Comparative Transcriptome Profiling of Human and Pig Intestinal Epithelial Cells After
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Predicting Hosts Based on Early SARS-Cov-2 Samples And
www.nature.com/scientificreports OPEN Predicting hosts based on early SARS‑CoV‑2 samples and analyzing the 2020 pandemic Qian Guo1,2,3,7, Mo Li4,7, Chunhui Wang4,7, Jinyuan Guo1,3,7, Xiaoqing Jiang1,2,5,7, Jie Tan1, Shufang Wu1,2, Peihong Wang1, Tingting Xiao6, Man Zhou1,2, Zhencheng Fang1,2, Yonghong Xiao6* & Huaiqiu Zhu1,2,3,5* The SARS‑CoV‑2 pandemic has raised concerns in the identifcation of the hosts of the virus since the early stages of the outbreak. To address this problem, we proposed a deep learning method, DeepHoF, based on extracting viral genomic features automatically, to predict the host likelihood scores on fve host types, including plant, germ, invertebrate, non‑human vertebrate and human, for novel viruses. DeepHoF made up for the lack of an accurate tool, reaching a satisfactory AUC of 0.975 in the fve‑ classifcation, and could make a reliable prediction for the novel viruses without close neighbors in phylogeny. Additionally, to fll the gap in the efcient inference of host species for SARS‑CoV‑2 using existing tools, we conducted a deep analysis on the host likelihood profle calculated by DeepHoF. Using the isolates sequenced in the earliest stage of the COVID‑19 pandemic, we inferred that minks, bats, dogs and cats were potential hosts of SARS‑CoV‑2, while minks might be one of the most noteworthy hosts. Several genes of SARS‑CoV‑2 demonstrated their signifcance in determining the host range. Furthermore, a large‑scale genome analysis, based on DeepHoF’s computation for the later pandemic in 2020, disclosed the uniformity of host range among SARS‑CoV‑2 samples and the strong association of SARS‑CoV‑2 between humans and minks. -
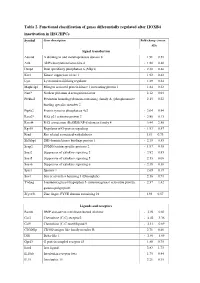
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Genome Organization of Canada Goose Coronavirus, a Novel
www.nature.com/scientificreports OPEN Genome Organization of Canada Goose Coronavirus, A Novel Species Identifed in a Mass Die-of of Received: 14 January 2019 Accepted: 25 March 2019 Canada Geese Published: xx xx xxxx Amber Papineau1,2, Yohannes Berhane1, Todd N. Wylie3,4, Kristine M. Wylie3,4, Samuel Sharpe5 & Oliver Lung 1,2 The complete genome of a novel coronavirus was sequenced directly from the cloacal swab of a Canada goose that perished in a die-of of Canada and Snow geese in Cambridge Bay, Nunavut, Canada. Comparative genomics and phylogenetic analysis indicate it is a new species of Gammacoronavirus, as it falls below the threshold of 90% amino acid similarity in the protein domains used to demarcate Coronaviridae. Additional features that distinguish the genome of Canada goose coronavirus include 6 novel ORFs, a partial duplication of the 4 gene and a presumptive change in the proteolytic processing of polyproteins 1a and 1ab. Viruses belonging to the Coronaviridae family have a single stranded positive sense RNA genome of 26–31 kb. Members of this family include both human pathogens, such as severe acute respiratory syn- drome virus (SARS-CoV)1, and animal pathogens, such as porcine epidemic diarrhea virus2. Currently, the International Committee on the Taxonomy of Viruses (ICTV) recognizes four genera in the Coronaviridae family: Alphacoronavirus, Betacoronavirus, Gammacoronavirus and Deltacoronavirus. While the reser- voirs of the Alphacoronavirus and Betacoronavirus genera are believed to be bats, the Gammacoronavirus and Deltacoronavirus genera have been shown to spread primarily through birds3. Te frst three species of the Deltacoronavirus genus were discovered in 20094 and recent work has vastly expanded the Deltacoronavirus genus, adding seven additional species3. -

Broad Receptor Engagement of an Emerging Global Coronavirus May Potentiate Its Diverse Cross-Species Transmissibility
Broad receptor engagement of an emerging global coronavirus may potentiate its diverse cross-species transmissibility Wentao Lia,1, Ruben J. G. Hulswita,1, Scott P. Kenneyb,1, Ivy Widjajaa, Kwonil Jungb, Moyasar A. Alhamob, Brenda van Dierena, Frank J. M. van Kuppevelda, Linda J. Saifb,2, and Berend-Jan Boscha,2 aVirology Division, Department of Infectious Diseases & Immunology, Faculty of Veterinary Medicine, Utrecht University, 3584 CL Utrecht, The Netherlands; and bDepartment of Veterinary Preventive Medicine, Food Animal Health Research Program, Ohio Agricultural Research and Development Center, The Ohio State University, Wooster, OH 44691 Contributed by Linda J. Saif, April 12, 2018 (sent for review February 15, 2018; reviewed by Tom Gallagher and Stefan Pöhlmann) Porcine deltacoronavirus (PDCoV), identified in 2012, is a common greatly increase the potential for successful adaptation to a new enteropathogen of swine with worldwide distribution. The source host (6, 12). A pivotal criterion of cross-species transmission and evolutionary history of this virus is, however, unknown. concerns the ability of a virus to engage a receptor within the PDCoV belongs to the Deltacoronavirus genus that comprises pre- novel host, which for CoVs, is determined by the receptor dominantly avian CoV. Phylogenetic analysis suggests that PDCoV specificity of the viral spike (S) entry protein. originated relatively recently from a host-switching event be- The porcine deltacoronavirus (PDCoV) is a recently discov- tween birds and mammals. Insight into receptor engagement by ered CoV of unknown origin. PDCoV (species name coronavirus PDCoV may shed light into such an exceptional phenomenon. Here HKU15) was identified in Hong Kong in pigs in the late 2000s we report that PDCoV employs host aminopeptidase N (APN) as an (13) and has since been detected in swine populations in various entry receptor and interacts with APN via domain B of its spike (S) countries worldwide (14–24). -

Global H3k4me3 Genome Mapping Reveals Alterations of Innate Immunity Signaling and Overexpression of JMJD3 in Human Myelodysplastic Syndrome CD34 Þ Cells
Leukemia (2013) 27, 2177–2186 & 2013 Macmillan Publishers Limited All rights reserved 0887-6924/13 www.nature.com/leu ORIGINAL ARTICLE Global H3K4me3 genome mapping reveals alterations of innate immunity signaling and overexpression of JMJD3 in human myelodysplastic syndrome CD34 þ cells YWei1, R Chen2, S Dimicoli1, C Bueso-Ramos3, D Neuberg4, S Pierce1, H Wang2, H Yang1, Y Jia1, H Zheng1, Z Fang1, M Nguyen3, I Ganan-Gomez1,5, B Ebert6, R Levine7, H Kantarjian1 and G Garcia-Manero1 The molecular bases of myelodysplastic syndromes (MDS) are not fully understood. Trimethylated histone 3 lysine 4 (H3K4me3) is present in promoters of actively transcribed genes and has been shown to be involved in hematopoietic differentiation. We performed a genome-wide H3K4me3 CHIP-Seq (chromatin immunoprecipitation coupled with whole genome sequencing) analysis of primary MDS bone marrow (BM) CD34 þ cells. This resulted in the identification of 36 genes marked by distinct higher levels of promoter H3K4me3 in MDS. A majority of these genes are involved in nuclear factor (NF)-kB activation and innate immunity signaling. We then analyzed expression of histone demethylases and observed significant overexpression of the JmjC-domain histone demethylase JMJD3 (KDM6b) in MDS CD34 þ cells. Furthermore, we demonstrate that JMJD3 has a positive effect on transcription of multiple CHIP-Seq identified genes involved in NF-kB activation. Inhibition of JMJD3 using shRNA in primary BM MDS CD34 þ cells resulted in an increased number of erythroid colonies in samples isolated from patients with lower-risk MDS. Taken together, these data indicate the deregulation of H3K4me3 and associated abnormal activation of innate immunity signals have a role in the pathogenesis of MDS and that targeting these signals may have potential therapeutic value in MDS. -

On the Coronaviruses and Their Associations with the Aquatic Environment and Wastewater
water Review On the Coronaviruses and Their Associations with the Aquatic Environment and Wastewater Adrian Wartecki 1 and Piotr Rzymski 2,* 1 Faculty of Medicine, Poznan University of Medical Sciences, 60-812 Pozna´n,Poland; [email protected] 2 Department of Environmental Medicine, Poznan University of Medical Sciences, 60-806 Pozna´n,Poland * Correspondence: [email protected] Received: 24 April 2020; Accepted: 2 June 2020; Published: 4 June 2020 Abstract: The outbreak of Coronavirus Disease 2019 (COVID-19), a severe respiratory disease caused by betacoronavirus SARS-CoV-2, in 2019 that further developed into a pandemic has received an unprecedented response from the scientific community and sparked a general research interest into the biology and ecology of Coronaviridae, a family of positive-sense single-stranded RNA viruses. Aquatic environments, lakes, rivers and ponds, are important habitats for bats and birds, which are hosts for various coronavirus species and strains and which shed viral particles in their feces. It is therefore of high interest to fully explore the role that aquatic environments may play in coronavirus spread, including cross-species transmissions. Besides the respiratory tract, coronaviruses pathogenic to humans can also infect the digestive system and be subsequently defecated. Considering this, it is pivotal to understand whether wastewater can play a role in their dissemination, particularly in areas with poor sanitation. This review provides an overview of the taxonomy, molecular biology, natural reservoirs and pathogenicity of coronaviruses; outlines their potential to survive in aquatic environments and wastewater; and demonstrates their association with aquatic biota, mainly waterfowl. It also calls for further, interdisciplinary research in the field of aquatic virology to explore the potential hotspots of coronaviruses in the aquatic environment and the routes through which they may enter it. -

Hydroxychloroquine Or Chloroquine for Treating Coronavirus Disease 2019 (COVID-19) – a PROTOCOL for a Systematic Review of Individual Participant Data
Hydroxychloroquine or Chloroquine for treating Coronavirus Disease 2019 (COVID-19) – a PROTOCOL for a systematic review of Individual Participant Data Authors Fontes LE, Riera R, Miranda E, Oke J, Heneghan CJ, Aronson JK, Pacheco RL, Martimbianco ALC, Nunan D BACKGROUND In the face of the pandemic of SARS CoV2, urgent research is needed to test potential therapeutic agents against the disease. Reliable research shall inform clinical decision makers. Currently, there are several studies testing the efficacy and safety profiles of different pharmacological interventions. Among these drugs, we can cite antimalarial, antivirals, biological drugs, interferon, etc. As of 6 April 2020 there are three published reportsand 100 ongoing trials testing hydroxychloroquine/chloroquine alone or in association with other drugs for COVID-19. This prospective systematic review with Individual Participant data aims to assess the rigour of the best-available evidence for hydroxychloroquine or chloroquine as treatment for COVID-19 infection. The PICO framework is: P: adults with COVID-19 infection I: chloroquine or hydroxychloroquine (alone or in association) C: placebo, other active treatments, usual standard care without antimalarials O: efficacy and safety outcomes OBJECTIVES To assess the effects (benefits and harms) of chloroquine or hydroxychloroquine for the treatment of COVID-19 infection. METHODS Criteria for considering studies for this review Types of studies We shall include randomized controlled trials (RCTs) with a parallel design. We intend to include even small trials (<50 participants), facing the urgent need for evidence to respond to the current pandemic. Quasi-randomized, non-randomized, or observational studies will be excluded due to a higher risk of confounding and selection bias (1). -

Identification of Key Pathways and Genes in Dementia Via Integrated Bioinformatics Analysis
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.18.440371; this version posted July 19, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Identification of Key Pathways and Genes in Dementia via Integrated Bioinformatics Analysis Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karnataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2021.04.18.440371; this version posted July 19, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract To provide a better understanding of dementia at the molecular level, this study aimed to identify the genes and key pathways associated with dementia by using integrated bioinformatics analysis. Based on the expression profiling by high throughput sequencing dataset GSE153960 derived from the Gene Expression Omnibus (GEO), the differentially expressed genes (DEGs) between patients with dementia and healthy controls were identified. With DEGs, we performed a series of functional enrichment analyses. Then, a protein–protein interaction (PPI) network, modules, miRNA-hub gene regulatory network and TF-hub gene regulatory network was constructed, analyzed and visualized, with which the hub genes miRNAs and TFs nodes were screened out. Finally, validation of hub genes was performed by using receiver operating characteristic curve (ROC) analysis. -

Genome Sequencing Unveils a Regulatory Landscape of Platelet Reactivity
ARTICLE https://doi.org/10.1038/s41467-021-23470-9 OPEN Genome sequencing unveils a regulatory landscape of platelet reactivity Ali R. Keramati 1,2,124, Ming-Huei Chen3,4,124, Benjamin A. T. Rodriguez3,4,5,124, Lisa R. Yanek 2,6, Arunoday Bhan7, Brady J. Gaynor 8,9, Kathleen Ryan8,9, Jennifer A. Brody 10, Xue Zhong11, Qiang Wei12, NHLBI Trans-Omics for Precision (TOPMed) Consortium*, Kai Kammers13, Kanika Kanchan14, Kruthika Iyer14, Madeline H. Kowalski15, Achilleas N. Pitsillides4,16, L. Adrienne Cupples 4,16, Bingshan Li 12, Thorsten M. Schlaeger7, Alan R. Shuldiner9, Jeffrey R. O’Connell8,9, Ingo Ruczinski17, Braxton D. Mitchell 8,9, ✉ Nauder Faraday2,18, Margaret A. Taub17, Lewis C. Becker1,2, Joshua P. Lewis 8,9,125 , 2,14,125✉ 3,4,125✉ 1234567890():,; Rasika A. Mathias & Andrew D. Johnson Platelet aggregation at the site of atherosclerotic vascular injury is the underlying patho- physiology of myocardial infarction and stroke. To build upon prior GWAS, here we report on 16 loci identified through a whole genome sequencing (WGS) approach in 3,855 NHLBI Trans-Omics for Precision Medicine (TOPMed) participants deeply phenotyped for platelet aggregation. We identify the RGS18 locus, which encodes a myeloerythroid lineage-specific regulator of G-protein signaling that co-localizes with expression quantitative trait loci (eQTL) signatures for RGS18 expression in platelets. Gene-based approaches implicate the SVEP1 gene, a known contributor of coronary artery disease risk. Sentinel variants at RGS18 and PEAR1 are associated with thrombosis risk and increased gastrointestinal bleeding risk, respectively. Our WGS findings add to previously identified GWAS loci, provide insights regarding the mechanism(s) by which genetics may influence cardiovascular disease risk, and underscore the importance of rare variant and regulatory approaches to identifying loci contributing to complex phenotypes. -

Untwining Anti-Tumor and Immunosuppressive Effects of JAK Inhibitors—A Strategy for Hematological Malignancies?
cancers Review Untwining Anti-Tumor and Immunosuppressive Effects of JAK Inhibitors—A Strategy for Hematological Malignancies? Klara Klein 1, Dagmar Stoiber 2, Veronika Sexl 1 and Agnieszka Witalisz-Siepracka 1,2,* 1 Department of Biomedical Science, Institute of Pharmacology and Toxicology, University of Veterinary Medicine Vienna, 1210 Vienna, Austria; [email protected] (K.K.); [email protected] (V.S.) 2 Department of Pharmacology, Physiology and Microbiology, Division Pharmacology, Karl Landsteiner University of Health Sciences, 3500 Krems, Austria; [email protected] * Correspondence: [email protected] or [email protected] Simple Summary: The Janus kinase-signal transducer and activator of transcription (JAK-STAT) pathway is aberrantly activated in many malignancies. Inhibition of this pathway via JAK inhibitors (JAKinibs) is therefore an attractive therapeutic strategy underlined by Ruxolitinib (JAK1/2 inhibitor) being approved for the treatment of myeloproliferative neoplasms. As a consequence of the crucial role of the JAK-STAT pathway in the regulation of immune responses, inhibition of JAKs suppresses the immune system. This review article provides a thorough overview of the current knowledge on JAKinibs’ effects on immune cells in the context of hematological malignancies. We also discuss the potential use of JAKinibs for the treatment of diseases in which lymphocytes are the source of the malignancy. Citation: Klein, K.; Stoiber, D.; Sexl, Abstract: The Janus kinase-signal transducer and activator of transcription (JAK-STAT) pathway V.; Witalisz-Siepracka, A. Untwining propagates signals from a variety of cytokines, contributing to cellular responses in health and disease. Anti-Tumor and Immunosuppressive Gain of function mutations in JAKs or STATs are associated with malignancies, with JAK2V617F being Effects of JAK Inhibitors—A Strategy the main driver mutation in myeloproliferative neoplasms (MPN). -

Complete Sections As Applicable
This form should be used for all taxonomic proposals. Please complete all those modules that are applicable (and then delete the unwanted sections). For guidance, see the notes written in blue and the separate document “Help with completing a taxonomic proposal” Please try to keep related proposals within a single document; you can copy the modules to create more than one genus within a new family, for example. MODULE 1: TITLE, AUTHORS, etc (to be completed by ICTV Code assigned: 2010.023a-dV officers) Short title: A new genus and three new species in the subfamily Coronavirinae (e.g. 6 new species in the genus Zetavirus) Modules attached 1 2 3 4 5 (modules 1 and 9 are required) 6 7 8 9 Author(s) with e-mail address(es) of the proposer: Raoul J. de Groot and Alexander E. Gorbalenya List the ICTV study group(s) that have seen this proposal: A list of study groups and contacts is provided at http://www.ictvonline.org/subcommittees.asp . If Submission on behalf of the Coronavirus Study in doubt, contact the appropriate subcommittee Group. Authors are CSG members (R.J de chair (fungal, invertebrate, plant, prokaryote or Groot, Chair) vertebrate viruses) ICTV-EC or Study Group comments and response of the proposer: Date first submitted to ICTV: Date of this revision (if different to above): Page 1 of 5 MODULE 2: NEW SPECIES Code 2010.023aV (assigned by ICTV officers) To create new species within: Genus: Deltacoronavirus (new) Subfamily: Coronavirinae Family: Coronaviridae Order: Nidovirales And name the new species: GenBank sequence accession number(s) of reference isolate: Bulbul coronavirus HKU11 [FJ376619] Thrush coronavirus HKU12 [FJ376621=NC_011549] Munia coronavirus HKU13 [FJ376622=NC_011550] Reasons to justify the creation and assignment of the new species: According to the demarcation criteria as outlined in Module 3 and agreed upon by the Coronavirus Study Group, the new coronaviruses isolated from Bulbul, Thrush and Munia are representatives of separates species. -

Coronaviruses in Avian Species – Review with Focus on Epidemiology and Diagnosis in Wild Birds
J Vet Res 62, 249-255, 2018 DOI:10.2478/jvetres-2018-0035 REVIEW ARTICLE Coronaviruses in avian species – review with focus on epidemiology and diagnosis in wild birds Justyna Miłek, Katarzyna Blicharz-Domańska Department of Poultry Diseases, National Veterinary Research Institute, 24-100 Puławy, Poland [email protected] Received: May 2, 2018 Accepted: September 19, 2018 Abstract Coronaviruses (CoVs) are a large group of enveloped viruses with a single-strand RNA genome, which continuously circulate in mammals and birds and pose a threat to livestock, companion animals, and humans. CoVs harboured by avian species are classified to the genera gamma- and deltacoronaviruses. Within the gamma-CoVs the main representative is avian coronavirus, a taxonomic name which includes the highly contagious infectious bronchitis viruses (IBVs) in chickens and similar viruses infecting other domestic birds such as turkeys, guinea fowls, or quails. Additionally, IBVs have been detected in healthy wild birds, demonstrating that they may act as the vector between domestic and free-living birds. Moreover, CoVs other than IBVs, are identified in wild birds, which suggests that wild birds play a key role in the epidemiology of other gammaCoVs and deltaCoVs. Development of molecular techniques has significantly improved knowledge of the prevalence of CoVs in avian species. The methods adopted in monitoring studies of CoVs in different avian species are mainly based on detection of conservative regions within the viral replicase, nucleocapsid genes, and 3’UTR or 5’UTR. The purpose of this review is to summarise recent discoveries in the areas of epidemiology and diagnosis of CoVs in avian species and to understand the role of wild birds in the virus distribution.