 Graphics processing unit (GPU) 100 14 Storage and archive 101 15 Images in different disciplines 109 15.1 Microscopy 109 360° 15.2 Medical imaging 114 15.3 Astronomical images 117 15.4 Industrial imaging 360° 118 thinking. 16 Selection of best digital images 120 References: thinking. 124 360° thinking . 360° thinking. Discover the truth at www.deloitte.ca/careers Discover the truth at www.deloitte.ca/careers © Deloitte & Touche LLP and affiliated entities. Discover the truth at www.deloitte.ca/careers © Deloitte & Touche LLP and affiliated entities. -

A 3D Interactive Multi-Object Segmentation Tool Using Local Robust Statistics Driven Active Contours
A 3D interactive multi-object segmentation tool using local robust statistics driven active contours The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Gao, Yi, Ron Kikinis, Sylvain Bouix, Martha Shenton, and Allen Tannenbaum. 2012. A 3D Interactive Multi-Object Segmentation Tool Using Local Robust Statistics Driven Active Contours. Medical Image Analysis 16, no. 6: 1216–1227. doi:10.1016/j.media.2012.06.002. Published Version doi:10.1016/j.media.2012.06.002 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:28548930 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA NIH Public Access Author Manuscript Med Image Anal. Author manuscript; available in PMC 2013 August 01. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Med Image Anal. 2012 August ; 16(6): 1216–1227. doi:10.1016/j.media.2012.06.002. A 3D Interactive Multi-object Segmentation Tool using Local Robust Statistics Driven Active Contours Yi Gaoa,*, Ron Kikinisb, Sylvain Bouixa, Martha Shentona, and Allen Tannenbaumc aPsychiatry Neuroimaging Laboratory, Brigham & Women's Hospital, Harvard Medical School, Boston, MA 02115 bSurgical Planning Laboratory, Brigham & Women's Hospital, Harvard Medical School, Boston, MA 02115 cDepartments of Electrical and Computer Engineering and Biomedical Engineering, Boston University, Boston, MA 02115 Abstract Extracting anatomical and functional significant structures renders one of the important tasks for both the theoretical study of the medical image analysis, and the clinical and practical community. -

An Open-Source Research Platform for Image-Guided Therapy
Int J CARS DOI 10.1007/s11548-015-1292-0 ORIGINAL ARTICLE CustusX: an open-source research platform for image-guided therapy Christian Askeland1,3 · Ole Vegard Solberg1 · Janne Beate Lervik Bakeng1 · Ingerid Reinertsen1 · Geir Arne Tangen1 · Erlend Fagertun Hofstad1 · Daniel Høyer Iversen1,2,3 · Cecilie Våpenstad1,2 · Tormod Selbekk1,3 · Thomas Langø1,3 · Toril A. Nagelhus Hernes2,3 · Håkon Olav Leira2,3 · Geirmund Unsgård2,3 · Frank Lindseth1,2,3 Received: 3 July 2015 / Accepted: 31 August 2015 © The Author(s) 2015. This article is published with open access at Springerlink.com Abstract Results The validation experiments show a navigation sys- Purpose CustusX is an image-guided therapy (IGT) research tem accuracy of <1.1mm, a frame rate of 20 fps, and latency platform dedicated to intraoperative navigation and ultra- of 285ms for a typical setup. The current platform is exten- sound imaging. In this paper, we present CustusX as a robust, sible, user-friendly and has a streamlined architecture and accurate, and extensible platform with full access to data and quality process. CustusX has successfully been used for algorithms and show examples of application in technologi- IGT research in neurosurgery, laparoscopic surgery, vascular cal and clinical IGT research. surgery, and bronchoscopy. Methods CustusX has been developed continuously for Conclusions CustusX is now a mature research platform more than 15years based on requirements from clinical for intraoperative navigation and ultrasound imaging and is and technological researchers within the framework of a ready for use by the IGT research community. CustusX is well-defined software quality process. The platform was open-source and freely available at http://www.custusx.org. -

Medical Image Processing Software
Wohlers Report 2018 Medical Image Processing Software Medical image Patient-specific medical devices and anatomical models are almost always produced using radiological imaging data. Medical image processing processing software is used to translate between radiology file formats and various software AM file formats. Theoretically, any volumetric radiological imaging dataset by Andy Christensen could be used to create these devices and models. However, without high- and Nicole Wake quality medical image data, the output from AM can be less than ideal. In this field, the old adage of “garbage in, garbage out” definitely applies. Due to the relative ease of image post-processing, computed tomography (CT) is the usual method for imaging bone structures and contrast- enhanced vasculature. In the dental field and for oral- and maxillofacial surgery, in-office cone-beam computed tomography (CBCT) has become popular. Another popular imaging technique that can be used to create anatomical models is magnetic resonance imaging (MRI). MRI is less useful for bone imaging, but its excellent soft tissue contrast makes it useful for soft tissue structures, solid organs, and cancerous lesions. Computed tomography: CT uses many X-ray projections through a subject to computationally reconstruct a cross-sectional image. As with traditional 2D X-ray imaging, a narrow X-ray beam is directed to pass through the subject and project onto an opposing detector. To create a cross-sectional image, the X-ray source and detector rotate around a stationary subject and acquire images at a number of angles. An image of the cross-section is then computed from these projections in a post-processing step. -

An Open Source Freeware Software for Ultrasound Imaging and Elastography
Proceedings of the eNTERFACE’07 Workshop on Multimodal Interfaces, Istanbul,˙ Turkey, July 16 - August 10, 2007 USIMAGTOOL: AN OPEN SOURCE FREEWARE SOFTWARE FOR ULTRASOUND IMAGING AND ELASTOGRAPHY Ruben´ Cardenes-Almeida´ 1, Antonio Tristan-Vega´ 1, Gonzalo Vegas-Sanchez-Ferrero´ 1, Santiago Aja-Fernandez´ 1, Veronica´ Garc´ıa-Perez´ 1, Emma Munoz-Moreno˜ 1, Rodrigo de Luis-Garc´ıa 1, Javier Gonzalez-Fern´ andez´ 2, Dar´ıo Sosa-Cabrera 2, Karl Krissian 2, Suzanne Kieffer 3 1 LPI, University of Valladolid, Spain 2 CTM, University of Las Palmas de Gran Canaria 3 TELE Laboratory, Universite´ catholique de Louvain, Louvain-la-Neuve, Belgium ABSTRACT • Open source code: to be able for everyone to modify and reuse the source code. UsimagTool will prepare specific software for the physician to change parameters for filtering and visualization in Ultrasound • Efficiency, robust and fast: using a standard object ori- Medical Imaging in general and in Elastography in particular, ented language such as C++. being the first software tool for researchers and physicians to • Modularity and flexibility for developers: in order to chan- compute elastography with integrated algorithms and modular ge or add functionalities as fast as possible. coding capabilities. It will be ready to implement in different • Multi-platform: able to run in many Operating systems ecographic systems. UsimagTool is based on C++, and VTK/ITK to be useful for more people. functions through a hidden layer, which means that participants may import their own functions and/or use the VTK/ITK func- • Usability: provided with an easy to use GUI to interact tions. as easy as possible with the end user. -
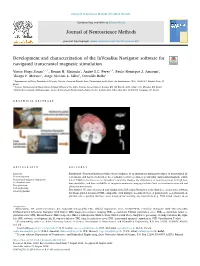
Development and Characterization of the Invesalius Navigator Software for Navigated Transcranial Magnetic Stimulation T ⁎ Victor Hugo Souzaa, ,1, Renan H
Journal of Neuroscience Methods 309 (2018) 109–120 Contents lists available at ScienceDirect Journal of Neuroscience Methods journal homepage: www.elsevier.com/locate/jneumeth Development and characterization of the InVesalius Navigator software for navigated transcranial magnetic stimulation T ⁎ Victor Hugo Souzaa, ,1, Renan H. Matsudaa, André S.C. Peresa,b, Paulo Henrique J. Amorimc, Thiago F. Moraesc, Jorge Vicente L. Silvac, Oswaldo Baffaa a Departamento de Física, Faculdade de Filosofia, Ciências e Letras de Ribeirão Preto, Universidade de São Paulo, Av. Bandeirantes, 3900, 14040-901, Ribeirão Preto, SP, Brazil b Instituto Internacional de Neurociência de Natal Edmond e Lily Safra, Instituto Santos Dumont, Rodovia RN 160 Km 03, 3003, 59280-000, Macaíba, RN, Brazil c Núcleo de Tecnologias Tridimensionais, Centro de Tecnologia da Informação Renato Archer, Rodovia Dom Pedro I Km 143, 13069-901, Campinas, SP, Brazil GRAPHICAL ABSTRACT ARTICLE INFO ABSTRACT Keywords: Background: Neuronavigation provides visual guidance of an instrument during procedures of neurological in- Neuronavigation terventions, and has been shown to be a valuable tool for accurately positioning transcranial magnetic stimu- Transcranial magnetic stimulation lation (TMS) coils relative to an individual’s anatomy. Despite the importance of neuronavigation, its high cost, Localization error low portability, and low availability of magnetic resonance imaging facilities limit its insertion in research and Co-registration clinical environments. Coil positioning New method: We have developed and validated the InVesalius Navigator as the first free, open-source software Surgical planning for image-guided navigated TMS, compatible with multiple tracking devices. A point-based, co-registration al- gorithm and a guiding interface were designed for tracking any instrument (e.g. -

Kitware Source Issue 21
SOFTWARE DEVELOPER’S QUARTERLY Issue 21 • April 2012 Editor’s Note ........................................................................... 1 PARAVIEW 3.14 RELEASED In late February, Kitware and the ParaView team released Recent Releases ..................................................................... 1 ParaView 3.14. This release features usability enhancements, improvements to the Plugin framework, new panels, and ParaView in Immersive Environments .................................. 3 more than 100 other resolved issues. Teaching Open Source .......................................................... 4 ParaView 3.14 features a redesigned Find Data dialog and Color Editor. The updated Find Data dialog makes it possible ParaView Query Selection Framework................................. 7 to use complex queries to select elements, including combin- ing multiple test cases with Boolean operations. The Color Ginkgo CADx: Open Source DICOM CADx Environment .... 8 Editor, used to edit lookup tables or color tables for scalar mapping, now enables independent editing of the color and Code Review, Topic Branches and VTK ................................. 9 opacity functions. Video Analysis on Business Intelligence, ParaView’s charting capabilities have been extended with a Studies in Computational Intelligence ............................... 11 new scatter plot matrix view and the ability to visualize mul- tiple dimensions of data in one compact form. This improved The Visible Patient .............................................................. -

Automated 3D Visualization of Brain Cancer
AUTOMATED 3D VISUALIZATION OF BRAIN CANCER AUTOMATED 3D VISUALIZATION OF BRAIN CANCER By MONA AL-REI, MSc. A Thesis Submitted to the School of Graduate Studies In Partial Fulfillment of the Requirements for the Degree Master of eHealth Program McMaster University @ Copyright by Mona Al-Rei, June 2017 McMaster University Master of eHealth (2017) Hamilton, Ontario TITLE: 3D Brain Cancer Visualization. AUTHOR: Mona Al-Rei. SUPERVISOR: Dr. Thomas Doyle. SUPERVISRORY COMMITTEE: Dr. Reza Samavi, Dr. David Koff. NUMBER OF PAGES: xvii, 119. ii To my beloved and wounded country Yemen iii Abstract Three-dimensional (3D) visualization in cancer control has seen recent progress due to the benefits it offers to the treatment, education, and understanding of the disease. This work identifies the need for an application that directly processes two-dimensional (2D) DICOM images for the segmentation of a brain tumor and the generation of an interactive 3D model suitable for enabling multisensory learning and visualization. A new software application (M-3Ds) was developed to meet these objectives with three modes of segmentation (manual, automatic, and hybrid) for evaluation. M-3Ds software was designed to mitigate the cognitive load and empower health care professionals in their decision making for improved patient outcomes and safety. Comparison of mode accuracy was evaluated. Industrial standard software programs were employed to verify and validate the results of M-3Ds using quantitative volumetric comparison. The study determined that M-3Ds‘ hybrid mode was the highest accuracy with least user intervention for brain tumor segmentation and suitable for the clinical workflow. This paper presents a novel approach to improve medical education, diagnosis, treatment for either surgical planning or radiotherapy of brain cancer. -

Software for Bioprinting Catherine Pakhomova1,2*, Dmitry Popov1, Eugenii Maltsev1, Iskander Akhatov1, Alexander Pasko1,3
REVIEW ARTICLE Software for Bioprinting Catherine Pakhomova1,2*, Dmitry Popov1, Eugenii Maltsev1, Iskander Akhatov1, Alexander Pasko1,3 1Center for Design, Manufacturing and Materials, Skolkovo Institute of Science and Technology, Moscow 2Institute of Engineering Physics for Biomedicine, NRNU Mephi, Moscow 3The National Centre for Computer Animation, Bournemouth University, UK Abstract: The bioprinting of heterogeneous organs is a crucial issue. To reach the complexity of such organs, there is a need for highly specialized software that will meet all requirements such as accuracy, complexity, and others. The primary objective of this review is to consider various software tools that are used in bioprinting and to reveal their capabilities. The sub-objective was to consider different approaches for the model creation using these software tools. Related articles on this topic were analyzed. Software tools are classified based on control tools, general computer-aided design (CAD) tools, tools to convert medical data to CAD formats, and a few highly specialized research-project tools. Different geometry representations are considered, and their advantages and disadvantages are considered applicable to heterogeneous volume modeling and bioprinting. The primary factor for the analysis is suitability of the software for heterogeneous volume modeling and bioprinting or multimaterial three- dimensional printing due to the commonality of these technologies. A shortage of specialized suitable software tools is revealed. There is a need to develop -

Evaluation of Imaging Software Accuracy for 3-Dimensional Analysis of the Mandibular Condyle
International Journal of Environmental Research and Public Health Article Evaluation of Imaging Software Accuracy for 3-Dimensional Analysis of the Mandibular Condyle. A Comparative Study Using a Surface-to-Surface Matching Technique Antonino Lo Giudice 1 , Vincenzo Quinzi 2 , Vincenzo Ronsivalle 1 , Marco Farronato 3 , Carmelo Nicotra 1, Francesco Indelicato 4 and Gaetano Isola 4,* 1 Department of General Surgery and Surgical-Medical Specialties, Section of Orthodontics, School of Dentistry, University of Catania, 95123 Catania, Italy; [email protected] (A.L.G.); [email protected] (V.R.); [email protected] (C.N.) 2 Post Graduate School of Orthodontics, Department of Life, Health and Environmental Sciences, University of L’Aquila, V.le San Salvatore, 67100 L’Aquila, Italy; [email protected] 3 Department of Medicine, Surgery and Dentistry, Section of Orthodontics, University of Milan, 20122 Milan, Italy; [email protected] 4 Department of General Surgery and Surgical-Medical Specialties, Section of Oral Surgery and Periodontology, School of Dentistry, University of Catania, 95123 Catania, Italy; [email protected] * Correspondence: [email protected]; Tel.: +39-095-3782453 Received: 29 May 2020; Accepted: 1 July 2020; Published: 3 July 2020 Abstract: The aim of this study was to assess the accuracy of 3D rendering of the mandibular condylar region obtained from different semi-automatic segmentation methodology. A total of 10 Cone beam computed tomography (CBCT) were selected to perform semi-automatic segmentation of the condyles by using three free-source software (Invesalius, version 3.0.0, Centro de Tecnologia da Informação Renato Archer, Campinas, SP, Brazil; ITK-Snap, version2.2.0; Slicer 3D, version 4.10.2) and one commercially available software Dolphin 3D (Dolphin Imaging, version 11.0, Chatsworth, CA, USA). -

Bioimagesuite Manual 95522 2
v2.6 c Copyright 2008 X. Papademetris, M. Jackowski, N. Rajeevan, R.T. Constable, and L.H Staib. Section of Bioimaging Sciences, Dept. of Diagnostic Radiology, Yale School of Medicine. All Rights Reserved ii Draft July 18, 2008 v2.6 Contents I A. Overview 1 1. Introduction 2 1.1. BioImage Suite Functionality . 3 1.2. BioImage Suite Software Infrastructure . 4 1.3. A Brief History . 4 2. Background 6 2.1. Applications of Medical Imaging Analysis: A Brief Overview . 6 2.2. Medical Image Processing & Analysis . 8 2.3. Software Development Related to Medical Image Analysis . 9 2.4. 3D Graphics and Volume Rendering . 11 3. Starting and Running BioImage Suite 18 3.1. Installation Overview . 18 3.2. Installation Instructions . 19 3.3. The Main BioImage Suite Menu . 22 3.4. Preferences Editor . 23 4. Application Structure 26 4.1. Application Structure Overview . 26 4.2. The File Menu . 27 4.3. The Display Menu . 31 5. Looking At Images 32 5.1. Image Formats . 32 5.2. The Viewers . 33 5.3. The Colormap Editor . 37 5.4. Coordinates for NeuroImaging . 39 5.5. Atlas Tools . 44 6. Advanced Image Visualization 47 6.1. 4D Images . 47 6.2. 3D Rendering Controls . 48 6.3. Volume Rendering . 48 6.4. Oblique Slices . 53 6.5. The Animation Tool . 54 iii Draft July 18, 2008 CONTENTS v2.6 II B. Anatomical Image Analysis 59 7. The Image Processing and Histogram Tools 60 7.1. Introduction . 60 7.2. “Image” and “Results” . 61 7.3. Histogram Control . 62 7.4. -

Slicermorph: an Open and Extensible Platform to Retrieve, Visualize And
bioRxiv preprint doi: https://doi.org/10.1101/2020.11.09.374926; this version posted November 9, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 SlicerMorph: An open and extensible platform to retrieve, visualize 2 and analyze 3D morphology 3 AUTHORS 4 Sara Rolfe 1,2 5 Steve Pieper 3 6 Arthur Porto 2 7 Kelly Diamond 2 8 Julie Winchester 4 9 Shan Shan 5 10 Henry Kirveslahti 6 11 Doug Boyer 4 12 Adam Summers 1 13 A. Murat Maga* 2,7 14 AFFILIATIONS 15 1. University of Washington, Friday Harbor Marine Laboratories, San Juan, WA 16 2. Seattle Children’s Research Institute, Center for Developmental Biology and 17 Regenerative Medicine, Seattle, WA 18 3. Isomics, Inc., Cambridge, MA 19 4. Duke University, Department of Evolutionary Anthropology, Durham, NC 20 5. Mount Holyoke College, Department of Mathematics, South Hadley, MA 21 6. Duke University, Department of Statistical Science, Durham, NC 22 7. University of Washington, Department of Pediatrics, Division of Craniofacial Medicine, 23 Seattle WA 24 *Corresponding author. 25 bioRxiv preprint doi: https://doi.org/10.1101/2020.11.09.374926; this version posted November 9, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license.