Genomic Characterization of Human Adenovirus Type 4 Strains Isolated
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Human Illness Caused by E. Coli O157:H7 from Food and Non-Food Sources
FRI BRIEFINGS Human Illness Caused by E. coli O157:H7 from Food and Non-food Sources M. Ellin Doyle1*, John Archer2, Charles W. Kaspar1, and Ronald Weiss1 1Food Research Institute, University of Wisconsin–Madison, Madison, WI 53706 2Wisconsin Division of Public Health, Bureau of Communicable Diseases and Preparedness, Communicable Disease Epidemiology Section, Madison, WI 53702 Contents Introduction ...................................................................................................................................2 Epidemiology of E. coli O157:H7..................................................................................................2 Outbreak Data ........................................................................................................................2 Reservoirs of E. coli O157:H7 ..............................................................................................3 Cattle—the primary reservoir ........................................................................................3 Other ruminants .............................................................................................................4 Other animals .................................................................................................................4 Transport Hosts......................................................................................................................4 Routes of Human Infection ....................................................................................................5 -

How Do Pathogenic Microorganisms Develop Cross-Kingdom Host Jumps? Peter Van Baarlen1, Alex Van Belkum2, Richard C
Molecular mechanisms of pathogenicity: how do pathogenic microorganisms develop cross-kingdom host jumps? Peter van Baarlen1, Alex van Belkum2, Richard C. Summerbell3, Pedro W. Crous3 & Bart P.H.J. Thomma1 1Laboratory of Phytopathology, Wageningen University, Wageningen, The Netherlands; 2Department of Medical Microbiology and Infectious Diseases, Erasmus MC, University Medical Centre Rotterdam, Rotterdam, The Netherlands; and 3CBS Fungal Biodiversity Centre, Utrecht, The Netherlands Correspondence: Bart P.H.J. Thomma, Abstract Downloaded from https://academic.oup.com/femsre/article/31/3/239/2367343 by guest on 27 September 2021 Laboratory of Phytopathology, Wageningen University, Binnenhaven 5, 6709 PD It is common knowledge that pathogenic viruses can change hosts, with avian Wageningen, The Netherlands. Tel.: 10031 influenza, the HIV, and the causal agent of variant Creutzfeldt–Jacob encephalitis 317 484536; fax: 10031 317 483412; as well-known examples. Less well known, however, is that host jumps also occur e-mail: [email protected] with more complex pathogenic microorganisms such as bacteria and fungi. In extreme cases, these host jumps even cross kingdom of life barriers. A number of Received 3 July 2006; revised 22 December requirements need to be met to enable a microorganism to cross such kingdom 2006; accepted 23 December 2006. barriers. Potential cross-kingdom pathogenic microorganisms must be able to First published online 26 February 2007. come into close and frequent contact with potential hosts, and must be able to overcome or evade host defences. Reproduction on, in, or near the new host will DOI:10.1111/j.1574-6976.2007.00065.x ensure the transmission or release of successful genotypes. -

Differences in Innate Immune Response Between Man and Mouse
Institute of Medical Physics and Biophysics Medical Department, Leipzig University Author Manuscript © 2014. This manuscript version is made available under the CC-BY-NC-ND 4.0 license http://creativecommons.org/licenses/by-nc-nd/4.0/ Published in final edited form as: Critical Reviews™ in Immunology, 2014; 34:433-54. Available at: http://dx.doi.org/ 10.1615/CritRevImmunol.2014011600. DIFFERENCES IN INNATE IMMUNE RESPONSE BETWEEN MAN AND MOUSE Josefin Zschaler 1,2,*, Denise Schlorke 1,2,* & Juergen Arnhold 1,2 1 Institute for Medical Physics and Biophysics, Medical Faculty, Leipzig University, Leipzig, Germany 2 Translational Centre for Regenerative Medicine (TRM), Leipzig University, Leipzig, Germany * Both authors contributed equally to this work. Abstract Mouse strains are frequently used to model human disease states, to test the efficiency of drugs and therapeutic principles. However, the direct translation of murine experimental data to human pathological events often fails due to sufficient differences in the organization of the immune system of both species. Here we give a short overview of the principle differences between mice and humans in defense strategies against pathogens and mechanisms involved in response to pathogenic microorganisms and other activators of the immune system. While in human blood mechanisms of immune resistance are highly prevailed, tolerance mechanisms dominate for the defense against pathogenic microorganisms in mouse blood. Further on, species-related differences of immune cells mainly involved in innate immune response as well as differences to maintain oxidative homeostasis are also considered. A number of disease scenarios in mice are critically reflected for their suitability to serve as a model for human pathologies. -

The Public Health Impact of Prion Diseases1
10 Feb 2005 13:10 AR AR238-PU26-08.tex AR238-PU26-08.sgm LaTeX2e(2002/01/18) P1: IBD 10.1146/annurev.publhealth.26.021304.144536 Annu. Rev. Public Health 2005. 26:191–212 doi: 10.1146/annurev.publhealth.26.021304.144536 Copyright ©c 2005 by Annual Reviews. All rights reserved First published online as a Review in Advance on December 8, 2004 THE PUBLIC HEALTH IMPACT OF PRION DISEASES1 Ermias D. Belay and Lawrence B. Schonberger Division of Viral and Rickettsial Diseases, National Center for Infectious Diseases, Centers for Disease Control and Prevention, Atlanta, Georgia 30333; email: [email protected] KeyWords transmissible spongiform encephalopathy, Creutzfeldt-Jakob disease, variant Creutzfeldt-Jakob disease, bovine spongiform encephalopathy, chronic wasting disease ■ Abstract Several prion disease–related human health risks from an exogenous source can be identified in the United States, including the iatrogenic transmission of Creutzfeldt-Jakob disease (CJD), the possible occurrence of variant CJD (vCJD), and potential zoonotic transmission of chronic wasting disease (CWD). Although cross- species transmission of prion diseases seems to be limited by an apparent “species barrier,” the occurrence of bovine spongiform encephalopathy (BSE) and its transmis- sion to humans indicate that animal prion diseases can pose a significant public health risk. Recent reports of secondary person-to-person spread of vCJD via blood products and detection of vCJD transmission in a patient heterozygous at codon 129 further illustrate the potential public health impacts of BSE. INTRODUCTION by IRMO/Information Center on 03/14/05. For personal use only. Prion diseases, also known as transmissible spongiform encephalopathies (TSEs), are a group of animal and human brain diseases that are uniformly fatal and often characterized by a long incubation period and a multifocal neuropathologic picture of neuronal loss, spongiform changes, and astrogliosis (3). -

Innate Immunity Part I: the Immediate Response Deborah A
Innate Immunity Part I: The Immediate Response Deborah A. Lebman, Ph.D. OBJECTIVES 1. Describe three mechanisms present at epithelial surfaces that protect the host from pathogens. 2. List 3 functions of complement proteins in innate immunity 3. Know the primary functions of macrophages and neutrophils 4. List the receptors on macrophages involved in recognition of pathogens 5. List the key toxic products of phagocytes READING • Parham, The Immune System, pg 227-238. TERMS • Defensin: antibacterial peptide produced by the host at epithelial surfaces • Complement: ubiquitously present proteins synthesized primarily in the liver that mediate a variety of responses to pathogens • Opsonin (opsonization): alteration of cell surface that facilitates phagocytosis • Phagocytosis: internalization of particulate matter by cells • CD14: LPS receptor • TLR-4: toll like receptor 4 I. Introduction A. The first response to pathogens occurs immediately B. The early response employs both mechanical and cell mediated mechanisms C. There is no cellular expansion, but there is cell recruitment D. A wide array of soluble mediators with both positive and negative effects, i.e. remove the pathogen but hurt the host, is released II. Overview of Pathogens A. There are four types of pathogen: bacteria, virus, fungus, parasite B. Table I: Examples of the Four Types of Common Human Pathogen Table 1: Examples of the Four Types of Common Human Pathogen Table 1 (Continued): Examples of the Four Types of Common Human Pathogen C. Pathogens can be intracellular or extracellular Table 2: Examples of intracellular and extracellular pathogens. Notice that both innate and adaptive mechanisms are involved in the response to both types of pathogen. -
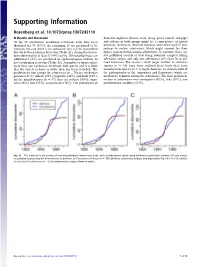
Supporting Information
Supporting Information Rosenberg et al. 10.1073/pnas.1307243110 SI Results and Discussion domestic ungulates (horses, cows, sheep, goats, camels, and pigs) Of the 83 arboviruses, nonhuman vertebrate hosts have been and rodents in both groups might be a consequence of spatial identified for 70 (84%); the remaining 13 are presumed to be proximity to humans. Sentinel monkeys were often used in pro- zoonoses because there is no indication they can be transmitted cedures to isolate arboviruses, which might account for their directly between humans by vectors (Table S1). Animal hosts have higher representation among arboviruses. In contrast, there are been identified for at least 57 (44%) of the 130 nonarboviruses; an few published records of bats being routinely sampled during additional 5 (8%) are presumed on epidemiological evidence to arbovirus studies, and only two arboviruses (3%) have been iso- have nonhuman reservoirs (Table S1). A number of viruses infect lated from bats. The reason a much larger number of arbovirus more than one nonhuman vertebrate host species and it is likely species (n = 16) have been isolated from birds than have that the variety of hosts is wider than has been recorded. The nonarbovirus species (n = 1) might, however, be characteristic of predominant host groups for arboviruses (n = 70) are nonhuman the pathogenicity of the togaviruses and flaviviruses, which are primates (31%), rodents (29%), ungulates (26%), and birds (23%); much more common among the arboviruses. The most prominent for the nonarboviruses (n = 57), they are rodents (30%), ungu- vectors of arboviruses were mosquitoes (67%), ticks (19%), and lates (26%), bats (23%), and primates (16%). -

Spillback in the Anthropocene: the Risk of Human-To-Wildlife Pathogen Transmission for Conservation and Public Health
Spillback in the Anthropocene: the risk of human-to-wildlife pathogen transmission for conservation and public health Anna C. Fagre*1, Lily E. Cohen2, Evan A. Eskew3, Max Farrell4, Emma Glennon5, Maxwell B. Joseph6, Hannah K. Frank7, Sadie Ryan8, 9,10, Colin J Carlson11,12, Gregory F Albery*13 *Corresponding authors: [email protected]; [email protected] 1: Department of Microbiology, Immunology, and Pathology, College of Veterinary Medicine and Biomedical Sciences, Colorado State University, Fort Collins, CO 2: Icahn School of Medicine at Mount Sinai, New York, NY 10029 3: Department of Biology, Pacific Lutheran University, Tacoma, WA, 98447 USA 4: Department of Ecology & Evolutionary Biology, University of Toronto 5: Disease Dynamics Unit, Department of Veterinary Medicine, University of Cambridge, Cambridge CB3 0ES, UK 6: Earth Lab, University of Colorado Boulder, Boulder, CO 80309 7: Department of Ecology and Evolutionary Biology, Tulane University, New Orleans, LA, 70118 USA 8: Quantitative Disease Ecology and Conservation (QDEC) Lab Group, Department of Geography, University of Florida, Gainesville, FL, 32610 USA 9: Emerging Pathogens Institute, University of Florida, Gainesville, FL, 32610 USA 10: School of Life Sciences, University of KwaZulu-Natal, Durban, 4041, South Africa 11: Center for Global Health Science and Security, Georgetown University Medical Center, Washington, DC, 20057 USA 12: Department of Microbiology and Immunology, Georgetown University Medical Center, Washington, DC, 20057 USA 13: Department of Biology, Georgetown University, Washington, DC, 20057 USA 1 Abstract The SARS-CoV-2 pandemic has led to increased concern over transmission of pathogens from humans to animals (“spillback”) and its potential to threaten conservation and public health. -

Risk of Human Pathogen Internalization in Leafy Vegetables During Lab-Scale Hydroponic Cultivation
horticulturae Review Risk of Human Pathogen Internalization in Leafy Vegetables During Lab-Scale Hydroponic Cultivation Gina M. Riggio 1, Sarah L. Jones 2 and Kristen E. Gibson 2,* 1 Cellular and Molecular Biology Program, Department of Food Science, University of Arkansas, Fayetteville, AR 72701, USA; [email protected] 2 Department of Food Science, University of Arkansas, Fayetteville, AR 72704, USA; [email protected] * Correspondence: [email protected]; Tel.: +1-479-575-6844 Received: 13 February 2019; Accepted: 7 March 2019; Published: 15 March 2019 Abstract: Controlled environment agriculture (CEA) is a growing industry for the production of leafy vegetables and fresh produce in general. Moreover, CEA is a potentially desirable alternative production system, as well as a risk management solution for the food safety challenges within the fresh produce industry. Here, we will focus on hydroponic leafy vegetable production (including lettuce, spinach, microgreens, and herbs), which can be categorized into six types: (1) nutrient film technique (NFT), (2) deep water raft culture (DWC), (3) flood and drain, (4) continuous drip systems, (5) the wick method, and (6) aeroponics. The first five are the most commonly used in the production of leafy vegetables. Each of these systems may confer different risks and advantages in the production of leafy vegetables. This review aims to (i) address the differences in current hydroponic system designs with respect to human pathogen internalization risk, and (ii) identify the preventive control points for reducing risks related to pathogen contamination in leafy greens and related fresh produce products. Keywords: hydroponic; leafy greens; internalization; pathogens; norovirus; Escherichia coli; Salmonella; Listeria spp.; preventive controls 1. -

Genomics-Based Re-Examination of the Taxonomy and Phylogeny Of
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Food and Drug Administration Papers U.S. Department of Health and Human Services 2020 Genomics-based re-examination of the taxonomy and phylogeny of human and simian Mastadenoviruses: an evolving whole genomes approach, revealing putative zoonosis, anthroponosis, and amphizoonosis June Kang George Mason University Ashrafali Mohamed Ismail Harvard Medical School Shoaleh Dehghan George Mason University Jaya Rajaiya Harvard Medical School FMarollowc W this. Allar andd additional works at: https://digitalcommons.unl.edu/usfda US Food & Drug Administration Part of the Dietetics and Clinical Nutrition Commons, Health and Medical Administration Commons, Health Services Administration Commons, Pharmaceutical Preparations Commons, and the Pharmacy AdministrSee next pageation, for Policy additional and Regulation authors Commons Kang, June; Ismail, Ashrafali Mohamed; Dehghan, Shoaleh; Rajaiya, Jaya; Allard, Marc W.; Lim, Haw Chuan; Dyer, David W.; Chodosh, James; and Seto, Donald, "Genomics-based re-examination of the taxonomy and phylogeny of human and simian Mastadenoviruses: an evolving whole genomes approach, revealing putative zoonosis, anthroponosis, and amphizoonosis" (2020). Food and Drug Administration Papers. 47. https://digitalcommons.unl.edu/usfda/47 This Article is brought to you for free and open access by the U.S. Department of Health and Human Services at DigitalCommons@University of Nebraska - Lincoln. It has been accepted for inclusion in Food and -

Laboratory Activities to Enhance the Study of Whole Blood Components and Their Role in the Immune Response
Laboratory Activities to Enhance the Study of Whole Blood Components and their Role in the Immune Response Thomas E. Schmit John H. Wallace Teacher Fellow Hamilton High School 327 Fairgrounds Road Hamilton, MT 59840 [email protected] Table Of Contents Teacher Guide Overview...................................................................................................................1 Science Background..................................................................................................1 Student Outcomes......................................................................................................4 Learning Objectives....................................................................................................4 Time Requirements.....................................................................................................4 Advance Preparation...................................................................................................4 Materials and Equipment............................................................................................4 Student Prior Knowledge............................................................................................6 What Is Expected From the Students..........................................................................7 Anticipated Results.....................................................................................................7 Classroom Discussion.................................................................................................8 -

Ecology of Pathogenic E.Coli - Stefano Morabito
MEDICAL SCIENCES - Ecology Of Pathogenic E.Coli - Stefano Morabito ECOLOGY OF PATHOGENIC E. coli Stefano Morabito European Reference Laboratory for Escherichia coli including VTEC. Foodborne zoonoses Unit; Department of Veterinary Public Health and Food Safety Istituto Superiore di Sanità, Viale Regina Elena 299, 00161, Rome. Italy. Keywords: Escherichia coli, pathogenicity, genomic plasticity, survival, environment, colonization, animal reservoirs, zoonosis. Contents 1. Introduction 2. Pathogenic E. coli types 3. Ecology of Pathogenic E. coli Circulating in Humans 4. The Zoonotic Connection 5. Ecology of Zoonotic E. coli 6. Pathogenic Escherichia coli Producing Biofilm 7. Signaling In the Gastro-Intestinal Tract and Pathogenesis 8. E. coli Pathotypes with Atypical Combination of Virulence Factors 9. Acknowledgments Glossary Bibliography Biographical Sketch Summary Escherichia coli is a ubiquitous bacterial species. It is part of the intestinal microflora of humans and warm-blooded animals and, at the same time, is among the most diffuse bacterial species in the environment. The pivotal aspect of this success is represented by an exceptional genomic plasticity. As a matter of fact, E. coli is capable to exchange genetic material with other bacteria, often belonging to different species, through horizontal gene transfer, a process involving the action of mobile genetic elements, such as bacteriophages, transposons, pathogenicity islands and plasmids. The relationships between E. coli and the human host are generally based on commensalism and yet this species exerts a beneficial effect in the gut, however, some strains evolved the capability to harm and cause disease either involving the gastro- intestinal tract or in other organism‘s districts. Most of the pathogenic E. -

The Characteristics of Pandemic Pathogens
Center for Health Security The Characteristics of PANDEMIC PATHOGENS Improving Pandemic Preparedness by Identifying the Attributes of Microorganisms Most Likely to Cause a Global Catastrophic Biological Event Center for Health Security The Characteristics of PANDEMIC PATHOGENS PROJECT TEAM Amesh A. Adalja, MD, Project Director Matthew Watson, BS Eric S. Toner, MD Anita Cicero, JD Thomas V. Inglesby, MD Copyright © 2018 by Johns Hopkins University EXECUTIVE SUMMARY The Characteristics of PANDEMIC PATHOGENS Background and Purpose of Report CONTENTS he Johns Hopkins Center for Health Security conducted this study to elucidate the characteristics T of naturally occurring microorganisms that constitute a global catastrophic biological risk (GCBR). 3 Executive Summary GCBRs are defined as “those events in which biological agents—whether naturally emerging or 8 Introduction 9 Purpose, Methods, and Analysis reemerging, deliberately created and released, or laboratory engineered and escaped—could lead to 10 Basis of Recommendations sudden, extraordinary, widespread disaster beyond the collective capability of national and international 18 Recommendations governments and the private sector to control. If unchecked, GCBRs would lead to great suffering, loss of 22 Future Directions life, and sustained damage to national governments, international relationships, economies, societal 22 Conclusion 23 References stability, or global security.” 26 Appendix A: List of Experts Interviewed 28 Appendix B: Meeting Participants The overarching aim of the study was to provide an inductive, microbe-agnostic analysis of the microbial world to identify fundamental principles that underlie this special category of microorganisms that have potential to cause global catastrophe. Such principles could refine pandemic preparedness by providing a new framework or lens through which to survey the threat landscape of infectious diseases in order to better anticipate, prepare for, and respond to GCBR threats.