Mouse Rgs18 Conditional Knockout Project (CRISPR/Cas9)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
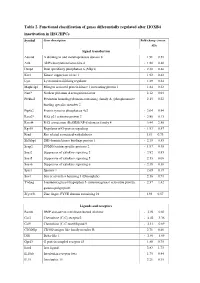
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Global H3k4me3 Genome Mapping Reveals Alterations of Innate Immunity Signaling and Overexpression of JMJD3 in Human Myelodysplastic Syndrome CD34 Þ Cells
Leukemia (2013) 27, 2177–2186 & 2013 Macmillan Publishers Limited All rights reserved 0887-6924/13 www.nature.com/leu ORIGINAL ARTICLE Global H3K4me3 genome mapping reveals alterations of innate immunity signaling and overexpression of JMJD3 in human myelodysplastic syndrome CD34 þ cells YWei1, R Chen2, S Dimicoli1, C Bueso-Ramos3, D Neuberg4, S Pierce1, H Wang2, H Yang1, Y Jia1, H Zheng1, Z Fang1, M Nguyen3, I Ganan-Gomez1,5, B Ebert6, R Levine7, H Kantarjian1 and G Garcia-Manero1 The molecular bases of myelodysplastic syndromes (MDS) are not fully understood. Trimethylated histone 3 lysine 4 (H3K4me3) is present in promoters of actively transcribed genes and has been shown to be involved in hematopoietic differentiation. We performed a genome-wide H3K4me3 CHIP-Seq (chromatin immunoprecipitation coupled with whole genome sequencing) analysis of primary MDS bone marrow (BM) CD34 þ cells. This resulted in the identification of 36 genes marked by distinct higher levels of promoter H3K4me3 in MDS. A majority of these genes are involved in nuclear factor (NF)-kB activation and innate immunity signaling. We then analyzed expression of histone demethylases and observed significant overexpression of the JmjC-domain histone demethylase JMJD3 (KDM6b) in MDS CD34 þ cells. Furthermore, we demonstrate that JMJD3 has a positive effect on transcription of multiple CHIP-Seq identified genes involved in NF-kB activation. Inhibition of JMJD3 using shRNA in primary BM MDS CD34 þ cells resulted in an increased number of erythroid colonies in samples isolated from patients with lower-risk MDS. Taken together, these data indicate the deregulation of H3K4me3 and associated abnormal activation of innate immunity signals have a role in the pathogenesis of MDS and that targeting these signals may have potential therapeutic value in MDS. -

Identification of Key Pathways and Genes in Dementia Via Integrated Bioinformatics Analysis
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.18.440371; this version posted July 19, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Identification of Key Pathways and Genes in Dementia via Integrated Bioinformatics Analysis Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karnataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2021.04.18.440371; this version posted July 19, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract To provide a better understanding of dementia at the molecular level, this study aimed to identify the genes and key pathways associated with dementia by using integrated bioinformatics analysis. Based on the expression profiling by high throughput sequencing dataset GSE153960 derived from the Gene Expression Omnibus (GEO), the differentially expressed genes (DEGs) between patients with dementia and healthy controls were identified. With DEGs, we performed a series of functional enrichment analyses. Then, a protein–protein interaction (PPI) network, modules, miRNA-hub gene regulatory network and TF-hub gene regulatory network was constructed, analyzed and visualized, with which the hub genes miRNAs and TFs nodes were screened out. Finally, validation of hub genes was performed by using receiver operating characteristic curve (ROC) analysis. -

Genome Sequencing Unveils a Regulatory Landscape of Platelet Reactivity
ARTICLE https://doi.org/10.1038/s41467-021-23470-9 OPEN Genome sequencing unveils a regulatory landscape of platelet reactivity Ali R. Keramati 1,2,124, Ming-Huei Chen3,4,124, Benjamin A. T. Rodriguez3,4,5,124, Lisa R. Yanek 2,6, Arunoday Bhan7, Brady J. Gaynor 8,9, Kathleen Ryan8,9, Jennifer A. Brody 10, Xue Zhong11, Qiang Wei12, NHLBI Trans-Omics for Precision (TOPMed) Consortium*, Kai Kammers13, Kanika Kanchan14, Kruthika Iyer14, Madeline H. Kowalski15, Achilleas N. Pitsillides4,16, L. Adrienne Cupples 4,16, Bingshan Li 12, Thorsten M. Schlaeger7, Alan R. Shuldiner9, Jeffrey R. O’Connell8,9, Ingo Ruczinski17, Braxton D. Mitchell 8,9, ✉ Nauder Faraday2,18, Margaret A. Taub17, Lewis C. Becker1,2, Joshua P. Lewis 8,9,125 , 2,14,125✉ 3,4,125✉ 1234567890():,; Rasika A. Mathias & Andrew D. Johnson Platelet aggregation at the site of atherosclerotic vascular injury is the underlying patho- physiology of myocardial infarction and stroke. To build upon prior GWAS, here we report on 16 loci identified through a whole genome sequencing (WGS) approach in 3,855 NHLBI Trans-Omics for Precision Medicine (TOPMed) participants deeply phenotyped for platelet aggregation. We identify the RGS18 locus, which encodes a myeloerythroid lineage-specific regulator of G-protein signaling that co-localizes with expression quantitative trait loci (eQTL) signatures for RGS18 expression in platelets. Gene-based approaches implicate the SVEP1 gene, a known contributor of coronary artery disease risk. Sentinel variants at RGS18 and PEAR1 are associated with thrombosis risk and increased gastrointestinal bleeding risk, respectively. Our WGS findings add to previously identified GWAS loci, provide insights regarding the mechanism(s) by which genetics may influence cardiovascular disease risk, and underscore the importance of rare variant and regulatory approaches to identifying loci contributing to complex phenotypes. -

Supplementary Information.Pdf
Supplementary Information Whole transcriptome profiling reveals major cell types in the cellular immune response against acute and chronic active Epstein‐Barr virus infection Huaqing Zhong1, Xinran Hu2, Andrew B. Janowski2, Gregory A. Storch2, Liyun Su1, Lingfeng Cao1, Jinsheng Yu3, and Jin Xu1 Department of Clinical Laboratory1, Children's Hospital of Fudan University, Minhang District, Shanghai 201102, China; Departments of Pediatrics2 and Genetics3, Washington University School of Medicine, Saint Louis, Missouri 63110, United States. Supplementary information includes the following: 1. Supplementary Figure S1: Fold‐change and correlation data for hyperactive and hypoactive genes. 2. Supplementary Table S1: Clinical data and EBV lab results for 110 study subjects. 3. Supplementary Table S2: Differentially expressed genes between AIM vs. Healthy controls. 4. Supplementary Table S3: Differentially expressed genes between CAEBV vs. Healthy controls. 5. Supplementary Table S4: Fold‐change data for 303 immune mediators. 6. Supplementary Table S5: Primers used in qPCR assays. Supplementary Figure S1. Fold‐change (a) and Pearson correlation data (b) for 10 cell markers and 61 hypoactive and hyperactive genes identified in subjects with acute EBV infection (AIM) in the primary cohort. Note: 23 up‐regulated hyperactive genes were highly correlated positively with cytotoxic T cell (Tc) marker CD8A and NK cell marker CD94 (KLRD1), and 38 down‐regulated hypoactive genes were highly correlated positively with B cell, conventional dendritic cell -

The Genetic Basis of Emotional Behaviour in Mice
European Journal of Human Genetics (2006) 14, 721–728 & 2006 Nature Publishing Group All rights reserved 1018-4813/06 $30.00 www.nature.com/ejhg REVIEW The genetic basis of emotional behaviour in mice Saffron AG Willis-Owen*,1 and Jonathan Flint1 1Wellcome Trust Centre for Human Genetics, Roosevelt Drive, Headington, Oxford, UK The last decade has witnessed a steady expansion in the number of quantitative trait loci (QTL) mapped for complex phenotypes. However, despite this proliferation, the number of successfully cloned QTL has remained surprisingly low, and to a great extent limited to large effect loci. In this review, we follow the progress of one complex trait locus; a low magnitude moderator of murine emotionality identified some 10 years ago in a simple two-strain intercross, and successively resolved using a variety of crosses and fear-related phenotypes. These experiments have revealed a complex underlying genetic architecture, whereby genetic effects fractionate into several separable QTL with some evidence of phenotype specificity. Ultimately, we describe a method of assessing gene candidacy, and show that given sufficient access to genetic diversity and recombination, progression from QTL to gene can be achieved even for low magnitude genetic effects. European Journal of Human Genetics (2006) 14, 721–728. doi:10.1038/sj.ejhg.5201569 Keywords: mouse; emotionality; quantitative trait locus; QTL Introduction tude genetic effects and their interactions (both with other Emotionality is a psychological trait of complex aetiology, genetic loci (ie epistasis) and nongenetic (environmental) which moderates an organism’s response to stress. Beha- factors) ultimately producing a quasi-continuously distrib- vioural evidence of emotionality has been documented uted phenotype. -

Biological Role and Disease Impact of Copy Number Variation in Complex Disease
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations 2014 Biological Role and Disease Impact of Copy Number Variation in Complex Disease Joseph Glessner University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Bioinformatics Commons, and the Genetics Commons Recommended Citation Glessner, Joseph, "Biological Role and Disease Impact of Copy Number Variation in Complex Disease" (2014). Publicly Accessible Penn Dissertations. 1286. https://repository.upenn.edu/edissertations/1286 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/1286 For more information, please contact [email protected]. Biological Role and Disease Impact of Copy Number Variation in Complex Disease Abstract In the human genome, DNA variants give rise to a variety of complex phenotypes. Ranging from single base mutations to copy number variations (CNVs), many of these variants are neutral in selection and disease etiology, making difficult the detection of true common orar r e frequency disease-causing mutations. However, allele frequency comparisons in cases, controls, and families may reveal disease associations. Single nucleotide polymorphism (SNP) arrays and exome sequencing are popular assays for genome-wide variant identification. oT limit bias between samples, uniform testing is crucial, including standardized platform versions and sample processing. Bases occupy single points while copy variants occupy segments. -

RGS18 (F-5): Sc-390908
SANTA CRUZ BIOTECHNOLOGY, INC. RGS18 (F-5): sc-390908 BACKGROUND APPLICATIONS The regulators of G protein signaling (RGS) proteins inhibit heterotrimeric G RGS18 (F-5) is recommended for detection of RGS18 of human origin by protein signaling. RGS proteins work by functioning as GTPase-activating Western Blotting (starting dilution 1:100, dilution range 1:100-1:1000), proteins (which increase the GTPase activity of G protein a subunits) thereby immunoprecipitation [1-2 µg per 100-500 µg of total protein (1 ml of cell driving G proteins into their inactive GDP-bound form. RGS18 is a 234 amino lysate)], immunofluorescence (starting dilution 1:50, dilution range 1:50- acid peptide expressed mainly in megakaryocyte cells, but also in hemato- 1:500) and solid phase ELISA (starting dilution 1:30, dilution range 1:30- poietic progenitor and myeloerythroid lineage cells. RGS18 expression is 1:3000). upregulated during megakaryocyte differentiation and may play an important Suitable for use as control antibody for RGS18 siRNA (h): sc-61468, RGS18 role in the mediation of megakaryocyte chemotaxis. Structurally, RGS18 con- shRNA Plasmid (h): sc-61468-SH and RGS18 shRNA (h) Lentiviral Particles: tains phosphorylation sites for casein kinase II, protein kinase C and protein sc-61468-V. kinase A. RGS18 specifically binds to two a subunits of the G protein, Ga i and Ga q. Molecular Weight of RGS18: 26 kDa. Positive Controls: human RGS18 transfected HEK293T whole cell lysate. REFERENCES 1. Yowe, D., et al. 2001. RGS18 is a myeloerythroid lineage-specific regula- RECOMMENDED SUPPORT REAGENTS tor of molecule highly expressed in megakaryocytes. -

Integrated Genetic Diagnosis of Neurofibromatosis Type 1 (NF1) and Molecular Characterization of One Case of Compound Heterozygosity
Integrated genetic diagnosis of Neurofibromatosis type 1 (NF1) and molecular characterization of one case of compound heterozygosity Coordinator: Prof. Andrea Biondi Tutor: Dr. Gaetano Finocchiaro Dr. Sara MOROSINI Matr. No. 745001 XXVI CYCLE ACADEMIC YEAR 2012-2013 2 A Fabio e Matteo 3 4 TABLE of CONTENTS CHAPTER 1 7 INTRODUCTION 7 Neurofibromatoses 7 Neurofibromatosis type 1 8 Genotype/Phenotype correlation 24 NF1 gene 26 NF1 Protein 30 SCOPE OF THE THESIS 34 REFERENCES 36 CHAPTER 2 49 A TEN YEAR OF ITALIAN EXPERIENCE IN NEUROFIBROMATOSIS TYPE 1: 102 NOVEL MUTATIONS 49 CHAPTER 3 94 INTEGRATED GENETIC STUDIES OF NEUROFIBROMATOSIS TYPE 1. 94 CHAPTER 4 113 A FAMILY CASE REPORT 113 CHAPTER 5 138 SUMMARY 138 5 CONCLUSION 140 FUTURE PERSPECTIVES 142 ACKNOWLEDGEMENTS 146 6 Chapter 1 INTRODUCTION Neurofibromatoses Historically the neurofibromatosis type 1 (NF1), type 2 (NF2) and schwannomatosis were referred with the common terms of Neurofibromatosis. Neurofibromatoses are a group of conditions that predispose to tumors of the nervous system and abnormal skin pigmentation. Each type is defined by the presence/absence of café au lait (CAL) spots and skinfold freckling, what kind of peripheral nerve tumor develops (neurofibromas vs. schwannomas) and other features, particularly in the eye, specific to each form. The neurofibromatoses, predisposing to multiple tumors of the peripheral nervous system, are often considered classical tumor suppressor diseases. Longitudinal care for individuals with neurofibromatosis aims at the early detection and symptomatic treatment of complications as they occur 1. Very much has been elucidated about the complex molecular mechanism leading to these diseases. In the last two decades of the last century, the modern methods of genetic research showed that NF1 and NF2 had distinct clinical and genetic features. -
Expression Analysis and Functional Characterization of the Dro1/Cl2 Gene During Zebrafish Somitogenesis
See discussions, stats, and author profiles for this publication at: https://www.researchgate.net/publication/271208226 Expression analysis and functional characterization of the dro1/Cl2 gene during zebrafish somitogenesis Conference Paper · July 2009 DOI: 10.13140/2.1.2295.8407 CITATIONS READS 0 159 11 authors, including: Isabella Della Noce Elena Turola Parco Tecnologico Padano Università degli Studi di Modena e Reggio E… 16 PUBLICATIONS 19 CITATIONS 36 PUBLICATIONS 705 CITATIONS SEE PROFILE SEE PROFILE Silvia Carra Rosina Critelli I.R.C.C.S. Istituto Auxologico Italiano Università degli Studi di Modena e Reggio E… 38 PUBLICATIONS 132 CITATIONS 30 PUBLICATIONS 470 CITATIONS SEE PROFILE SEE PROFILE Some of the authors of this publication are also working on these related projects: Developmental Biology View project Mitogenome analysis of Sparidae species View project All content following this page was uploaded by Carlo De Lorenzo on 23 January 2015. The user has requested enhancement of the downloaded file. 6th European Zebrafish Genetics and Development Meeting Rome 15th - 19th July 2009 Program & Abstracts *UKVTUDMJDLFE DMJDLTBXBZGSPNQVSF%/" 8IZTBZEFPYZSJCPOVDMFJDBDJEXIFO%/"EPFTUIFKPC 8IZXPSLNJOVUFTPO%/"FYUSBDUJPOXIFOZPVDBOEPJUJO 8IZTQFOENPSFUIBOEPVCMFPOTPNFUIJOHMFTTUIBO(FOF.PMF (FOF.PMFJTBOJEFBMTZTUFNGPSBVUPNBUFE%/"BOE3/"FYUSBDUJPO 4JNQMFUPCVZ TJNQMFUPVTF BOETJNQMFUPNBJOUBJO (FOF.PMFJTWFSJ¾FEGPS[FCSB¾TITBNQMFTCZXPSMEMFBEJOHSFTFBSDIFST 6th European Zebrafish Genetics and Development Meeting Rome 15th - 19th July 2009 Program -

Receptor Signaling Through Osteoclast-Associated Monocyte
Downloaded from http://www.jimmunol.org/ by guest on September 29, 2021 is online at: average * The Journal of Immunology The Journal of Immunology , 20 of which you can access for free at: 2015; 194:3169-3179; Prepublished online 27 from submission to initial decision 4 weeks from acceptance to publication February 2015; doi: 10.4049/jimmunol.1402800 http://www.jimmunol.org/content/194/7/3169 Collagen Induces Maturation of Human Monocyte-Derived Dendritic Cells by Signaling through Osteoclast-Associated Receptor Heidi S. Schultz, Louise M. Nitze, Louise H. Zeuthen, Pernille Keller, Albrecht Gruhler, Jesper Pass, Jianhe Chen, Li Guo, Andrew J. Fleetwood, John A. Hamilton, Martin W. Berchtold and Svetlana Panina J Immunol cites 43 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Author Choice option Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Freely available online through http://www.jimmunol.org/content/suppl/2015/02/27/jimmunol.140280 0.DCSupplemental This article http://www.jimmunol.org/content/194/7/3169.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Author Choice Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2015 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. -

NF1) Liang Shang1†, Zhen Fang1†, Jin Liu2†, Fengying Du1†, Haiyan Jing3, Yali Xu3, Kangdi Dong1, Xiaoman Zhang4, Hao Wu1, Changqing Jing1,5* and Leping Li1,5*
Shang et al. BMC Cancer (2019) 19:1196 https://doi.org/10.1186/s12885-019-6375-9 CASE REPORT Open Access Case report of ascending colon cancer and multiple jejunal GISTs in a patient with neurofibromatosis type 1 (NF1) Liang Shang1†, Zhen Fang1†, Jin Liu2†, Fengying Du1†, Haiyan Jing3, Yali Xu3, Kangdi Dong1, Xiaoman Zhang4, Hao Wu1, Changqing Jing1,5* and Leping Li1,5* Abstract Background: NF1(Neurofibromatosis type 1) is an autosomal dominant genetic disorder. Patients with NF1 have an increased risk of developing benign or malignant tumours, such as gastrointestinal stromal tumours (GISTs). However, the coexistence of NF1, GIST and colon cancer is very rare, and few cases have been reported in the literature. Case presentation: We admitted a case of a 64-year-old man with type 1 neurofibromatosis, GISTs, and ascending colon cancer. This case was characterized by café-au-lait macules, discrete cutaneous neurofibromas, nodular neurofibromas, multiple jejunal tumours, and ascending colon cancer. Laparoscopic exploration revealed ascending colon cancer and multiple jejunal tumours. Laparoscopic right hemicolectomy and local excision of the jejunal tumours were performed successfully. The pathological results confirmed moderate differentiated adenocarcinoma of the ascending colon with multiple jejunal GISTs (low risk, very low risk). Moreover, the immunohistochemistry results of multiple jejunal GISTs suggest that NF1 is positive. Whole-exome sequencing (WES) of colon cancer revealed mutations in more than 20 genes, including KRAS, PIK3CA, APC, SMAD4, etc. The results of whole-exome sequencing (WES) of jejunal GISTs revealed an NF1 mutation and no KIT or PDGFR gene mutation. Conclusion: We report a rare case of simultaneous NF1, GIST and colon adenocarcinoma.