Ecology of Pathogenic E.Coli - Stefano Morabito
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Human Illness Caused by E. Coli O157:H7 from Food and Non-Food Sources
FRI BRIEFINGS Human Illness Caused by E. coli O157:H7 from Food and Non-food Sources M. Ellin Doyle1*, John Archer2, Charles W. Kaspar1, and Ronald Weiss1 1Food Research Institute, University of Wisconsin–Madison, Madison, WI 53706 2Wisconsin Division of Public Health, Bureau of Communicable Diseases and Preparedness, Communicable Disease Epidemiology Section, Madison, WI 53702 Contents Introduction ...................................................................................................................................2 Epidemiology of E. coli O157:H7..................................................................................................2 Outbreak Data ........................................................................................................................2 Reservoirs of E. coli O157:H7 ..............................................................................................3 Cattle—the primary reservoir ........................................................................................3 Other ruminants .............................................................................................................4 Other animals .................................................................................................................4 Transport Hosts......................................................................................................................4 Routes of Human Infection ....................................................................................................5 -

How Do Pathogenic Microorganisms Develop Cross-Kingdom Host Jumps? Peter Van Baarlen1, Alex Van Belkum2, Richard C
Molecular mechanisms of pathogenicity: how do pathogenic microorganisms develop cross-kingdom host jumps? Peter van Baarlen1, Alex van Belkum2, Richard C. Summerbell3, Pedro W. Crous3 & Bart P.H.J. Thomma1 1Laboratory of Phytopathology, Wageningen University, Wageningen, The Netherlands; 2Department of Medical Microbiology and Infectious Diseases, Erasmus MC, University Medical Centre Rotterdam, Rotterdam, The Netherlands; and 3CBS Fungal Biodiversity Centre, Utrecht, The Netherlands Correspondence: Bart P.H.J. Thomma, Abstract Downloaded from https://academic.oup.com/femsre/article/31/3/239/2367343 by guest on 27 September 2021 Laboratory of Phytopathology, Wageningen University, Binnenhaven 5, 6709 PD It is common knowledge that pathogenic viruses can change hosts, with avian Wageningen, The Netherlands. Tel.: 10031 influenza, the HIV, and the causal agent of variant Creutzfeldt–Jacob encephalitis 317 484536; fax: 10031 317 483412; as well-known examples. Less well known, however, is that host jumps also occur e-mail: [email protected] with more complex pathogenic microorganisms such as bacteria and fungi. In extreme cases, these host jumps even cross kingdom of life barriers. A number of Received 3 July 2006; revised 22 December requirements need to be met to enable a microorganism to cross such kingdom 2006; accepted 23 December 2006. barriers. Potential cross-kingdom pathogenic microorganisms must be able to First published online 26 February 2007. come into close and frequent contact with potential hosts, and must be able to overcome or evade host defences. Reproduction on, in, or near the new host will DOI:10.1111/j.1574-6976.2007.00065.x ensure the transmission or release of successful genotypes. -

Bacterial Dissimilation of Citric Acid Carl Robert Brewer Iowa State College
Iowa State University Capstones, Theses and Retrospective Theses and Dissertations Dissertations 1939 Bacterial dissimilation of citric acid Carl Robert Brewer Iowa State College Follow this and additional works at: https://lib.dr.iastate.edu/rtd Part of the Microbiology Commons Recommended Citation Brewer, Carl Robert, "Bacterial dissimilation of citric acid " (1939). Retrospective Theses and Dissertations. 13227. https://lib.dr.iastate.edu/rtd/13227 This Dissertation is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Retrospective Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. INFORMATION TO USERS This manuscript has been reproduced from the microfilm master. UMI films the text directly from the original or copy submitted. Thus, some thesis and dissertation copies are in typewriter face, while others may be from any type of computer printer. The quality of this reproduction is dependent upon the quality of the copy submitted. Brol<en or indistinct print, colored or poor quality illustrations and photographs, print bleedthrough, substandard margins, and improper alignment can adversely affect reproduction. In the unlikely event that the author did not send UMI a complete manuscript and there are missing pages, these will be noted. Also, if unauthorized copyright material had to be removed, a note will indicate the deletion. Oversize materials (e.g., maps, drawings, charts) are reproduced by sectioning the original, beginning at the upper left-hand comer and continuing from left to right in equal sections with small overlaps. -

Differences in Innate Immune Response Between Man and Mouse
Institute of Medical Physics and Biophysics Medical Department, Leipzig University Author Manuscript © 2014. This manuscript version is made available under the CC-BY-NC-ND 4.0 license http://creativecommons.org/licenses/by-nc-nd/4.0/ Published in final edited form as: Critical Reviews™ in Immunology, 2014; 34:433-54. Available at: http://dx.doi.org/ 10.1615/CritRevImmunol.2014011600. DIFFERENCES IN INNATE IMMUNE RESPONSE BETWEEN MAN AND MOUSE Josefin Zschaler 1,2,*, Denise Schlorke 1,2,* & Juergen Arnhold 1,2 1 Institute for Medical Physics and Biophysics, Medical Faculty, Leipzig University, Leipzig, Germany 2 Translational Centre for Regenerative Medicine (TRM), Leipzig University, Leipzig, Germany * Both authors contributed equally to this work. Abstract Mouse strains are frequently used to model human disease states, to test the efficiency of drugs and therapeutic principles. However, the direct translation of murine experimental data to human pathological events often fails due to sufficient differences in the organization of the immune system of both species. Here we give a short overview of the principle differences between mice and humans in defense strategies against pathogens and mechanisms involved in response to pathogenic microorganisms and other activators of the immune system. While in human blood mechanisms of immune resistance are highly prevailed, tolerance mechanisms dominate for the defense against pathogenic microorganisms in mouse blood. Further on, species-related differences of immune cells mainly involved in innate immune response as well as differences to maintain oxidative homeostasis are also considered. A number of disease scenarios in mice are critically reflected for their suitability to serve as a model for human pathologies. -

The Public Health Impact of Prion Diseases1
10 Feb 2005 13:10 AR AR238-PU26-08.tex AR238-PU26-08.sgm LaTeX2e(2002/01/18) P1: IBD 10.1146/annurev.publhealth.26.021304.144536 Annu. Rev. Public Health 2005. 26:191–212 doi: 10.1146/annurev.publhealth.26.021304.144536 Copyright ©c 2005 by Annual Reviews. All rights reserved First published online as a Review in Advance on December 8, 2004 THE PUBLIC HEALTH IMPACT OF PRION DISEASES1 Ermias D. Belay and Lawrence B. Schonberger Division of Viral and Rickettsial Diseases, National Center for Infectious Diseases, Centers for Disease Control and Prevention, Atlanta, Georgia 30333; email: [email protected] KeyWords transmissible spongiform encephalopathy, Creutzfeldt-Jakob disease, variant Creutzfeldt-Jakob disease, bovine spongiform encephalopathy, chronic wasting disease ■ Abstract Several prion disease–related human health risks from an exogenous source can be identified in the United States, including the iatrogenic transmission of Creutzfeldt-Jakob disease (CJD), the possible occurrence of variant CJD (vCJD), and potential zoonotic transmission of chronic wasting disease (CWD). Although cross- species transmission of prion diseases seems to be limited by an apparent “species barrier,” the occurrence of bovine spongiform encephalopathy (BSE) and its transmis- sion to humans indicate that animal prion diseases can pose a significant public health risk. Recent reports of secondary person-to-person spread of vCJD via blood products and detection of vCJD transmission in a patient heterozygous at codon 129 further illustrate the potential public health impacts of BSE. INTRODUCTION by IRMO/Information Center on 03/14/05. For personal use only. Prion diseases, also known as transmissible spongiform encephalopathies (TSEs), are a group of animal and human brain diseases that are uniformly fatal and often characterized by a long incubation period and a multifocal neuropathologic picture of neuronal loss, spongiform changes, and astrogliosis (3). -

Innate Immunity Part I: the Immediate Response Deborah A
Innate Immunity Part I: The Immediate Response Deborah A. Lebman, Ph.D. OBJECTIVES 1. Describe three mechanisms present at epithelial surfaces that protect the host from pathogens. 2. List 3 functions of complement proteins in innate immunity 3. Know the primary functions of macrophages and neutrophils 4. List the receptors on macrophages involved in recognition of pathogens 5. List the key toxic products of phagocytes READING • Parham, The Immune System, pg 227-238. TERMS • Defensin: antibacterial peptide produced by the host at epithelial surfaces • Complement: ubiquitously present proteins synthesized primarily in the liver that mediate a variety of responses to pathogens • Opsonin (opsonization): alteration of cell surface that facilitates phagocytosis • Phagocytosis: internalization of particulate matter by cells • CD14: LPS receptor • TLR-4: toll like receptor 4 I. Introduction A. The first response to pathogens occurs immediately B. The early response employs both mechanical and cell mediated mechanisms C. There is no cellular expansion, but there is cell recruitment D. A wide array of soluble mediators with both positive and negative effects, i.e. remove the pathogen but hurt the host, is released II. Overview of Pathogens A. There are four types of pathogen: bacteria, virus, fungus, parasite B. Table I: Examples of the Four Types of Common Human Pathogen Table 1: Examples of the Four Types of Common Human Pathogen Table 1 (Continued): Examples of the Four Types of Common Human Pathogen C. Pathogens can be intracellular or extracellular Table 2: Examples of intracellular and extracellular pathogens. Notice that both innate and adaptive mechanisms are involved in the response to both types of pathogen. -

Quorum Sensing: Understanding the Role of Bacteria in Meat Spoilage
CORE Metadata, citation and similar papers at core.ac.uk Provided by Cranfield CERES CRANFIELD UNIVERSITY Cranfield Health PhD THESIS VASILIKI A. BLANA QUORUM SENSING: UNDERSTANDING THE ROLE OF BACTERIA IN MEAT SPOILAGE Supervisors: 1st Professor Naresh Magan and 2nd Professor George-John Nychas 2010 ABSTRACT Quorum sensing is a fundamental process to all of microbiology since it is ubiquitous in the bacterial world, where bacterial cells communicate with each other using low molecular weight signal molecules called autoinducers. Despite the fact that quorum sensing regulates numerous bacterial behaviours, very few studies have addressed the role of this phenomenon in foods. The microbial association of beef consists mainly of pseudomonads, Enterobacteriaceae, Brochothrix thermosphacta and lactic acid bacteria as revealed by minced beef samples purchased from retail shops, which fluctuates according to the storage conditions. Certain members of the microbial association, which are considered to produce signal molecules, have been found to be major contributors to meat spoilage. Pseudomonas fragi and Enterobacteriaceae strains, i.e., Hafnia alvei and Serratia liquefaciens are among the most common quorum sensing signal producers recovered from various food environments. N-acyl homoserine lactone (AHL) and autoinducer-2 (AI-2) signal molecules were found to be present in meat stored under different conditions (i.e., temperature and packaging), and correlated with the ephemeral spoilage organisms that comprise the microbial community generally -

Citrobacter Koseri, Levinea Malonatica
INTERNATIONALJOURNAL OF SYSTEMATICBACTERIOLOGY, Jan. 1990, p. 107-108 Vol. 40, No. 1 0020-7713/90/010107-02$02.oo/o Copyright 0 1990, International Union of Microbiological Societies Correct Names of the Species Citrobacter koseri, Levinea malonatica , and Citrobacter diversus Request for an Opinion WILHELM FREDERIKSEN Department of Diagnostic Bacteriology and Antibiotics, Statens Seruminstitut, DK-2300 Copenhagen S,Denmark The single species carrying the three names Citrobacter koseri, Levinea malonatica, and Citrobacter diversus differs by at least eight characteristics from Citrobacter diversum as described by Werkman and Gillen in 1932. It is obviously not the same organism. Accordingly, the species should not carry the name proposed by Werkman and Gillen. I request that the name Citrobacter diversus be placed on the list of nomina rejicienda. Citrobacter koseri is the correct name. Levinea koseri is a correct combination when the genus Levinea is accepted. The epithet maZoonatica is a later synonym of the epithet koseri. In 1932 Werkman and Gillen (7) proposed a new genus, taxon C. diversum described by Werkman and Gillen in the Citrobacter, containing seven species. The description of following characteristics: motility, production of H,S, and Citrobacter diversum (sic) of Werkman and Gillen was based production of acid from inositol and raffinose. To this can be on two strains. These strains have not been kept available in added production of acid from glycogen, melizitose, starch, collections, and the name C. diversum did not come into and galactose, as Table 1 shows. general use. Ewing and Davis obviously accepted four deviations to In 1970 Frederiksen (4) described a new species which he lead to the statement about reactions “similar to those named Citrobacter koseri. -

Biomedical Applications of Bacteria-Derived Polymers
polymers Review Biomedical Applications of Bacteria-Derived Polymers Jonathan David Hinchliffe, Alakananda Parassini Madappura, Syed Mohammad Daniel Syed Mohamed and Ipsita Roy * Department of Materials Science and Engineering, Faculty of Engineering, University of Sheffield, Sheffield S1 3JD, UK; jhinchliffe3@sheffield.ac.uk (J.D.H.); [email protected] (A.P.M.); smdsyedmohamed1@sheffield.ac.uk (S.M.D.S.M.) * Correspondence: I.Roy@sheffield.ac.uk; Tel.: +44-11-4222-5962 Abstract: Plastics have found widespread use in the fields of cosmetic, engineering, and medical sciences due to their wide-ranging mechanical and physical properties, as well as suitability in biomedical applications. However, in the light of the environmental cost of further upscaling current methods of synthesizing many plastics, work has recently focused on the manufacture of these polymers using biological methods (often bacterial fermentation), which brings with them the advantages of both low temperature synthesis and a reduced reliance on potentially toxic and non-eco-friendly compounds. This can be seen as a boon in the biomaterials industry, where there is a need for highly bespoke, biocompatible, processable polymers with unique biological properties, for the regeneration and replacement of a large number of tissue types, following disease. However, barriers still remain to the mass-production of some of these polymers, necessitating new research. This review attempts a critical analysis of the contemporary literature concerning the use of a number of bacteria-derived polymers in the context of biomedical applications, including the biosynthetic Citation: Hinchliffe, J.D.; Parassini pathways and organisms involved, as well as the challenges surrounding their mass production. -

Chapter 4 Antimicrobial Properties of Organosulfur Compounds
Chapter 4 Antimicrobial Properties of Organosulfur Compounds Osman Sagdic and Fatih Tornuk Abstract Organosulfur compounds are defi ned as organic molecules containing one or more carbon-sulfur bonds. These compounds are present particularly in Allium and Brassica vegetables and are converted to a variety of other sulfur con- taining compounds via hydrolysis by several herbal enzymes when the intact bulbs are damaged or cut. Sulfur containing hydrolysis products constitute very diverse chemical structures and exhibit several bioactive properties as well as antimicrobial. The antimicrobial activity of organosulfur compounds has been reported against a wide spectrum of bacteria, fungi and viruses. Despite the wide antimicrobial spec- trum, their pungent fl avor/odor is the most considerable factor restricting their com- mon use in foods as antimicrobial additives. However, meat products might be considered as the most suitable food materials in this respect since Allium and Brassica vegetables especially garlic and onion have been used as fl avoring and preservative agents in meat origin foods. In this chapter, the chemical diversity and in vitro and in food antimicrobial activity of the organosulfur compounds of Allium and Brassica plants are summarized. Keywords Organosulfur compounds • Garlic • Onion • Allium • Brassica • Thiosulfi nates • Glucosinolates O. Sagdic (*) Department of Food Engineering, Faculty of Chemical and Metallurgical Engineering , Yildiz Teknik University , 34220 Esenler , Istanbul , Turkey e-mail: [email protected] F. Tornuk S a fi ye Cikrikcioglu Vocational College , Erciyes University , 38039 Kayseri , Turkey A.K. Patra (ed.), Dietary Phytochemicals and Microbes, 127 DOI 10.1007/978-94-007-3926-0_4, © Springer Science+Business Media Dordrecht 2012 128 O. -

Wedding Higher Taxonomic Ranks with Metabolic Signatures Coded in Prokaryotic Genomes
Wedding higher taxonomic ranks with metabolic signatures coded in prokaryotic genomes Gregorio Iraola*, Hugo Naya* Corresponding authors: E-mail: [email protected], [email protected] This PDF file includes: Supplementary Table 1 Supplementary Figures 1 to 4 Supplementary Methods SUPPLEMENTARY TABLES Supplementary Tab. 1 Supplementary Tab. 1. Full prediction for the set of 108 external genomes used as test. genome domain phylum class order family genus prediction alphaproteobacterium_LFTY0 Bacteria Proteobacteria Alphaproteobacteria Rhodobacterales Rhodobacteraceae Unknown candidatus_nasuia_deltocephalinicola_PUNC_CP013211 Bacteria Proteobacteria Gammaproteobacteria Unknown Unknown Unknown candidatus_sulcia_muelleri_PUNC_CP013212 Bacteria Bacteroidetes Flavobacteriia Flavobacteriales NA Candidatus Sulcia deinococcus_grandis_ATCC43672_BCMS0 Bacteria Deinococcus-Thermus Deinococci Deinococcales Deinococcaceae Deinococcus devosia_sp_H5989_CP011300 Bacteria Proteobacteria Unknown Unknown Unknown Unknown micromonospora_RV43_LEKG0 Bacteria Actinobacteria Actinobacteria Micromonosporales Micromonosporaceae Micromonospora nitrosomonas_communis_Nm2_CP011451 Bacteria Proteobacteria Betaproteobacteria Nitrosomonadales Nitrosomonadaceae Unknown nocardia_seriolae_U1_BBYQ0 Bacteria Actinobacteria Actinobacteria Corynebacteriales Nocardiaceae Nocardia nocardiopsis_RV163_LEKI01 Bacteria Actinobacteria Actinobacteria Streptosporangiales Nocardiopsaceae Nocardiopsis oscillatoriales_cyanobacterium_MTP1_LNAA0 Bacteria Cyanobacteria NA Oscillatoriales -
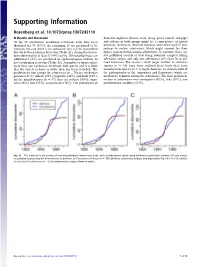
Supporting Information
Supporting Information Rosenberg et al. 10.1073/pnas.1307243110 SI Results and Discussion domestic ungulates (horses, cows, sheep, goats, camels, and pigs) Of the 83 arboviruses, nonhuman vertebrate hosts have been and rodents in both groups might be a consequence of spatial identified for 70 (84%); the remaining 13 are presumed to be proximity to humans. Sentinel monkeys were often used in pro- zoonoses because there is no indication they can be transmitted cedures to isolate arboviruses, which might account for their directly between humans by vectors (Table S1). Animal hosts have higher representation among arboviruses. In contrast, there are been identified for at least 57 (44%) of the 130 nonarboviruses; an few published records of bats being routinely sampled during additional 5 (8%) are presumed on epidemiological evidence to arbovirus studies, and only two arboviruses (3%) have been iso- have nonhuman reservoirs (Table S1). A number of viruses infect lated from bats. The reason a much larger number of arbovirus more than one nonhuman vertebrate host species and it is likely species (n = 16) have been isolated from birds than have that the variety of hosts is wider than has been recorded. The nonarbovirus species (n = 1) might, however, be characteristic of predominant host groups for arboviruses (n = 70) are nonhuman the pathogenicity of the togaviruses and flaviviruses, which are primates (31%), rodents (29%), ungulates (26%), and birds (23%); much more common among the arboviruses. The most prominent for the nonarboviruses (n = 57), they are rodents (30%), ungu- vectors of arboviruses were mosquitoes (67%), ticks (19%), and lates (26%), bats (23%), and primates (16%).