2019 Strain Engineering for Mi
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Glycerol Dehydrogenase from Gluconobacter Industrius
Agric. Biol Chem., 49 (4), 1001 -1010, 1985 1001 Solubilization, Purification and Properties of Membrane-bound Glycerol Dehydrogenase from Gluconobacter industrius Minoru Ameyama,Emiko Shinagawa, Kazunobu Matsushita and Osao Adachi Laboratory of Applied Microbiology, Department of Agricultural Chemistry, Faculty of Agriculture, Yamaguchi University, Yamaguchi 753, Japan Received July 30, 1984 Membrane-bound glycerol dehydrogenase was solubilized and purified about 100-fold from the membraneof Gluconobacter industrius IFO 3260 grown on a glycerol-glutamate medium. Solubilization of the enzyme was successfully achieved by use of 0.5% dimethyldodecylamineoxide in 0.05 m Tris-HCl, pH 8.0. Alcohol dehydrogenase and D-glucose dehydrogenase, which were abundantly formed in the same bacterial membrane, were eliminated on solubilization. Glycerol dehydrogenase was further purified through fractionation with polyethylene glycol 6000. The enzymeshowed a broad substrate specificity and various kinds of polyhydroxyl alcohols, in addition to glycerol, were rapidly oxidized in the presence of 2,6-dichlorophenolindophenoi and phenazine methosulfate as the electron acceptor but NADand NADPwere inert. The enzyme was proved to be a quinoprotein in which pyrroloquinoline quinone functioned as the prosthetic group. The first report on microbial oxidation of localization of the oxidase system in cells of G. glycerol to dihydroxyacetone was by Bertrand liquefaciens and found that the oxidation of with a strain capable of L-sorbose fermen- glycerol and raeso-erythritol -

<I>Lactobacillus Reuteri</I>
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Faculty Publications in Food Science and Food Science and Technology Department Technology 2014 From prediction to function using evolutionary genomics: Human-specific ecotypes of Lactobacillus reuteri have diverse probiotic functions Jennifer K. Spinler Texas Children’s Hospital, [email protected] Amrita Sontakke Baylor College of Medicine Emily B. Hollister Baylor College of Medicine Susan F. Venable Baylor College of Medicine Phaik Lyn Oh University of Nebraska, Lincoln See next page for additional authors Follow this and additional works at: http://digitalcommons.unl.edu/foodsciefacpub Spinler, Jennifer K.; Sontakke, Amrita; Hollister, Emily B.; Venable, Susan F.; Oh, Phaik Lyn; Balderas, Miriam A.; Saulnier, Delphine M.A.; Mistretta, Toni-Ann; Devaraj, Sridevi; Walter, Jens; Versalovic, James; and Highlander, Sarah K., "From prediction to function using evolutionary genomics: Human-specific ce otypes of Lactobacillus reuteri have diverse probiotic functions" (2014). Faculty Publications in Food Science and Technology. 132. http://digitalcommons.unl.edu/foodsciefacpub/132 This Article is brought to you for free and open access by the Food Science and Technology Department at DigitalCommons@University of Nebraska - Lincoln. It has been accepted for inclusion in Faculty Publications in Food Science and Technology by an authorized administrator of DigitalCommons@University of Nebraska - Lincoln. Authors Jennifer K. Spinler, Amrita Sontakke, Emily B. Hollister, -

Medium and Long-Term Opportunities and Risks of the Biotechnological Production of Bulk Chemicals from Renewable Resources
Medium and Long-term Opportunities and Risks of the Biotechnological Production of Bulk Chemicals from Renewable Resources - The Potential of White Biotechnology The BREW Project Final report Prepared under the European Commission’s GROWTH Programme (DG Research) Project team: Academy Utrecht University (UU), Dept. of Science, Technology and Society (STS), Utrecht, Netherlands Fraunhofer Institute for Systems and Innovation Research (FhG-ISI), Karlsruhe, Germany Universidad Complutense de Madrid (UCM), Dept. of Chemical Engineering, Madrid, Spain Plant Research International (PRI), Wageningen, Netherlands CERISS (Centro per l'Educazione, la Ricerca, l'Informazione su Scienza e Società), Milan, Italy A&F (Agrotechnology and Food Innovations) Wageningen, Netherlands Industry partners BP Chemicals, Hull, United Kingdom Degussa AG, Hanau, Germany DSM NV, Heerlen, Netherlands DuPont, Bad Homburg, Germany NatureWorks, Naarden, Netherlands Novozymes A/S, Bagsvaerd, Denmark Roquette Frères, Lestrem, France Shell International Chemicals BV, Amsterdam, Netherlands Uniqema, Wilton/Redcar, United Kingdom Utrecht, September 2006 Authors Dr. Martin Patel (project co-ordinator) Utrecht University (UU) Manuela Crank, BE Chem Department of Science, Technology Dr. Veronika Dornburg and Society (STS) Barbara Hermann, M.Sc. Heidelberglaan 2 Lex Roes, M.Sc. NL-3584 CS Utrecht, Netherlands Tel. +31 (0) 30 253-7600 Fax +31 (0) 30 253-7601 [email protected] Dr. Bärbel Hüsing Fraunhofer Institute for Systems and Innovation Research (FhG-ISI), Karlsruhe, Germany Dr. Leo Overbeek Plant Research International (PRI) Wageningen, Netherlands Dr. Fabio Terragni CERISS (Centro per l'Educazione, la Dr. Elena Recchia Ricerca, l'Informazione su Scienza e Società), Milan, Italy Contributors Academy Dr. Ruud Weusthuis A&F (Agrotechnology and Food Innovations) Wageningen, Netherlands Prof. -

Evolution of Coenzyme BI2 Synthesis Among Enteric Bacteria
Copyright 0 1996 by the Genetics Society of America Evolution of Coenzyme BI2Synthesis Among Enteric Bacteria: Evidence for Loss and Reacquisition of a Multigene Complex Jeffrey G. Lawrence and John R. Roth Department of Biology, University of Utah, Salt Lake City, Utah 84112 Manuscript received June 16, 1995 Accepted for publication October 4, 1995 ABSTRACT We have examined the distribution of cobalamin (coenzyme BI2) synthetic ability and cobalamin- dependent metabolism among entericbacteria. Most species of enteric bacteria tested synthesize cobala- min under both aerobic and anaerobic conditions and ferment glycerol in a cobalamindependent fashion. The group of species including Escha'chia coli and Salmonella typhimurium cannot ferment glyc- erol. E. coli strains cannot synthesize cobalamin de novo, and Salmonella spp. synthesize cobalamin only under anaerobic conditions. In addition, the cobalamin synthetic genes of Salmonella spp. (cob) show a regulatory pattern different from that of other enteric taxa tested. We propose that the cobalamin synthetic genes, as well asgenes providing cobalamindependent diol dehydratase, were lostby a common ancestor of E. coli and Salmonella spp. and were reintroduced as a single fragment into the Salmonella lineage from an exogenous source. Consistent with this hypothesis, the S. typhimurium cob genes do not hybridize with the genomes of other enteric species. The Salmonella cob operon may represent a class of genes characterized by periodic loss and reacquisition by host genomes. This process may be an important aspect of bacterial population genetics and evolution. OBALAMIN (coenzyme BIZ) is a large evolution- The cobalamin biosynthetic genes have been charac- C arily ancient molecule ( GEORGOPAPADAKOUand terized in S. -

Supplementary Information
Supplementary information (a) (b) Figure S1. Resistant (a) and sensitive (b) gene scores plotted against subsystems involved in cell regulation. The small circles represent the individual hits and the large circles represent the mean of each subsystem. Each individual score signifies the mean of 12 trials – three biological and four technical. The p-value was calculated as a two-tailed t-test and significance was determined using the Benjamini-Hochberg procedure; false discovery rate was selected to be 0.1. Plots constructed using Pathway Tools, Omics Dashboard. Figure S2. Connectivity map displaying the predicted functional associations between the silver-resistant gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S3. Connectivity map displaying the predicted functional associations between the silver-sensitive gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S4. Metabolic overview of the pathways in Escherichia coli. The pathways involved in silver-resistance are coloured according to respective normalized score. Each individual score represents the mean of 12 trials – three biological and four technical. Amino acid – upward pointing triangle, carbohydrate – square, proteins – diamond, purines – vertical ellipse, cofactor – downward pointing triangle, tRNA – tee, and other – circle. -

Electron Transport Phosphorylation in Rumen Butyrivibrios: Unprecedented ATP Yield for Glucose Fermentation to Butyrate
HYPOTHESIS AND THEORY published: 24 June 2015 doi: 10.3389/fmicb.2015.00622 Electron transport phosphorylation in rumen butyrivibrios: unprecedented ATP yield for glucose fermentation to butyrate Timothy J. Hackmann1 and Jeffrey L. Firkins2* 1 Department of Animal Sciences, University of Florida, Gainesville, FL, USA, 2 Department of Animal Sciences, The Ohio State University, Columbus, OH, USA From a genomic analysis of rumen butyrivibrios (Butyrivibrio and Pseudobutyrivibrio sp.), we have re-evaluated the contribution of electron transport phosphorylation (ETP) to ATP formation in this group. This group is unique in that most (76%) genomes were predicted to possess genes for both Ech and Rnf transmembrane ion pumps. These pumps act in concert with the NifJ and Bcd-Etf to form a electrochemical potential (μH+ and μNa+), which drives ATP synthesis by ETP. Of the 62 total butyrivibrio genomes currently available from the Hungate 1000 project, all 62 were predicted to Edited by: possess NifJ, which reduces oxidized ferredoxin (Fdox) during pyruvate conversion to Emilio M. Ungerfeld, Instituto de Investigaciones acetyl-CoA. All 62 possessed all subunits of Bcd-Etf, which reduces Fdox and oxidizes Agropecuarias, Chile reduced NAD during crotonyl-CoA reduction. Additionally, 61 genomes possessed all Reviewed by: subunits of the Rnf, which generates μH+ or μNa+ from oxidation of reduced Fd Wolfgang Buckel, (Fdred) and reduction of oxidized NAD. Further, 47 genomes possessed all six subunits Philipps-Universität Marburg, + Germany of the Ech, which generates μH from oxidation of Fdred. For glucose fermentation Robert J. Wallace, to butyrate and H2, the electrochemical potential established should drive synthesis University of Aberdeen, UK of ∼1.5 ATP by the F0F1-ATP synthase (possessed by all 62 genomes). -
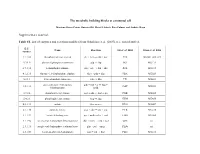
The Metabolic Building Blocks of a Minimal Cell Supplementary
The metabolic building blocks of a minimal cell Mariana Reyes-Prieto, Rosario Gil, Mercè Llabrés, Pere Palmer and Andrés Moya Supplementary material. Table S1. List of enzymes and reactions modified from Gabaldon et. al. (2007). n.i.: non identified. E.C. Name Reaction Gil et. al. 2004 Glass et. al. 2006 number 2.7.1.69 phosphotransferase system glc + pep → g6p + pyr PTS MG041, 069, 429 5.3.1.9 glucose-6-phosphate isomerase g6p ↔ f6p PGI MG111 2.7.1.11 6-phosphofructokinase f6p + atp → fbp + adp PFK MG215 4.1.2.13 fructose-1,6-bisphosphate aldolase fbp ↔ gdp + dhp FBA MG023 5.3.1.1 triose-phosphate isomerase gdp ↔ dhp TPI MG431 glyceraldehyde-3-phosphate gdp + nad + p ↔ bpg + 1.2.1.12 GAP MG301 dehydrogenase nadh 2.7.2.3 phosphoglycerate kinase bpg + adp ↔ 3pg + atp PGK MG300 5.4.2.1 phosphoglycerate mutase 3pg ↔ 2pg GPM MG430 4.2.1.11 enolase 2pg ↔ pep ENO MG407 2.7.1.40 pyruvate kinase pep + adp → pyr + atp PYK MG216 1.1.1.27 lactate dehydrogenase pyr + nadh ↔ lac + nad LDH MG460 1.1.1.94 sn-glycerol-3-phosphate dehydrogenase dhp + nadh → g3p + nad GPS n.i. 2.3.1.15 sn-glycerol-3-phosphate acyltransferase g3p + pal → mag PLSb n.i. 2.3.1.51 1-acyl-sn-glycerol-3-phosphate mag + pal → dag PLSc MG212 acyltransferase 2.7.7.41 phosphatidate cytidyltransferase dag + ctp → cdp-dag + pp CDS MG437 cdp-dag + ser → pser + 2.7.8.8 phosphatidylserine synthase PSS n.i. cmp 4.1.1.65 phosphatidylserine decarboxylase pser → peta PSD n.i. -

High-Yield Anaerobic Succinate Production by Strategically
Meng et al. Microb Cell Fact (2016) 15:141 DOI 10.1186/s12934-016-0536-1 Microbial Cell Factories RESEARCH Open Access High‑yield anaerobic succinate production by strategically regulating multiple metabolic pathways based on stoichiometric maximum in Escherichia coli Jiao Meng1,2,3†, Baiyun Wang1,2,3†, Dingyu Liu1,2,3, Tao Chen1,2,3,4, Zhiwen Wang1,2,3* and Xueming Zhao1,2,3 Abstract Background: Succinate has been identified by the U.S. Department of Energy as one of the top 12 building block chemicals, which can be used as a specialty chemical in the agricultural, food, and pharmaceutical industries. Escheri- chia coli are now one of the most important succinate producing candidates. However, the stoichiometric maximum succinate yield under anaerobic conditions through the reductive branch of the TCA cycle is restricted by NADH sup- ply in E. coli. Results: In the present work, we report a rational approach to increase succinate yield by regulating NADH supply via pentose phosphate (PP) pathway and enhancing flux towards succinate. The deregulated genes zwf243 (encod- ing glucose-6-phosphate dehydrogenase) and gnd361 (encoding 6-phosphogluconate dehydrogenase) involved in NADPH generation from Corynebacterium glutamicum were firstly introduced into E. coli for succinate production. Co-expression of beneficial mutated dehydrogenases, which removed feedback inhibition in the oxidative part of the PP pathway, increased succinate yield from 1.01 to 1.16 mol/mol glucose. Three critical genes, pgl (encoding 6-phos- phogluconolactonase), tktA (encoding transketolase) and talB (encoding transaldolase) were then overexpressed to redirect more carbon flux towards PP pathway and further improved succinate yield to 1.21 mol/mol glucose. -

The Diversity of Microbial Aldo/Keto Reductases from Escherichia Coli K12
Lapthorn, A. J., Zhu, X., and Ellis, E. M. (2013) The diversity of microbial aldo/keto reductases from Escherichia coli K12. Chemico-Biological Interactions, 202(1-3), pp. 168-177. (doi:10.1016/j.cbi.2012.10.008) There may be differences between this version and the published version. You are advised to consult the publisher’s version if you wish to cite from it. http://eprints.gla.ac.uk/124204/ Deposited on: 07 October 2016 Enlighten – Research publications by members of the University of Glasgow http://eprints.gla.ac.uk The diversity of microbial aldo/keto reductases from Escherichia coli K12 Adrian J. Lapthorn 1, Xiaofeng Zhu 2,3 and Elizabeth M. Ellis 3 1 School of Chemistry, Joseph Black Building, University of Glasgow, Glasgow G12 8QQ 2 College of Life Science and State Key Laboratory of Biotherapy and Cancer Centre, Sichuan University, Chengdu, China 3 Strathclyde Institute of Pharmacy and Biomedical Sciences, 161 Cathedral Street, Glasgow, G4 0RE Corresponding author: Adrian J. Lapthorn School of Chemistry, Joseph Black Building, University of Glasgow, Glasgow G12 8QQ Tel: +44 141-330 5940 Fax: +44 141-330 4888 E-mail: [email protected] Key Words: Aldo-Keto reductases, methylglyoxal reductase, 2,5-diketo-D-gluconate reductase, tyrosine auxotrophy suppressor protein, L-glyceraldehyde 3-phosphate reductase, AKR quaternary structure. Abstract The genome of Escherichia coli K12 contains 9 open reading frames encoding aldo/keto reductases (AKR) that are differentially regulated and sequence diverse. A significant amount of data is available for the E. coli AKRs through the availability of gene knockouts and gene expression studies, which adds to the biochemical and kinetic data. -
Q 297 Suppl USE
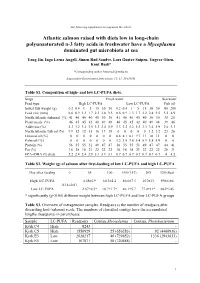
The following supplement accompanies the article Atlantic salmon raised with diets low in long-chain polyunsaturated n-3 fatty acids in freshwater have a Mycoplasma dominated gut microbiota at sea Yang Jin, Inga Leena Angell, Simen Rød Sandve, Lars Gustav Snipen, Yngvar Olsen, Knut Rudi* *Corresponding author: [email protected] Aquaculture Environment Interactions 11: 31–39 (2019) Table S1. Composition of high- and low LC-PUFA diets. Stage Fresh water Sea water Feed type High LC-PUFA Low LC-PUFA Fish oil Initial fish weight (g) 0.2 0.4 1 5 15 30 50 0.2 0.4 1 5 15 30 50 80 200 Feed size (mm) 0.6 0.9 1.3 1.7 2.2 2.8 3.5 0.6 0.9 1.3 1.7 2.2 2.8 3.5 3.5 4.9 North Atlantic fishmeal (%) 41 40 40 40 40 30 30 41 40 40 40 40 30 30 35 25 Plant meals (%) 46 45 45 42 40 49 48 46 45 45 42 40 49 48 39 46 Additives (%) 3.3 3.2 3.2 3.5 3.3 3.4 3.9 3.3 3.2 3.2 3.5 3.3 3.4 3.9 2.6 3.3 North Atlantic fish oil (%) 9.9 12 12 15 16 17 18 0 0 0 0 0 1.2 1.2 23 26 Linseed oil (%) 0 0 0 0 0 0 0 6.8 8.1 8.1 9.7 11 10 11 0 0 Palm oil (%) 0 0 0 0 0 0 0 3.2 3.8 3.8 5.4 5.9 5.8 5.9 0 0 Protein (%) 56 55 55 51 49 47 47 56 55 55 51 49 47 47 44 41 Fat (%) 16 18 18 21 22 22 22 16 18 18 21 22 22 22 28 31 EPA+DHA (% diet) 2.2 2.4 2.4 2.9 3.1 3.1 3.1 0.7 0.7 0.7 0.7 0.7 0.7 0.7 4 4.2 Table S2. -

Supplementary Informations SI2. Supplementary Table 1
Supplementary Informations SI2. Supplementary Table 1. M9, soil, and rhizosphere media composition. LB in Compound Name Exchange Reaction LB in soil LBin M9 rhizosphere H2O EX_cpd00001_e0 -15 -15 -10 O2 EX_cpd00007_e0 -15 -15 -10 Phosphate EX_cpd00009_e0 -15 -15 -10 CO2 EX_cpd00011_e0 -15 -15 0 Ammonia EX_cpd00013_e0 -7.5 -7.5 -10 L-glutamate EX_cpd00023_e0 0 -0.0283302 0 D-glucose EX_cpd00027_e0 -0.61972444 -0.04098397 0 Mn2 EX_cpd00030_e0 -15 -15 -10 Glycine EX_cpd00033_e0 -0.0068175 -0.00693094 0 Zn2 EX_cpd00034_e0 -15 -15 -10 L-alanine EX_cpd00035_e0 -0.02780553 -0.00823049 0 Succinate EX_cpd00036_e0 -0.0056245 -0.12240603 0 L-lysine EX_cpd00039_e0 0 -10 0 L-aspartate EX_cpd00041_e0 0 -0.03205557 0 Sulfate EX_cpd00048_e0 -15 -15 -10 L-arginine EX_cpd00051_e0 -0.0068175 -0.00948672 0 L-serine EX_cpd00054_e0 0 -0.01004986 0 Cu2+ EX_cpd00058_e0 -15 -15 -10 Ca2+ EX_cpd00063_e0 -15 -100 -10 L-ornithine EX_cpd00064_e0 -0.0068175 -0.00831712 0 H+ EX_cpd00067_e0 -15 -15 -10 L-tyrosine EX_cpd00069_e0 -0.0068175 -0.00233919 0 Sucrose EX_cpd00076_e0 0 -0.02049199 0 L-cysteine EX_cpd00084_e0 -0.0068175 0 0 Cl- EX_cpd00099_e0 -15 -15 -10 Glycerol EX_cpd00100_e0 0 0 -10 Biotin EX_cpd00104_e0 -15 -15 0 D-ribose EX_cpd00105_e0 -0.01862144 0 0 L-leucine EX_cpd00107_e0 -0.03596182 -0.00303228 0 D-galactose EX_cpd00108_e0 -0.25290619 -0.18317325 0 L-histidine EX_cpd00119_e0 -0.0068175 -0.00506825 0 L-proline EX_cpd00129_e0 -0.01102953 0 0 L-malate EX_cpd00130_e0 -0.03649016 -0.79413596 0 D-mannose EX_cpd00138_e0 -0.2540567 -0.05436649 0 Co2 EX_cpd00149_e0 -

Engineering of Glycerol Utilization in Gluconobacter Oxydans 621H For
Yan et al. Microb Cell Fact (2018) 17:158 https://doi.org/10.1186/s12934-018-1001-0 Microbial Cell Factories RESEARCH Open Access Engineering of glycerol utilization in Gluconobacter oxydans 621H for biocatalyst preparation in a low‑cost way Jinxin Yan1, Jing Xu1,3, Menghao Cao1, Zhong Li1, Chengpeng Xu1, Xinyu Wang1, Chunyu Yang1, Ping Xu2, Chao Gao1 and Cuiqing Ma1* Abstract Background: Whole cells of Gluconobacter oxydans are widely used in various biocatalytic processes. Sorbitol at high concentrations is commonly used in complex media to prepare biocatalysts. Exploiting an alternative process for preparation of biocatalysts with low cost substrates is of importance for industrial applications. Results: G. oxydans 621H was confrmed to have the ability to grow in mineral salts medium with glycerol, an inevitable waste generated from industry of biofuels, as the sole carbon source. Based on the glycerol utilization mechanism elucidated in this study, the major polyol dehydrogenase (GOX0854) and the membrane-bound alcohol dehydrogenase (GOX1068) can competitively utilize glycerol but play no obvious roles in the biocatalyst prepara- tion. Thus, the genes related to these two enzymes were deleted. Whole cells of G. oxydans ∆GOX1068∆GOX0854 can be prepared from glycerol with a 2.4-fold higher biomass yield than that of G. oxydans 621H. Using whole cells of G. 1 1 oxydans ∆GOX1068∆GOX0854 as the biocatalyst, 61.6 g L− xylonate was produced from 58.4 g L− xylose at a yield of 1 1.05 g g− . Conclusion: This process is an example of efcient preparation of whole cells of G. oxydans with reduced cost.