SPRED1 Germline Mutations Caused a Neurofibromatosis Type 1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Wt1) – an Effector in Leukemogenesis?
WILMS’ TUMOUR GENE 1 PROTEIN (WT1) – AN EFFECTOR IN LEUKEMOGENESIS? Vidovic, Karina 2010 Link to publication Citation for published version (APA): Vidovic, K. (2010). WILMS’ TUMOUR GENE 1 PROTEIN (WT1) – AN EFFECTOR IN LEUKEMOGENESIS?. Lund University: Faculty of Medicine. Total number of authors: 1 General rights Unless other specific re-use rights are stated the following general rights apply: Copyright and moral rights for the publications made accessible in the public portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. • Users may download and print one copy of any publication from the public portal for the purpose of private study or research. • You may not further distribute the material or use it for any profit-making activity or commercial gain • You may freely distribute the URL identifying the publication in the public portal Read more about Creative commons licenses: https://creativecommons.org/licenses/ Take down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. LUND UNIVERSITY PO Box 117 221 00 Lund +46 46-222 00 00 Download date: 07. Oct. 2021 Division of Hematology and Transfusion Medicine Lund University, Lund, Sweden WILMS’ TUMOUR GENE 1 PROTEIN (WT1) – AN EFFECTOR IN LEUKEMOGENESIS? Karina Vidovic Thesis 2010 Contact adress Karina Vidovic Division of Hematology and Transfusion Medicine BMC, C14 Klinikgatan 28 SE- 221 84 Lund Sweden Phone +46 46 222 07 30 e-mail: [email protected] ISBN 978-91-86443-99-3 ! Karina Vidovic Printed by Media-Tryck, Lund, Sweden 2 ”The journey of a thousand miles begins with one step” Lao Tzu (Chinese taoist), 600-531 BC 3 4 LIST OF PAPERS This thesis is based on the following papers, referred to in the text by their Roman numerals I. -

Piggybac Mutagenesis and Exome Sequencing Identify Genetic Driver
Noorani et al. Genome Biology (2020) 21:181 https://doi.org/10.1186/s13059-020-02092-2 RESEARCH Open Access PiggyBac mutagenesis and exome sequencing identify genetic driver landscapes and potential therapeutic targets of EGFR-mutant gliomas Imran Noorani1,2* , Jorge de la Rosa1†, Yoon Ha Choi1,3†, Alexander Strong1, Hannes Ponstingl1, M. S. Vijayabaskar1, Jusung Lee3, Eunmin Lee3, Angela Richard-Londt4, Mathias Friedrich1,5, Federica Furlanetto5, Rocio Fuente1, Ruby Banerjee1, Fengtang Yang1, Frances Law1, Colin Watts2,6, Roland Rad5, George Vassiliou1, Jong Kyoung Kim3, Thomas Santarius2, Sebastian Brandner4 and Allan Bradley1* * Correspondence: [email protected]; [email protected] Abstract †Jorge de la Rosa and Yoonha Choi contributed equally to this work. Background: Glioma is the most common intrinsic brain tumor and also occurs in 1The Wellcome Trust Sanger the spinal cord. Activating EGFR mutations are common in IDH1 wild-type gliomas. Institute, Wellcome Trust Genome However, the cooperative partners of EGFR driving gliomagenesis remain poorly Campus, Hinxton, Cambridgeshire CB10 1SA, UK understood. Full list of author information is Results: We explore EGFR-mutant glioma evolution in conditional mutant mice by available at the end of the article whole-exome sequencing, transposon mutagenesis forward genetic screening, and transcriptomics. We show mutant EGFR is sufficient to initiate gliomagenesis in vivo, both in the brain and spinal cord. We identify significantly recurrent somatic alterations in these gliomas including mutant EGFR amplifications and Sub1, Trp53, and Tead2 loss-of-function mutations. Comprehensive functional characterization of 96 gliomas by genome-wide piggyBac insertional mutagenesis in vivo identifies 281 known and novel EGFR-cooperating driver genes, including Cdkn2a, Nf1, Spred1, and Nav3. -

(NF1) Defects Are Common in Human Ovarian Serous Carcinomas and Co-Occur with TP53 Mutations
Volume 10 Number 12 December 2008 pp. 1362–1372 1362 www.neoplasia.com Neurofibromin 1 (NF1) Defects Navneet Sangha*, Rong Wu*, Rork Kuick†, Scott Powers‡, David Mu§, Diane Fiander*, Are Common in Human Ovarian Kit Yuen*, Hidetaka Katabuchi¶, Hironori Tashiro¶, Serous Carcinomas and Co-occur Eric R. Fearon*,†,# and Kathleen R. Cho*,†,# 1,2 with TP53 Mutations *Department of Pathology, The University of Michigan Medical School, Ann Arbor, MI 48109, USA; †The Comprehensive Cancer Center, The University of Michigan Medical School, Ann Arbor, MI 48109, USA; ‡Cold Spring Harbor Laboratory, Cold Spring Harbor, NY 11724, USA; §Department of Pathology, Penn State University College of Medicine, Hershey, PA 17033, USA; ¶Department of Gynecology, Kumamoto University, Kumamoto, 860-8556, Japan; #Department of Internal Medicine, The University of Michigan Medical School, Ann Arbor, MI 48109, USA Abstract Ovarian serous carcinoma (OSC) is the most common and lethal histologic type of ovarian epithelial malignancy. Mutations of TP53 and dysfunction of the Brca1 and/or Brca2 tumor-suppressor proteins have been implicated in the molecular pathogenesis of a large fraction of OSCs, but frequent somatic mutations in other well-established tumor-suppressor genes have not been identified. Using a genome-wide screen of DNA copy number alterations in 36 primary OSCs, we identified two tumors with apparent homozygous deletions of the NF1 gene. Subsequently, 18 ovarian carcinoma–derived cell lines and 41 primary OSCs were evaluated for NF1 alterations. Markedly re- duced or absent expression of Nf1 protein was observed in 6 of the 18 cell lines, and using the protein truncation test and sequencing of cDNA and genomic DNA, NF1 mutations resulting in deletion of exons and/or aberrant splicing of NF1 transcripts were detected in 5 of the 6 cell lines with loss of NF1 expression. -

Biomarker Testing in Non- Small Cell Lung Cancer (NSCLC)
The biopharma business of Merck KGaA, Darmstadt, Germany operates as EMD Serono in the U.S. and Canada. Biomarker testing in non- small cell lung cancer (NSCLC) Copyright © 2020 EMD Serono, Inc. All rights reserved. US/TEP/1119/0018(1) Lung cancer in the US: Incidence, mortality, and survival Lung cancer is the second most common cancer diagnosed annually and the leading cause of mortality in the US.2 228,820 20.5% 57% Estimated newly 5-year Advanced or 1 survival rate1 metastatic at diagnosed cases in 2020 diagnosis1 5.8% 5-year relative 80-85% 2 135,720 survival with NSCLC distant disease1 Estimated deaths in 20201 2 NSCLC, non-small cell lung cancer; US, United States. 1. National Institutes of Health (NIH), National Cancer Institute. Cancer Stat Facts: Lung and Bronchus Cancer website. www.seer.cancer.gov/statfacts/html/lungb.html. Accessed May 20, 2020. 2. American Cancer Society. What is Lung Cancer? website. https://www.cancer.org/cancer/non-small-cell-lung-cancer/about/what-is-non-small-cell-lung-cancer.html. Accessed May 20, 2020. NSCLC is both histologically and genetically diverse 1-3 NSCLC distribution by histology Prevalence of genetic alterations in NSCLC4 PTEN 10% DDR2 3% OTHER 25% PIK3CA 12% LARGE CELL CARCINOMA 10% FGFR1 20% SQUAMOUS CELL CARCINOMA 25% Oncogenic drivers in adenocarcinoma Other or ADENOCARCINOMA HER2 1.9% 40% KRAS 25.5% wild type RET 0.7% 55% NTRK1 1.7% ROS1 1.7% Oncogenic drivers in 0% 20% 40% 60% RIT1 2.2% squamous cell carcinoma Adenocarcinoma DDR2 2.9% Squamous cell carcinoma NRG1 3.2% Large cell carcinoma -

Essential Role of Autophagy in Protecting Neonatal Haematopoietic Stem Cells from Oxidative Stress in a P62-Independent Manner
www.nature.com/scientificreports OPEN Essential role of autophagy in protecting neonatal haematopoietic stem cells from oxidative stress in a p62‑independent manner Naho Nomura1,7,10, Chiaki Ito1,10, Takako Ooshio1,8, Yuko Tadokoro1,2, Susumu Kohno3, Masaya Ueno1,2, Masahiko Kobayashi1,2, Atsuko Kasahara4, Yusuke Takase1,9, Kenta Kurayoshi1, Sha Si1,2, Chiaki Takahashi3, Masaaki Komatsu5, Toru Yanagawa6 & Atsushi Hirao1,2* Autophagy is a cellular degradation system contributing to homeostasis of tissue stem cells including haematopoietic stem cells (HSCs). It plays pleiotropic roles in HSC characteristics throughout life, but its stage‑specifc roles in HSC self‑renewal are unclear. To investigate the efects of Atg5 deletion on stage‑specifc HSC functions, we compared the repopulating capacity of HSCs in Atg5f/f;Vavi-cre mice from postnatal day (P) 0–7 weeks of age. Interestingly, Atg5 defciency led to no remarkable abnormality in the HSC self‑renewal capacity at P0, but signifcant defects at P7, followed by severe defects. Induction of Atg5 deletion at P5 by tamoxifen administration to Atg5f/f;Rosa26-Cre-ERT2 mice resulted in normal haematopoiesis, including the HSC population, until around 1 year, suggesting that Atg5 in the early neonatal period was critical for haematopoiesis in adults. Mitochondrial oxidative stress was increased by Atg5 loss in neonatal HSC/progenitor cells. Although p62 had accumulated in immature bone marrow cells of Atg5f/f;Vavi-cre mice, p62 deletion did not restore defective HSC functions, indicating that Atg5‑dependent haematopoietic regulation in the developmental period was independent of p62. This study proposes a critical role of autophagy in HSC protection against harsh environments in the early neonatal stage, which is essential for healthy long‑term haematopoiesis. -

Sprouty Proteins, Masterminds of Receptor Tyrosine Kinase Signaling
CORE Metadata, citation and similar papers at core.ac.uk Provided by RERO DOC Digital Library Angiogenesis (2008) 11:53–62 DOI 10.1007/s10456-008-9089-1 ORIGINAL PAPER Sprouty proteins, masterminds of receptor tyrosine kinase signaling Miguel A. Cabrita Æ Gerhard Christofori Received: 14 December 2007 / Accepted: 7 January 2008 / Published online: 25 January 2008 Ó Springer Science+Business Media B.V. 2008 Abstract Angiogenesis relies on endothelial cells prop- Abbreviations erly processing signals from growth factors provided in Ang Angiopoietin both an autocrine and a paracrine manner. These mitogens c-Cbl Cellular homologue of Casitas B-lineage bind to their cognate receptor tyrosine kinases (RTKs) on lymphoma proto-oncogene product the cell surface, thereby activating a myriad of complex EGF Epidermal growth factor intracellular signaling pathways whose outputs include cell EGFR EGF receptor growth, migration, and morphogenesis. Understanding how eNOS Endothelial nitric oxide synthase these cascades are precisely controlled will provide insight ERK Extracellular signal-regulated kinase into physiological and pathological angiogenesis. The FGF Fibroblast growth factor Sprouty (Spry) family of proteins is a highly conserved FGFR FGF receptor group of negative feedback loop modulators of growth GDNF Glial-derived neurotrophic factor factor-mediated mitogen-activated protein kinase (MAPK) Grb2 Growth factor receptor-bound protein 2 activation originally described in Drosophila. There are HMVEC Human microvascular endothelial cell four mammalian orthologs (Spry1-4) whose modulation of Hrs Hepatocyte growth factor-regulated tyrosine RTK-induced signaling pathways is growth factor – and kinase substrate cell context – dependant. Endothelial cells are a group of HUVEC Human umbilical vein endothelial cell highly differentiated cell types necessary for defining the MAPK Mitogen-activated protein kinase mammalian vasculature. -

T Cell Depending on the Differentiation State of the Dual
Dual Effects of Sprouty1 on TCR Signaling Depending on the Differentiation State of the T Cell This information is current as Heonsik Choi, Sung-Yup Cho, Ronald H. Schwartz and of September 29, 2021. Kyungho Choi J Immunol 2006; 176:6034-6045; ; doi: 10.4049/jimmunol.176.10.6034 http://www.jimmunol.org/content/176/10/6034 Downloaded from References This article cites 63 articles, 29 of which you can access for free at: http://www.jimmunol.org/content/176/10/6034.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication by guest on September 29, 2021 *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2006 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Dual Effects of Sprouty1 on TCR Signaling Depending on the Differentiation State of the T Cell1 Heonsik Choi,* Sung-Yup Cho,† Ronald H. Schwartz,* and Kyungho Choi2* Sprouty (Spry) is known to be a negative feedback inhibitor of growth factor receptor signaling through inhibition of the Ras/ MAPK pathway. -

Identification of Transcriptional Mechanisms Downstream of Nf1 Gene Defeciency in Malignant Peripheral Nerve Sheath Tumors Daochun Sun Wayne State University
Wayne State University DigitalCommons@WayneState Wayne State University Dissertations 1-1-2012 Identification of transcriptional mechanisms downstream of nf1 gene defeciency in malignant peripheral nerve sheath tumors Daochun Sun Wayne State University, Follow this and additional works at: http://digitalcommons.wayne.edu/oa_dissertations Recommended Citation Sun, Daochun, "Identification of transcriptional mechanisms downstream of nf1 gene defeciency in malignant peripheral nerve sheath tumors" (2012). Wayne State University Dissertations. Paper 558. This Open Access Dissertation is brought to you for free and open access by DigitalCommons@WayneState. It has been accepted for inclusion in Wayne State University Dissertations by an authorized administrator of DigitalCommons@WayneState. IDENTIFICATION OF TRANSCRIPTIONAL MECHANISMS DOWNSTREAM OF NF1 GENE DEFECIENCY IN MALIGNANT PERIPHERAL NERVE SHEATH TUMORS by DAOCHUN SUN DISSERTATION Submitted to the Graduate School of Wayne State University, Detroit, Michigan in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY 2012 MAJOR: MOLECULAR BIOLOGY AND GENETICS Approved by: _______________________________________ Advisor Date _______________________________________ _______________________________________ _______________________________________ © COPYRIGHT BY DAOCHUN SUN 2012 All Rights Reserved DEDICATION This work is dedicated to my parents and my wife Ze Zheng for their continuous support and understanding during the years of my education. I could not achieve my goal without them. ii ACKNOWLEDGMENTS I would like to express tremendous appreciation to my mentor, Dr. Michael Tainsky. His guidance and encouragement throughout this project made this dissertation come true. I would also like to thank my committee members, Dr. Raymond Mattingly and Dr. John Reiners Jr. for their sustained attention to this project during the monthly NF1 group meetings and committee meetings, Dr. -

Transcriptomic and Epigenomic Characterization of the Developing Bat Wing
ARTICLES OPEN Transcriptomic and epigenomic characterization of the developing bat wing Walter L Eckalbar1,2,9, Stephen A Schlebusch3,9, Mandy K Mason3, Zoe Gill3, Ash V Parker3, Betty M Booker1,2, Sierra Nishizaki1,2, Christiane Muswamba-Nday3, Elizabeth Terhune4,5, Kimberly A Nevonen4, Nadja Makki1,2, Tara Friedrich2,6, Julia E VanderMeer1,2, Katherine S Pollard2,6,7, Lucia Carbone4,8, Jeff D Wall2,7, Nicola Illing3 & Nadav Ahituv1,2 Bats are the only mammals capable of powered flight, but little is known about the genetic determinants that shape their wings. Here we generated a genome for Miniopterus natalensis and performed RNA-seq and ChIP-seq (H3K27ac and H3K27me3) analyses on its developing forelimb and hindlimb autopods at sequential embryonic stages to decipher the molecular events that underlie bat wing development. Over 7,000 genes and several long noncoding RNAs, including Tbx5-as1 and Hottip, were differentially expressed between forelimb and hindlimb, and across different stages. ChIP-seq analysis identified thousands of regions that are differentially modified in forelimb and hindlimb. Comparative genomics found 2,796 bat-accelerated regions within H3K27ac peaks, several of which cluster near limb-associated genes. Pathway analyses highlighted multiple ribosomal proteins and known limb patterning signaling pathways as differentially regulated and implicated increased forelimb mesenchymal condensation in differential growth. In combination, our work outlines multiple genetic components that likely contribute to bat wing formation, providing insights into this morphological innovation. The order Chiroptera, commonly known as bats, is the only group of To characterize the genetic differences that underlie divergence in mammals to have evolved the capability of flight. -
Beyond Traditional Morphological Characterization of Lung
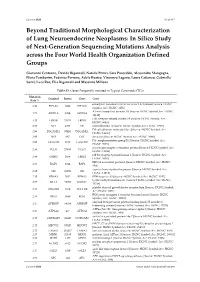
Cancers 2020 S1 of S15 Beyond Traditional Morphological Characterization of Lung Neuroendocrine Neoplasms: In Silico Study of Next-Generation Sequencing Mutations Analysis across the Four World Health Organization Defined Groups Giovanni Centonze, Davide Biganzoli, Natalie Prinzi, Sara Pusceddu, Alessandro Mangogna, Elena Tamborini, Federica Perrone, Adele Busico, Vincenzo Lagano, Laura Cattaneo, Gabriella Sozzi, Luca Roz, Elia Biganzoli and Massimo Milione Table S1. Genes Frequently mutated in Typical Carcinoids (TCs). Mutation Original Entrez Gene Gene Rate % eukaryotic translation initiation factor 1A X-linked [Source: HGNC 4.84 EIF1AX 1964 EIF1AX Symbol; Acc: HGNC: 3250] AT-rich interaction domain 1A [Source: HGNC Symbol;Acc: HGNC: 4.71 ARID1A 8289 ARID1A 11110] LDL receptor related protein 1B [Source: HGNC Symbol; Acc: 4.35 LRP1B 53353 LRP1B HGNC: 6693] 3.53 NF1 4763 NF1 neurofibromin 1 [Source: HGNC Symbol;Acc: HGNC: 7765] DS cell adhesion molecule like 1 [Source: HGNC Symbol; Acc: 2.90 DSCAML1 57453 DSCAML1 HGNC: 14656] 2.90 DST 667 DST dystonin [Source: HGNC Symbol;Acc: HGNC: 1090] FA complementation group D2 [Source: HGNC Symbol; Acc: 2.90 FANCD2 2177 FANCD2 HGNC: 3585] piccolo presynaptic cytomatrix protein [Source: HGNC Symbol; Acc: 2.90 PCLO 27445 PCLO HGNC: 13406] erb-b2 receptor tyrosine kinase 2 [Source: HGNC Symbol; Acc: 2.44 ERBB2 2064 ERBB2 HGNC: 3430] BRCA1 associated protein 1 [Source: HGNC Symbol; Acc: HGNC: 2.35 BAP1 8314 BAP1 950] capicua transcriptional repressor [Source: HGNC Symbol; Acc: 2.35 CIC 23152 CIC HGNC: -

The NF2 Tumor Suppressor Merlin Interacts with Ras and Rasgap, Which May Modulate Ras Signaling
Oncogene (2019) 38:6370–6381 https://doi.org/10.1038/s41388-019-0883-6 ARTICLE The NF2 tumor suppressor merlin interacts with Ras and RasGAP, which may modulate Ras signaling 1 1 2 1 1 1 Yan Cui ● Susann Groth ● Scott Troutman ● Annemarie Carlstedt ● Tobias Sperka ● Lars Björn Riecken ● 2 3 1 Joseph L. Kissil ● Hongchuan Jin ● Helen Morrison Received: 5 July 2018 / Revised: 31 March 2019 / Accepted: 1 May 2019 / Published online: 16 July 2019 © The Author(s) 2019. This article is published with open access Abstract Inactivation of the tumor suppressor NF2/merlin underlies neurofibromatosis type 2 (NF2) and some sporadic tumors. Previous studies have established that merlin mediates contact inhibition of proliferation; however, the exact mechanisms remain obscure and multiple pathways have been implicated. We have previously reported that merlin inhibits Ras and Rac activity during contact inhibition, but how merlin regulates Ras activity has remained elusive. Here we demonstrate that merlin can directly interact with both Ras and p120RasGAP (also named RasGAP). While merlin does not increase the catalytic activity of RasGAP, the interactions with Ras and RasGAP may fine-tune Ras signaling. In vivo, loss of RasGAP in 1234567890();,: 1234567890();,: Schwann cells, unlike the loss of merlin, failed to promote tumorigenic growth in an orthotopic model. Therefore, modulation of Ras signaling through RasGAP likely contributes to, but is not sufficient to account for, merlin’s tumor suppressor activity. Our study provides new insight into the mechanisms of merlin-dependent Ras regulation and may have additional implications for merlin-dependent regulation of other small GTPases. Introduction ependymomas, and astrocytomas [1]. -

Mir-126-3P Contributes to Sorafenib Resistance in Hepatocellular Carcinoma Via Downregulating SPRED1
38 Original Article Page 1 of 14 miR-126-3p contributes to sorafenib resistance in hepatocellular carcinoma via downregulating SPRED1 Wenliang Tan1,2#, Zhirong Lin1,2#, Xianqing Chen3, Wenxin Li4, Sicong Zhu1,5, Yingcheng Wei1,2, Liyun Huo1,2, Yajin Chen1,2, Changzhen Shang1,2 1Guangdong Provincial Key Laboratory of Malignant Tumor Epigenetics and Gene Regulation, Sun Yat-sen Memorial Hospital, Sun Yat-Sen University, Guangzhou, China; 2Department of Hepatobiliary Surgery, Sun Yat-sen Memorial Hospital, Sun Yat-sen University, Guangzhou, China; 3Department of Hepatobiliary Surgery, the Eighth Affiliated Hospital, Sun Yat-sen University, Shenzhen, China; 4Department of Cardiology, the Eighth Affiliated Hospital, Sun Yat-sen University, Shenzhen, China; 5Department of Surgical Intensive Care Unit, Sun Yat-sen Memorial Hospital, Sun Yat-sen University, Guangzhou, China Contributions: (I) Conception and design: C Shang, Y Chen; (II) Administrative support: C Shang, Y Chen; (III) Provision of study materials or patients: W Tan, Z Lin; (IV) Collection and assembly of data: S Zhu, Y Wei, L Huo; (V) Data analysis and interpretation: X Chen, W Li; (VI) Manuscript writing: All authors; (VII) Final approval of manuscript: All authors. #These authors contributed equally to this work. Correspondence to: Changzhen Shang; Yajin Chen. Department of Hepatobiliary Surgery, Sun Yat-sen Memorial Hospital of Sun Yat-sen University, Guangzhou 510120, China. Email: [email protected]; [email protected]. Background: Sorafenib can prolong the survival of patients with advanced hepatocellular carcinoma (HCC). However, drug resistance remains the main obstacle to improving its efficiency. This study aimed to explore the likely molecular mechanism of sorafenib resistance. Methods: Differentially expressed microRNAs (miRNAs) related to sorafenib response were analyzed with the Limma package in R software.