Biological Imaging Software Tools Pu to K.W.E
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Management of Large Sets of Image Data Capture, Databases, Image Processing, Storage, Visualization Karol Kozak
Management of large sets of image data Capture, Databases, Image Processing, Storage, Visualization Karol Kozak Download free books at Karol Kozak Management of large sets of image data Capture, Databases, Image Processing, Storage, Visualization Download free eBooks at bookboon.com 2 Management of large sets of image data: Capture, Databases, Image Processing, Storage, Visualization 1st edition © 2014 Karol Kozak & bookboon.com ISBN 978-87-403-0726-9 Download free eBooks at bookboon.com 3 Management of large sets of image data Contents Contents 1 Digital image 6 2 History of digital imaging 10 3 Amount of produced images – is it danger? 18 4 Digital image and privacy 20 5 Digital cameras 27 5.1 Methods of image capture 31 6 Image formats 33 7 Image Metadata – data about data 39 8 Interactive visualization (IV) 44 9 Basic of image processing 49 Download free eBooks at bookboon.com 4 Click on the ad to read more Management of large sets of image data Contents 10 Image Processing software 62 11 Image management and image databases 79 12 Operating system (os) and images 97 13 Graphics processing unit (GPU) 100 14 Storage and archive 101 15 Images in different disciplines 109 15.1 Microscopy 109 360° 15.2 Medical imaging 114 15.3 Astronomical images 117 15.4 Industrial imaging 360° 118 thinking. 16 Selection of best digital images 120 References: thinking. 124 360° thinking . 360° thinking. Discover the truth at www.deloitte.ca/careers Discover the truth at www.deloitte.ca/careers © Deloitte & Touche LLP and affiliated entities. Discover the truth at www.deloitte.ca/careers © Deloitte & Touche LLP and affiliated entities. -

Bioimage Analysis Tools
Bioimage Analysis Tools Kota Miura, Sébastien Tosi, Christoph Möhl, Chong Zhang, Perrine Paul-Gilloteaux, Ulrike Schulze, Simon Norrelykke, Christian Tischer, Thomas Pengo To cite this version: Kota Miura, Sébastien Tosi, Christoph Möhl, Chong Zhang, Perrine Paul-Gilloteaux, et al.. Bioimage Analysis Tools. Kota Miura. Bioimage Data Analysis, Wiley-VCH, 2016, 978-3-527-80092-6. hal- 02910986 HAL Id: hal-02910986 https://hal.archives-ouvertes.fr/hal-02910986 Submitted on 3 Aug 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. 2 Bioimage Analysis Tools 1 2 3 4 5 6 Kota Miura, Sébastien Tosi, Christoph Möhl, Chong Zhang, Perrine Pau/-Gilloteaux, - Ulrike Schulze,7 Simon F. Nerrelykke,8 Christian Tischer,9 and Thomas Penqo'" 1 European Molecular Biology Laboratory, Meyerhofstraße 1, 69117 Heidelberg, Germany National Institute of Basic Biology, Okazaki, 444-8585, Japan 2/nstitute for Research in Biomedicine ORB Barcelona), Advanced Digital Microscopy, Parc Científic de Barcelona, dBaldiri Reixac 1 O, 08028 Barcelona, Spain 3German Center of Neurodegenerative -

Other Departments and Institutes Courses 1
Other Departments and Institutes Courses 1 Other Departments and Institutes Courses About Course Numbers: 02-250 Introduction to Computational Biology Each Carnegie Mellon course number begins with a two-digit prefix that Spring: 12 units designates the department offering the course (i.e., 76-xxx courses are This class provides a general introduction to computational tools for biology. offered by the Department of English). Although each department maintains The course is divided into two halves. The first half covers computational its own course numbering practices, typically, the first digit after the prefix molecular biology and genomics. It examines important sources of biological indicates the class level: xx-1xx courses are freshmen-level, xx-2xx courses data, how they are archived and made available to researchers, and what are sophomore level, etc. Depending on the department, xx-6xx courses computational tools are available to use them effectively in research. may be either undergraduate senior-level or graduate-level, and xx-7xx In the process, it covers basic concepts in statistics, mathematics, and courses and higher are graduate-level. Consult the Schedule of Classes computer science needed to effectively use these resources and understand (https://enr-apps.as.cmu.edu/open/SOC/SOCServlet/) each semester for their results. Specific topics covered include sequence data, searching course offerings and for any necessary pre-requisites or co-requisites. and alignment, structural data, genome sequencing, genome analysis, genetic variation, gene and protein expression, and biological networks and pathways. The second half covers computational cell biology, including biological modeling and image analysis. It includes homework requiring Computational Biology Courses modification of scripts to perform computational analyses. -

A 3D Interactive Multi-Object Segmentation Tool Using Local Robust Statistics Driven Active Contours
A 3D interactive multi-object segmentation tool using local robust statistics driven active contours The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Gao, Yi, Ron Kikinis, Sylvain Bouix, Martha Shenton, and Allen Tannenbaum. 2012. A 3D Interactive Multi-Object Segmentation Tool Using Local Robust Statistics Driven Active Contours. Medical Image Analysis 16, no. 6: 1216–1227. doi:10.1016/j.media.2012.06.002. Published Version doi:10.1016/j.media.2012.06.002 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:28548930 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA NIH Public Access Author Manuscript Med Image Anal. Author manuscript; available in PMC 2013 August 01. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Med Image Anal. 2012 August ; 16(6): 1216–1227. doi:10.1016/j.media.2012.06.002. A 3D Interactive Multi-object Segmentation Tool using Local Robust Statistics Driven Active Contours Yi Gaoa,*, Ron Kikinisb, Sylvain Bouixa, Martha Shentona, and Allen Tannenbaumc aPsychiatry Neuroimaging Laboratory, Brigham & Women's Hospital, Harvard Medical School, Boston, MA 02115 bSurgical Planning Laboratory, Brigham & Women's Hospital, Harvard Medical School, Boston, MA 02115 cDepartments of Electrical and Computer Engineering and Biomedical Engineering, Boston University, Boston, MA 02115 Abstract Extracting anatomical and functional significant structures renders one of the important tasks for both the theoretical study of the medical image analysis, and the clinical and practical community. -
![Downloaded from the Cellprofiler Site [31] to Provide a Starting Point for New Analyses](https://docslib.b-cdn.net/cover/6758/downloaded-from-the-cellprofiler-site-31-to-provide-a-starting-point-for-new-analyses-626758.webp)
Downloaded from the Cellprofiler Site [31] to Provide a Starting Point for New Analyses
Open Access Software2006CarpenteretVolume al. 7, Issue 10, Article R100 CellProfiler: image analysis software for identifying and quantifying comment cell phenotypes Anne E Carpenter*, Thouis R Jones*†, Michael R Lamprecht*, Colin Clarke*†, In Han Kang†, Ola Friman‡, David A Guertin*, Joo Han Chang*, Robert A Lindquist*, Jason Moffat*, Polina Golland† and David M Sabatini*§ reviews Addresses: *Whitehead Institute for Biomedical Research, Cambridge, MA 02142, USA. †Computer Sciences and Artificial Intelligence Laboratory, Massachusetts Institute of Technology, Cambridge, MA 02142, USA. ‡Department of Radiology, Brigham and Women's Hospital, Boston, MA 02115, USA. §Department of Biology, Massachusetts Institute of Technology, Cambridge, MA 02142, USA. Correspondence: David M Sabatini. Email: [email protected] Published: 31 October 2006 Received: 15 September 2006 Accepted: 31 October 2006 reports Genome Biology 2006, 7:R100 (doi:10.1186/gb-2006-7-10-r100) The electronic version of this article is the complete one and can be found online at http://genomebiology.com/2006/7/10/R100 © 2006 Carpenter et al.; licensee BioMed Central Ltd. This is an open access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. deposited research Cell<p>CellProfiler, image analysis the software first free, open-source system for flexible and high-throughput cell image analysis is described.</p> Abstract Biologists can now prepare and image thousands of samples per day using automation, enabling chemical screens and functional genomics (for example, using RNA interference). Here we describe the first free, open-source system designed for flexible, high-throughput cell image analysis, research refereed CellProfiler. -

Cellprofiler: Image Analysis Software for Identifying and Quantifying Cell Phenotypes
CellProfiler: image analysis software for identifying and quantifying cell phenotypes The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Genome Biology. 2006 Oct 31;7(10):R100 As Published http://dx.doi.org/10.1186/gb-2006-7-10-r100 Publisher BioMed Central Ltd Version Final published version Citable link http://hdl.handle.net/1721.1/58762 Terms of Use Creative Commons Attribution Detailed Terms http://creativecommons.org/licenses/by/2.0 1 2 3 Table of Contents Getting Started: MaskImage . 96 Introduction . 6 Morph . 110 Installation . 7 OverlayOutlines . 111 Getting Started with CellProfiler . 9 PlaceAdjacent . 112 RescaleIntensity . 115 Resize . 116 Help: Rotate . 118 BatchProcessing . 11 Smooth . 122 Colormaps. .16 Subtract . 125 DefaultImageFolder . 17 SubtractBackground . 126 DefaultOutputFolder . 18 Tile.............................................127 DeveloperInfo . 19 FastMode . 28 MatlabCrash . 29 Object Processing modules: OutputFilename . 30 ClassifyObjects . 41 PixelSize . 31 ClassifyObjectsByTwoMeasurements . 42 Preferences. .32 ConvertToImage . 45 SkipErrors . 33 Exclude . 65 TechDiagnosis . 34 ExpandOrShrink . 66 FilterByObjectMeasurement . 71 IdentifyObjectsInGrid . 76 File Processing modules: IdentifyPrimAutomatic . 77 CreateBatchFiles . 51 IdentifyPrimManual . 83 ExportToDatabase . 68 IdentifySecondary . 84 ExportToExcel. .70 IdentifyTertiarySubregion. .88 LoadImages . 90 Relate . 113 LoadSingleImage . 94 LoadText. .95 RenameOrRenumberFiles . -

An Open-Source Research Platform for Image-Guided Therapy
Int J CARS DOI 10.1007/s11548-015-1292-0 ORIGINAL ARTICLE CustusX: an open-source research platform for image-guided therapy Christian Askeland1,3 · Ole Vegard Solberg1 · Janne Beate Lervik Bakeng1 · Ingerid Reinertsen1 · Geir Arne Tangen1 · Erlend Fagertun Hofstad1 · Daniel Høyer Iversen1,2,3 · Cecilie Våpenstad1,2 · Tormod Selbekk1,3 · Thomas Langø1,3 · Toril A. Nagelhus Hernes2,3 · Håkon Olav Leira2,3 · Geirmund Unsgård2,3 · Frank Lindseth1,2,3 Received: 3 July 2015 / Accepted: 31 August 2015 © The Author(s) 2015. This article is published with open access at Springerlink.com Abstract Results The validation experiments show a navigation sys- Purpose CustusX is an image-guided therapy (IGT) research tem accuracy of <1.1mm, a frame rate of 20 fps, and latency platform dedicated to intraoperative navigation and ultra- of 285ms for a typical setup. The current platform is exten- sound imaging. In this paper, we present CustusX as a robust, sible, user-friendly and has a streamlined architecture and accurate, and extensible platform with full access to data and quality process. CustusX has successfully been used for algorithms and show examples of application in technologi- IGT research in neurosurgery, laparoscopic surgery, vascular cal and clinical IGT research. surgery, and bronchoscopy. Methods CustusX has been developed continuously for Conclusions CustusX is now a mature research platform more than 15years based on requirements from clinical for intraoperative navigation and ultrasound imaging and is and technological researchers within the framework of a ready for use by the IGT research community. CustusX is well-defined software quality process. The platform was open-source and freely available at http://www.custusx.org. -

Medical Image Processing Software
Wohlers Report 2018 Medical Image Processing Software Medical image Patient-specific medical devices and anatomical models are almost always produced using radiological imaging data. Medical image processing processing software is used to translate between radiology file formats and various software AM file formats. Theoretically, any volumetric radiological imaging dataset by Andy Christensen could be used to create these devices and models. However, without high- and Nicole Wake quality medical image data, the output from AM can be less than ideal. In this field, the old adage of “garbage in, garbage out” definitely applies. Due to the relative ease of image post-processing, computed tomography (CT) is the usual method for imaging bone structures and contrast- enhanced vasculature. In the dental field and for oral- and maxillofacial surgery, in-office cone-beam computed tomography (CBCT) has become popular. Another popular imaging technique that can be used to create anatomical models is magnetic resonance imaging (MRI). MRI is less useful for bone imaging, but its excellent soft tissue contrast makes it useful for soft tissue structures, solid organs, and cancerous lesions. Computed tomography: CT uses many X-ray projections through a subject to computationally reconstruct a cross-sectional image. As with traditional 2D X-ray imaging, a narrow X-ray beam is directed to pass through the subject and project onto an opposing detector. To create a cross-sectional image, the X-ray source and detector rotate around a stationary subject and acquire images at a number of angles. An image of the cross-section is then computed from these projections in a post-processing step. -

Titel Untertitel
KNIME Image Processing Nycomed Chair for Bioinformatics and Information Mining Department of Computer and Information Science Konstanz University, Germany Why Image Processing with KNIME? KNIME UGM 2013 2 The “Zoo” of Image Processing Tools Development Processing UI Handling ImgLib2 ImageJ OMERO OpenCV ImageJ2 BioFormats MatLab Fiji … NumPy CellProfiler VTK Ilastik VIGRA CellCognition … Icy Photoshop … = Single, individual, case specific, incompatible solutions KNIME UGM 2013 3 The “Zoo” of Image Processing Tools Development Processing UI Handling ImgLib2 ImageJ OMERO OpenCV ImageJ2 BioFormats MatLab Fiji … NumPy CellProfiler VTK Ilastik VIGRA CellCognition … Icy Photoshop … → Integration! KNIME UGM 2013 4 KNIME as integration platform KNIME UGM 2013 5 Integration: What and How? KNIME UGM 2013 6 Integration ImgLib2 • Developed at MPI-CBG Dresden • Generic framework for data (image) processing algoritms and data-structures • Generic design of algorithms for n-dimensional images and labelings • http://fiji.sc/wiki/index.php/ImgLib2 → KNIME: used as image representation (within the data cells); basis for algorithms KNIME UGM 2013 7 Integration ImageJ/Fiji • Popular, highly interactive image processing tool • Huge base of available plugins • Fiji: Extension of ImageJ1 with plugin-update mechanism and plugins • http://rsb.info.nih.gov/ij/ & http://fiji.sc/ → KNIME: ImageJ Macro Node KNIME UGM 2013 8 Integration ImageJ2 • Next-generation version of ImageJ • Complete re-design of ImageJ while maintaining backwards compatibility • Based on ImgLib2 -

9. Biomedical Imaging Informatics Daniel L
9. Biomedical Imaging Informatics Daniel L. Rubin, Hayit Greenspan, and James F. Brinkley After reading this chapter, you should know the answers to these questions: 1. What makes images a challenging type of data to be processed by computers, as opposed to non-image clinical data? 2. Why are there many different imaging modalities, and by what major two characteristics do they differ? 3. How are visual and knowledge content in images represented computationally? How are these techniques similar to representation of non-image biomedical data? 4. What sort of applications can be developed to make use of the semantic image content made accessible using the Annotation and Image Markup model? 5. Describe four different types of image processing methods. Why are such methods assembled into a pipeline when creating imaging applications? 6. Give an example of an imaging modality with high spatial resolution. Give an example of a modality that provides functional information. Why are most imaging modalities not capable of providing both? 7. What is the goal in performing segmentation in image analysis? Why is there more than one segmentation method? 8. What are two types of quantitative information in images? What are two types of semantic information in images? How might this information be used in medical applications? 9. What is the difference between image registration and image fusion? Given an example of each. 1 9.1. Introduction Imaging plays a central role in the healthcare process. Imaging is crucial not only to health care, but also to medical communication and education, as well as in research. -

An Open Source Freeware Software for Ultrasound Imaging and Elastography
Proceedings of the eNTERFACE’07 Workshop on Multimodal Interfaces, Istanbul,˙ Turkey, July 16 - August 10, 2007 USIMAGTOOL: AN OPEN SOURCE FREEWARE SOFTWARE FOR ULTRASOUND IMAGING AND ELASTOGRAPHY Ruben´ Cardenes-Almeida´ 1, Antonio Tristan-Vega´ 1, Gonzalo Vegas-Sanchez-Ferrero´ 1, Santiago Aja-Fernandez´ 1, Veronica´ Garc´ıa-Perez´ 1, Emma Munoz-Moreno˜ 1, Rodrigo de Luis-Garc´ıa 1, Javier Gonzalez-Fern´ andez´ 2, Dar´ıo Sosa-Cabrera 2, Karl Krissian 2, Suzanne Kieffer 3 1 LPI, University of Valladolid, Spain 2 CTM, University of Las Palmas de Gran Canaria 3 TELE Laboratory, Universite´ catholique de Louvain, Louvain-la-Neuve, Belgium ABSTRACT • Open source code: to be able for everyone to modify and reuse the source code. UsimagTool will prepare specific software for the physician to change parameters for filtering and visualization in Ultrasound • Efficiency, robust and fast: using a standard object ori- Medical Imaging in general and in Elastography in particular, ented language such as C++. being the first software tool for researchers and physicians to • Modularity and flexibility for developers: in order to chan- compute elastography with integrated algorithms and modular ge or add functionalities as fast as possible. coding capabilities. It will be ready to implement in different • Multi-platform: able to run in many Operating systems ecographic systems. UsimagTool is based on C++, and VTK/ITK to be useful for more people. functions through a hidden layer, which means that participants may import their own functions and/or use the VTK/ITK func- • Usability: provided with an easy to use GUI to interact tions. as easy as possible with the end user. -
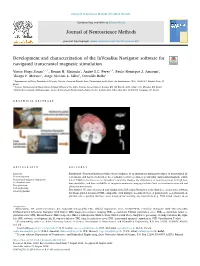
Development and Characterization of the Invesalius Navigator Software for Navigated Transcranial Magnetic Stimulation T ⁎ Victor Hugo Souzaa, ,1, Renan H
Journal of Neuroscience Methods 309 (2018) 109–120 Contents lists available at ScienceDirect Journal of Neuroscience Methods journal homepage: www.elsevier.com/locate/jneumeth Development and characterization of the InVesalius Navigator software for navigated transcranial magnetic stimulation T ⁎ Victor Hugo Souzaa, ,1, Renan H. Matsudaa, André S.C. Peresa,b, Paulo Henrique J. Amorimc, Thiago F. Moraesc, Jorge Vicente L. Silvac, Oswaldo Baffaa a Departamento de Física, Faculdade de Filosofia, Ciências e Letras de Ribeirão Preto, Universidade de São Paulo, Av. Bandeirantes, 3900, 14040-901, Ribeirão Preto, SP, Brazil b Instituto Internacional de Neurociência de Natal Edmond e Lily Safra, Instituto Santos Dumont, Rodovia RN 160 Km 03, 3003, 59280-000, Macaíba, RN, Brazil c Núcleo de Tecnologias Tridimensionais, Centro de Tecnologia da Informação Renato Archer, Rodovia Dom Pedro I Km 143, 13069-901, Campinas, SP, Brazil GRAPHICAL ABSTRACT ARTICLE INFO ABSTRACT Keywords: Background: Neuronavigation provides visual guidance of an instrument during procedures of neurological in- Neuronavigation terventions, and has been shown to be a valuable tool for accurately positioning transcranial magnetic stimu- Transcranial magnetic stimulation lation (TMS) coils relative to an individual’s anatomy. Despite the importance of neuronavigation, its high cost, Localization error low portability, and low availability of magnetic resonance imaging facilities limit its insertion in research and Co-registration clinical environments. Coil positioning New method: We have developed and validated the InVesalius Navigator as the first free, open-source software Surgical planning for image-guided navigated TMS, compatible with multiple tracking devices. A point-based, co-registration al- gorithm and a guiding interface were designed for tracking any instrument (e.g.