Biological Role and Disease Impact of Copy Number Variation in Complex Disease
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

PARSANA-DISSERTATION-2020.Pdf
DECIPHERING TRANSCRIPTIONAL PATTERNS OF GENE REGULATION: A COMPUTATIONAL APPROACH by Princy Parsana A dissertation submitted to The Johns Hopkins University in conformity with the requirements for the degree of Doctor of Philosophy Baltimore, Maryland July, 2020 © 2020 Princy Parsana All rights reserved Abstract With rapid advancements in sequencing technology, we now have the ability to sequence the entire human genome, and to quantify expression of tens of thousands of genes from hundreds of individuals. This provides an extraordinary opportunity to learn phenotype relevant genomic patterns that can improve our understanding of molecular and cellular processes underlying a trait. The high dimensional nature of genomic data presents a range of computational and statistical challenges. This dissertation presents a compilation of projects that were driven by the motivation to efficiently capture gene regulatory patterns in the human transcriptome, while addressing statistical and computational challenges that accompany this data. We attempt to address two major difficulties in this domain: a) artifacts and noise in transcriptomic data, andb) limited statistical power. First, we present our work on investigating the effect of artifactual variation in gene expression data and its impact on trans-eQTL discovery. Here we performed an in-depth analysis of diverse pre-recorded covariates and latent confounders to understand their contribution to heterogeneity in gene expression measurements. Next, we discovered 673 trans-eQTLs across 16 human tissues using v6 data from the Genotype Tissue Expression (GTEx) project. Finally, we characterized two trait-associated trans-eQTLs; one in Skeletal Muscle and another in Thyroid. Second, we present a principal component based residualization method to correct gene expression measurements prior to reconstruction of co-expression networks. -
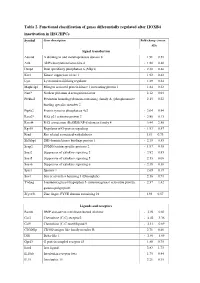
Table 2. Functional Classification of Genes Differentially Regulated After HOXB4 Inactivation in HSC/Hpcs
Table 2. Functional classification of genes differentially regulated after HOXB4 inactivation in HSC/HPCs Symbol Gene description Fold-change (mean ± SD) Signal transduction Adam8 A disintegrin and metalloprotease domain 8 1.91 ± 0.51 Arl4 ADP-ribosylation factor-like 4 - 1.80 ± 0.40 Dusp6 Dual specificity phosphatase 6 (Mkp3) - 2.30 ± 0.46 Ksr1 Kinase suppressor of ras 1 1.92 ± 0.42 Lyst Lysosomal trafficking regulator 1.89 ± 0.34 Mapk1ip1 Mitogen activated protein kinase 1 interacting protein 1 1.84 ± 0.22 Narf* Nuclear prelamin A recognition factor 2.12 ± 0.04 Plekha2 Pleckstrin homology domain-containing. family A. (phosphoinosite 2.15 ± 0.22 binding specific) member 2 Ptp4a2 Protein tyrosine phosphatase 4a2 - 2.04 ± 0.94 Rasa2* RAS p21 activator protein 2 - 2.80 ± 0.13 Rassf4 RAS association (RalGDS/AF-6) domain family 4 3.44 ± 2.56 Rgs18 Regulator of G-protein signaling - 1.93 ± 0.57 Rrad Ras-related associated with diabetes 1.81 ± 0.73 Sh3kbp1 SH3 domain kinase bindings protein 1 - 2.19 ± 0.53 Senp2 SUMO/sentrin specific protease 2 - 1.97 ± 0.49 Socs2 Suppressor of cytokine signaling 2 - 2.82 ± 0.85 Socs5 Suppressor of cytokine signaling 5 2.13 ± 0.08 Socs6 Suppressor of cytokine signaling 6 - 2.18 ± 0.38 Spry1 Sprouty 1 - 2.69 ± 0.19 Sos1 Son of sevenless homolog 1 (Drosophila) 2.16 ± 0.71 Ywhag 3-monooxygenase/tryptophan 5- monooxygenase activation protein. - 2.37 ± 1.42 gamma polypeptide Zfyve21 Zinc finger. FYVE domain containing 21 1.93 ± 0.57 Ligands and receptors Bambi BMP and activin membrane-bound inhibitor - 2.94 ± 0.62 -

Identification of the Binding Partners for Hspb2 and Cryab Reveals
Brigham Young University BYU ScholarsArchive Theses and Dissertations 2013-12-12 Identification of the Binding arP tners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non- Redundant Roles for Small Heat Shock Proteins Kelsey Murphey Langston Brigham Young University - Provo Follow this and additional works at: https://scholarsarchive.byu.edu/etd Part of the Microbiology Commons BYU ScholarsArchive Citation Langston, Kelsey Murphey, "Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non-Redundant Roles for Small Heat Shock Proteins" (2013). Theses and Dissertations. 3822. https://scholarsarchive.byu.edu/etd/3822 This Thesis is brought to you for free and open access by BYU ScholarsArchive. It has been accepted for inclusion in Theses and Dissertations by an authorized administrator of BYU ScholarsArchive. For more information, please contact [email protected], [email protected]. Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactions and Non-Redundant Roles for Small Heat Shock Proteins Kelsey Langston A thesis submitted to the faculty of Brigham Young University in partial fulfillment of the requirements for the degree of Master of Science Julianne H. Grose, Chair William R. McCleary Brian Poole Department of Microbiology and Molecular Biology Brigham Young University December 2013 Copyright © 2013 Kelsey Langston All Rights Reserved ABSTRACT Identification of the Binding Partners for HspB2 and CryAB Reveals Myofibril and Mitochondrial Protein Interactors and Non-Redundant Roles for Small Heat Shock Proteins Kelsey Langston Department of Microbiology and Molecular Biology, BYU Master of Science Small Heat Shock Proteins (sHSP) are molecular chaperones that play protective roles in cell survival and have been shown to possess chaperone activity. -

Genetic Variants in Humanin Nuclear Isoform Gene Regions Show No Association with Coronary Artery Disease
Genetic variants in humanin nuclear isoform gene regions show no association with coronary artery disease Mall Eltermaa ( [email protected] ) University of Tartu https://orcid.org/0000-0002-9651-4107 Maili Jakobson University of Tartu Meeme Utt University of Tartu Sulev Kõks Perron Institute for Neurological and Translational Science Reedik Mägi University of Tartu Joel Starkopf Tartu University Hospital Research article Keywords: humanin, humanin-like, coronary artery disease, association study, peptide Posted Date: June 25th, 2019 DOI: https://doi.org/10.21203/rs.2.10582/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Version of Record: A version of this preprint was published at BMC Research Notes on November 21st, 2019. See the published version at https://doi.org/10.1186/s13104-019-4807-x. Loading [MathJax]/jax/output/CommonHTML/jax.js Page 1/7 Abstract Background Coronary artery disease contributes to noncommunicable disease deaths worldwide. In order to make preventive methods more accurate, we need to know more about the development and progress of this pathology, including the genetic aspects. Humanin is a small peptide known for its cytoprotective and anti-apoptotic properties. Our study looked for genomic associations between humanin-like nuclear isoform genes and coronary artery disease using CARDIoGRAMplusC4D Consortium data. Results Lookup from meta-analysis datasets gave single nucleotide polymorphisms in all 13 humanin-like nuclear isoform genes with the lowest P-value for rs6151662 from the MTRNR2L2 gene including the 50 kb anking region in both directions (P-value = 0.0037). Within the gene region alone the top variant was rs78083998 from the MTRNR2L13 region (meta-analysis P-value=0.042). -

PGENETICS-D-11-00413 Title: Integrating
Editorial Manager(tm) for PLoS Genetics Manuscript Draft Manuscript Number: PGENETICS-D-11-00413 Title: Integrating genetic and gene expression evidence into genome-wide association analysis of gene sets Short Title: Integrative association analysis of gene sets Article Type: Research Article Section/Category: Natural Variation Keywords: gene set analysis; eQTL; integrative genomics Corresponding Author: Sayan Mukherjee Corresponding Author's Institution: Duke University First Author: Qing Xiong Order of Authors: Qing Xiong;Nicola Ancona;Elizabeth Hauser;Sayan Mukherjee;Terrence Furey Abstract: Background: Single variant or single gene analyses generally account for only a small proportion of the phenotypic variation in complex traits. Alternatively, gene set or pathway association analyses are playing an increasingly important role in uncovering genetic architectures of complex traits through the identification of systematic genetic interactions. Two dominant paradigms for gene set analyses are association analyses based on SNP genotypes and those based on gene expression profiles. However, gene-disease association can manifest in many ways such as alterations of gene expression, genotype and copy number, thus an integrative approach combining multiple forms of evidence can more accurately and comprehensively capture pathway associations. Methodology: We have developed a single statistical framework, Gene Set Association Analysis (GSAA), that simultaneously measures genome-wide patterns of genetic variation and gene expression variation -

Transcriptomic Characterization of Fibrolamellar Hepatocellular
Transcriptomic characterization of fibrolamellar PNAS PLUS hepatocellular carcinoma Elana P. Simona, Catherine A. Freijeb, Benjamin A. Farbera,c, Gadi Lalazara, David G. Darcya,c, Joshua N. Honeymana,c, Rachel Chiaroni-Clarkea, Brian D. Dilld, Henrik Molinad, Umesh K. Bhanote, Michael P. La Quagliac, Brad R. Rosenbergb,f, and Sanford M. Simona,1 aLaboratory of Cellular Biophysics, The Rockefeller University, New York, NY 10065; bPresidential Fellows Laboratory, The Rockefeller University, New York, NY 10065; cDivision of Pediatric Surgery, Department of Surgery, Memorial Sloan-Kettering Cancer Center, New York, NY 10065; dProteomics Resource Center, The Rockefeller University, New York, NY 10065; ePathology Core Facility, Memorial Sloan-Kettering Cancer Center, New York, NY 10065; and fJohn C. Whitehead Presidential Fellows Program, The Rockefeller University, New York, NY 10065 Edited by Susan S. Taylor, University of California, San Diego, La Jolla, CA, and approved September 22, 2015 (received for review December 29, 2014) Fibrolamellar hepatocellular carcinoma (FLHCC) tumors all carry a exon of DNAJB1 and all but the first exon of PRKACA. This deletion of ∼400 kb in chromosome 19, resulting in a fusion of the produced a chimeric RNA transcript and a translated chimeric genes for the heat shock protein, DNAJ (Hsp40) homolog, subfam- protein that retains the full catalytic activity of wild-type PKA. ily B, member 1, DNAJB1, and the catalytic subunit of protein ki- This chimeric protein was found in 15 of 15 FLHCC patients nase A, PRKACA. The resulting chimeric transcript produces a (21) in the absence of any other recurrent mutations in the DNA fusion protein that retains kinase activity. -

A Clinicopathological and Molecular Genetic Analysis of Low-Grade Glioma in Adults
A CLINICOPATHOLOGICAL AND MOLECULAR GENETIC ANALYSIS OF LOW-GRADE GLIOMA IN ADULTS Presented by ANUSHREE SINGH MSc A thesis submitted in partial fulfilment of the requirements of the University of Wolverhampton for the degree of Doctor of Philosophy Brain Tumour Research Centre Research Institute in Healthcare Sciences Faculty of Science and Engineering University of Wolverhampton November 2014 i DECLARATION This work or any part thereof has not previously been presented in any form to the University or to any other body whether for the purposes of assessment, publication or for any other purpose (unless otherwise indicated). Save for any express acknowledgments, references and/or bibliographies cited in the work, I confirm that the intellectual content of the work is the result of my own efforts and of no other person. The right of Anushree Singh to be identified as author of this work is asserted in accordance with ss.77 and 78 of the Copyright, Designs and Patents Act 1988. At this date copyright is owned by the author. Signature: Anushree Date: 30th November 2014 ii ABSTRACT The aim of the study was to identify molecular markers that can determine progression of low grade glioma. This was done using various approaches such as IDH1 and IDH2 mutation analysis, MGMT methylation analysis, copy number analysis using array comparative genomic hybridisation and identification of differentially expressed miRNAs using miRNA microarray analysis. IDH1 mutation was present at a frequency of 71% in low grade glioma and was identified as an independent marker for improved OS in a multivariate analysis, which confirms the previous findings in low grade glioma studies. -

Alters Enhancer Function in the Developing Telencephalon Meis1
Downloaded from genome.cshlp.org on March 19, 2014 - Published by Cold Spring Harbor Laboratory Press Restless Legs Syndrome-associated intronic common variant in Meis1 alters enhancer function in the developing telencephalon Derek Spieler, Maria Kaffe, Franziska Knauf, et al. Genome Res. published online March 18, 2014 Access the most recent version at doi:10.1101/gr.166751.113 Supplemental http://genome.cshlp.org/content/suppl/2014/02/10/gr.166751.113.DC1.html Material P<P Published online March 18, 2014 in advance of the print journal. Open Access Freely available online through the Genome Research Open Access option. Creative This article, published in Genome Research, is available under a Creative Commons Commons License (Attribution-NonCommercial 3.0 Unported), as described at License http://creativecommons.org/licenses/by-nc/3.0/. Email Alerting Receive free email alerts when new articles cite this article - sign up in the box at the Service top right corner of the article or click here. Advance online articles have been peer reviewed and accepted for publication but have not yet appeared in the paper journal (edited, typeset versions may be posted when available prior to final publication). Advance online articles are citable and establish publication priority; they are indexed by PubMed from initial publication. Citations to Advance online articles must include the digital object identifier (DOIs) and date of initial publication. To subscribe to Genome Research go to: http://genome.cshlp.org/subscriptions © 2014 Spieler et al.; Published by Cold Spring Harbor Laboratory Press Downloaded from genome.cshlp.org on March 19, 2014 - Published by Cold Spring Harbor Laboratory Press Research Restless Legs Syndrome-associated intronic common variant in Meis1 alters enhancer function in the developing telencephalon Derek Spieler,1,25 Maria Kaffe,1,2,25 Franziska Knauf,1,25 Jose´ Bessa,3 Juan J. -

A Misplaced Lncrna Causes Brachydactyly in Humans
A misplaced lncRNA causes brachydactyly in humans Philipp G. Maass, … , Friedrich C. Luft, Sylvia Bähring J Clin Invest. 2012;122(11):3990-4002. https://doi.org/10.1172/JCI65508. Research Article Translocations are chromosomal rearrangements that are frequently associated with a variety of disease states and developmental disorders. We identified 2 families with brachydactyly type E (BDE) resulting from different translocations affecting chromosome 12p. Both translocations caused downregulation of the parathyroid hormone-like hormone (PTHLH) gene by disrupting the cis-regulatory landscape. Using chromosome conformation capturing, we identified a regulator on chromosome 12q that interacts in cis with PTHLH over a 24.4-megabase distance and in trans with the sex- determining region Y-box 9 (SOX9) gene on chromosome 17q. The element also harbored a long noncoding RNA (lncRNA). Silencing of the lncRNA, PTHLH, or SOX9 revealed a feedback mechanism involving an expression-dependent network in humans. In the BDE patients, the human lncRNA was upregulated by the disrupted chromosomal association. Moreover, the lncRNA occupancy at the PTHLH locus was reduced. Our results document what we believe to be a novel in cis– and in trans–acting DNA and lncRNA regulatory feedback element that is reciprocally regulated by coding genes. Furthermore, our findings provide a systematic and combinatorial view of how enhancers encoding lncRNAs may affect gene expression in normal development. Find the latest version: https://jci.me/65508/pdf Research article Related Commentary, page 3837 A misplaced lncRNA causes brachydactyly in humans Philipp G. Maass,1,2 Andreas Rump,3 Herbert Schulz,2 Sigmar Stricker,4 Lisanne Schulze,1,2 Konrad Platzer,3 Atakan Aydin,1,2 Sigrid Tinschert,3 Mary B. -

Global H3k4me3 Genome Mapping Reveals Alterations of Innate Immunity Signaling and Overexpression of JMJD3 in Human Myelodysplastic Syndrome CD34 Þ Cells
Leukemia (2013) 27, 2177–2186 & 2013 Macmillan Publishers Limited All rights reserved 0887-6924/13 www.nature.com/leu ORIGINAL ARTICLE Global H3K4me3 genome mapping reveals alterations of innate immunity signaling and overexpression of JMJD3 in human myelodysplastic syndrome CD34 þ cells YWei1, R Chen2, S Dimicoli1, C Bueso-Ramos3, D Neuberg4, S Pierce1, H Wang2, H Yang1, Y Jia1, H Zheng1, Z Fang1, M Nguyen3, I Ganan-Gomez1,5, B Ebert6, R Levine7, H Kantarjian1 and G Garcia-Manero1 The molecular bases of myelodysplastic syndromes (MDS) are not fully understood. Trimethylated histone 3 lysine 4 (H3K4me3) is present in promoters of actively transcribed genes and has been shown to be involved in hematopoietic differentiation. We performed a genome-wide H3K4me3 CHIP-Seq (chromatin immunoprecipitation coupled with whole genome sequencing) analysis of primary MDS bone marrow (BM) CD34 þ cells. This resulted in the identification of 36 genes marked by distinct higher levels of promoter H3K4me3 in MDS. A majority of these genes are involved in nuclear factor (NF)-kB activation and innate immunity signaling. We then analyzed expression of histone demethylases and observed significant overexpression of the JmjC-domain histone demethylase JMJD3 (KDM6b) in MDS CD34 þ cells. Furthermore, we demonstrate that JMJD3 has a positive effect on transcription of multiple CHIP-Seq identified genes involved in NF-kB activation. Inhibition of JMJD3 using shRNA in primary BM MDS CD34 þ cells resulted in an increased number of erythroid colonies in samples isolated from patients with lower-risk MDS. Taken together, these data indicate the deregulation of H3K4me3 and associated abnormal activation of innate immunity signals have a role in the pathogenesis of MDS and that targeting these signals may have potential therapeutic value in MDS. -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

Noelia Díaz Blanco
Effects of environmental factors on the gonadal transcriptome of European sea bass (Dicentrarchus labrax), juvenile growth and sex ratios Noelia Díaz Blanco Ph.D. thesis 2014 Submitted in partial fulfillment of the requirements for the Ph.D. degree from the Universitat Pompeu Fabra (UPF). This work has been carried out at the Group of Biology of Reproduction (GBR), at the Department of Renewable Marine Resources of the Institute of Marine Sciences (ICM-CSIC). Thesis supervisor: Dr. Francesc Piferrer Professor d’Investigació Institut de Ciències del Mar (ICM-CSIC) i ii A mis padres A Xavi iii iv Acknowledgements This thesis has been made possible by the support of many people who in one way or another, many times unknowingly, gave me the strength to overcome this "long and winding road". First of all, I would like to thank my supervisor, Dr. Francesc Piferrer, for his patience, guidance and wise advice throughout all this Ph.D. experience. But above all, for the trust he placed on me almost seven years ago when he offered me the opportunity to be part of his team. Thanks also for teaching me how to question always everything, for sharing with me your enthusiasm for science and for giving me the opportunity of learning from you by participating in many projects, collaborations and scientific meetings. I am also thankful to my colleagues (former and present Group of Biology of Reproduction members) for your support and encouragement throughout this journey. To the “exGBRs”, thanks for helping me with my first steps into this world. Working as an undergrad with you Dr.