The NF2 Tumor Suppressor Merlin Interacts with Ras and Rasgap, Which May Modulate Ras Signaling
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Retinoblastoma Tumor-Suppressor Gene, the Exception That Proves the Rule
Oncogene (2006) 25, 5233–5243 & 2006 Nature Publishing Group All rights reserved 0950-9232/06 $30.00 www.nature.com/onc REVIEW The retinoblastoma tumor-suppressor gene, the exception that proves the rule DW Goodrich Department of Pharmacology & Therapeutics, Roswell Park Cancer Institute, Buffalo, NY, USA The retinoblastoma tumor-suppressor gene (Rb1)is transmission of one mutationally inactivated Rb1 allele centrally important in cancer research. Mutational and loss of the remaining wild-type allele in somatic inactivation of Rb1 causes the pediatric cancer retino- retinal cells. Hence hereditary retinoblastoma typically blastoma, while deregulation ofthe pathway in which it has an earlier onset and a greater number of tumor foci functions is common in most types of human cancer. The than sporadic retinoblastoma where both Rb1 alleles Rb1-encoded protein (pRb) is well known as a general cell must be inactivated in somatic retinal cells. To this day, cycle regulator, and this activity is critical for pRb- Rb1 remains an exception among cancer-associated mediated tumor suppression. The main focus of this genes in that its mutation is apparently both necessary review, however, is on more recent evidence demonstrating and sufficient, or at least rate limiting, for the genesis of the existence ofadditional, cell type-specific pRb func- a human cancer. The simple genetics of retinoblastoma tions in cellular differentiation and survival. These has spawned the hope that a complete molecular additional functions are relevant to carcinogenesis sug- understanding of the Rb1-encoded protein (pRb) would gesting that the net effect of Rb1 loss on the behavior of lead to deeper insight into the processes of neoplastic resulting tumors is highly dependent on biological context. -

Wt1) – an Effector in Leukemogenesis?
WILMS’ TUMOUR GENE 1 PROTEIN (WT1) – AN EFFECTOR IN LEUKEMOGENESIS? Vidovic, Karina 2010 Link to publication Citation for published version (APA): Vidovic, K. (2010). WILMS’ TUMOUR GENE 1 PROTEIN (WT1) – AN EFFECTOR IN LEUKEMOGENESIS?. Lund University: Faculty of Medicine. Total number of authors: 1 General rights Unless other specific re-use rights are stated the following general rights apply: Copyright and moral rights for the publications made accessible in the public portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. • Users may download and print one copy of any publication from the public portal for the purpose of private study or research. • You may not further distribute the material or use it for any profit-making activity or commercial gain • You may freely distribute the URL identifying the publication in the public portal Read more about Creative commons licenses: https://creativecommons.org/licenses/ Take down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. LUND UNIVERSITY PO Box 117 221 00 Lund +46 46-222 00 00 Download date: 07. Oct. 2021 Division of Hematology and Transfusion Medicine Lund University, Lund, Sweden WILMS’ TUMOUR GENE 1 PROTEIN (WT1) – AN EFFECTOR IN LEUKEMOGENESIS? Karina Vidovic Thesis 2010 Contact adress Karina Vidovic Division of Hematology and Transfusion Medicine BMC, C14 Klinikgatan 28 SE- 221 84 Lund Sweden Phone +46 46 222 07 30 e-mail: [email protected] ISBN 978-91-86443-99-3 ! Karina Vidovic Printed by Media-Tryck, Lund, Sweden 2 ”The journey of a thousand miles begins with one step” Lao Tzu (Chinese taoist), 600-531 BC 3 4 LIST OF PAPERS This thesis is based on the following papers, referred to in the text by their Roman numerals I. -

DNA Microarrays (Gene Chips) and Cancer
DNA Microarrays (Gene Chips) and Cancer Cancer Education Project University of Rochester DNA Microarrays (Gene Chips) and Cancer http://www.biosci.utexas.edu/graduate/plantbio/images/spot/microarray.jpg http://www.affymetrix.com Part 1 Gene Expression and Cancer Nucleus Proteins DNA RNA Cell membrane All your cells have the same DNA Sperm Embryo Egg Fertilized Egg - Zygote How do cells that have the same DNA (genes) end up having different structures and functions? DNA in the nucleus Genes Different genes are turned on in different cells. DIFFERENTIAL GENE EXPRESSION GENE EXPRESSION (Genes are “on”) Transcription Translation DNA mRNA protein cell structure (Gene) and function Converts the DNA (gene) code into cell structure and function Differential Gene Expression Different genes Different genes are turned on in different cells make different mRNA’s Differential Gene Expression Different genes are turned Different genes Different mRNA’s on in different cells make different mRNA’s make different Proteins An example of differential gene expression White blood cell Stem Cell Platelet Red blood cell Bone marrow stem cells differentiate into specialized blood cells because different genes are expressed during development. Normal Differential Gene Expression Genes mRNA mRNA Expression of different genes results in the cell developing into a red blood cell or a white blood cell Cancer and Differential Gene Expression mRNA Genes But some times….. Mutations can lead to CANCER CELL some genes being Abnormal gene expression more or less may result -

P14ARF Inhibits Human Glioblastoma–Induced Angiogenesis by Upregulating the Expression of TIMP3
P14ARF inhibits human glioblastoma–induced angiogenesis by upregulating the expression of TIMP3 Abdessamad Zerrouqi, … , Daniel J. Brat, Erwin G. Van Meir J Clin Invest. 2012;122(4):1283-1295. https://doi.org/10.1172/JCI38596. Research Article Oncology Malignant gliomas are the most common and the most lethal primary brain tumors in adults. Among malignant gliomas, 60%–80% show loss of P14ARF tumor suppressor activity due to somatic alterations of the INK4A/ARF genetic locus. The tumor suppressor activity of P14ARF is in part a result of its ability to prevent the degradation of P53 by binding to and sequestering HDM2. However, the subsequent finding of P14ARF loss in conjunction with TP53 gene loss in some tumors suggests the protein may have other P53-independent tumor suppressor functions. Here, we report what we believe to be a novel tumor suppressor function for P14ARF as an inhibitor of tumor-induced angiogenesis. We found that P14ARF mediates antiangiogenic effects by upregulating expression of tissue inhibitor of metalloproteinase–3 (TIMP3) in a P53-independent fashion. Mechanistically, this regulation occurred at the gene transcription level and was controlled by HDM2-SP1 interplay, where P14ARF relieved a dominant negative interaction of HDM2 with SP1. P14ARF-induced expression of TIMP3 inhibited endothelial cell migration and vessel formation in response to angiogenic stimuli produced by cancer cells. The discovery of this angiogenesis regulatory pathway may provide new insights into P53-independent P14ARF tumor-suppressive mechanisms that have implications for the development of novel therapies directed at tumors and other diseases characterized by vascular pathology. Find the latest version: https://jci.me/38596/pdf Research article P14ARF inhibits human glioblastoma–induced angiogenesis by upregulating the expression of TIMP3 Abdessamad Zerrouqi,1 Beata Pyrzynska,1,2 Maria Febbraio,3 Daniel J. -

Review Article PTEN Gene: a Model for Genetic Diseases in Dermatology
The Scientific World Journal Volume 2012, Article ID 252457, 8 pages The cientificWorldJOURNAL doi:10.1100/2012/252457 Review Article PTEN Gene: A Model for Genetic Diseases in Dermatology Corrado Romano1 and Carmelo Schepis2 1 Unit of Pediatrics and Medical Genetics, I.R.C.C.S. Associazione Oasi Maria Santissima, 94018 Troina, Italy 2 Unit of Dermatology, I.R.C.C.S. Associazione Oasi Maria Santissima, 94018 Troina, Italy Correspondence should be addressed to Carmelo Schepis, [email protected] Received 19 October 2011; Accepted 4 January 2012 Academic Editors: G. Vecchio and H. Zitzelsberger Copyright © 2012 C. Romano and C. Schepis. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. PTEN gene is considered one of the most mutated tumor suppressor genes in human cancer, and it’s likely to become the first one in the near future. Since 1997, its involvement in tumor suppression has smoothly increased, up to the current importance. Germline mutations of PTEN cause the PTEN hamartoma tumor syndrome (PHTS), which include the past-called Cowden, Bannayan- Riley-Ruvalcaba, Proteus, Proteus-like, and Lhermitte-Duclos syndromes. Somatic mutations of PTEN have been observed in glioblastoma, prostate cancer, and brest cancer cell lines, quoting only the first tissues where the involvement has been proven. The negative regulation of cell interactions with the extracellular matrix could be the way PTEN phosphatase acts as a tumor suppressor. PTEN gene plays an essential role in human development. A recent model sees PTEN function as a stepwise gradation, which can be impaired not only by heterozygous mutations and homozygous losses, but also by other molecular mechanisms, such as transcriptional regression, epigenetic silencing, regulation by microRNAs, posttranslational modification, and aberrant localization. -

(NF1) Defects Are Common in Human Ovarian Serous Carcinomas and Co-Occur with TP53 Mutations
Volume 10 Number 12 December 2008 pp. 1362–1372 1362 www.neoplasia.com Neurofibromin 1 (NF1) Defects Navneet Sangha*, Rong Wu*, Rork Kuick†, Scott Powers‡, David Mu§, Diane Fiander*, Are Common in Human Ovarian Kit Yuen*, Hidetaka Katabuchi¶, Hironori Tashiro¶, Serous Carcinomas and Co-occur Eric R. Fearon*,†,# and Kathleen R. Cho*,†,# 1,2 with TP53 Mutations *Department of Pathology, The University of Michigan Medical School, Ann Arbor, MI 48109, USA; †The Comprehensive Cancer Center, The University of Michigan Medical School, Ann Arbor, MI 48109, USA; ‡Cold Spring Harbor Laboratory, Cold Spring Harbor, NY 11724, USA; §Department of Pathology, Penn State University College of Medicine, Hershey, PA 17033, USA; ¶Department of Gynecology, Kumamoto University, Kumamoto, 860-8556, Japan; #Department of Internal Medicine, The University of Michigan Medical School, Ann Arbor, MI 48109, USA Abstract Ovarian serous carcinoma (OSC) is the most common and lethal histologic type of ovarian epithelial malignancy. Mutations of TP53 and dysfunction of the Brca1 and/or Brca2 tumor-suppressor proteins have been implicated in the molecular pathogenesis of a large fraction of OSCs, but frequent somatic mutations in other well-established tumor-suppressor genes have not been identified. Using a genome-wide screen of DNA copy number alterations in 36 primary OSCs, we identified two tumors with apparent homozygous deletions of the NF1 gene. Subsequently, 18 ovarian carcinoma–derived cell lines and 41 primary OSCs were evaluated for NF1 alterations. Markedly re- duced or absent expression of Nf1 protein was observed in 6 of the 18 cell lines, and using the protein truncation test and sequencing of cDNA and genomic DNA, NF1 mutations resulting in deletion of exons and/or aberrant splicing of NF1 transcripts were detected in 5 of the 6 cell lines with loss of NF1 expression. -

Wnt-Independent and Wnt-Dependent Effects of APC Loss on the Chemotherapeutic Response
International Journal of Molecular Sciences Review Wnt-Independent and Wnt-Dependent Effects of APC Loss on the Chemotherapeutic Response Casey D. Stefanski 1,2 and Jenifer R. Prosperi 1,2,3,* 1 Department of Biological Sciences, University of Notre Dame, Notre Dame, IN 46617, USA; [email protected] 2 Mike and Josie Harper Cancer Research Institute, South Bend, IN 46617, USA 3 Department of Biochemistry and Molecular Biology, Indiana University School of Medicine-South Bend, South Bend, IN 46617, USA * Correspondence: [email protected]; Tel.: +1-574-631-4002 Received: 30 September 2020; Accepted: 20 October 2020; Published: 22 October 2020 Abstract: Resistance to chemotherapy occurs through mechanisms within the epithelial tumor cells or through interactions with components of the tumor microenvironment (TME). Chemoresistance and the development of recurrent tumors are two of the leading factors of cancer-related deaths. The Adenomatous Polyposis Coli (APC) tumor suppressor is lost in many different cancers, including colorectal, breast, and prostate cancer, and its loss correlates with a decreased overall survival in cancer patients. While APC is commonly known for its role as a negative regulator of the WNT pathway, APC has numerous binding partners and functional roles. Through APC’s interactions with DNA repair proteins, DNA replication proteins, tubulin, and other components, recent evidence has shown that APC regulates the chemotherapy response in cancer cells. In this review article, we provide an overview of some of the cellular processes in which APC participates and how they impact chemoresistance through both epithelial- and TME-derived mechanisms. Keywords: adenomatous polyposis coli; chemoresistance; WNT signaling 1. -

Teacher Background on P53 Tumor Suppressor Protein
Cancer Lab p53 – Teacher Background on p53 Tumor Suppressor Protein Note: The Teacher Background Section is meant to provide information for the teacher about the topic and is tied very closely to the PowerPoint slide show. For greater understanding, the teacher may want to play the slide show as he/she reads the background section. For the students, the slide show can be used in its entirety or can be edited as necessary for a given class. What Is p53 and Where Is the Gene Located? While commonly known as p53, the official name of this gene is Tumor Protein p53 and its official symbol is TP53. TheTP53 gene codes for the TP53 (p53) protein which acts as a tumor suppressor and works in response to DNA damage to orchestrate the repair of damaged DNA. If the DNA cannot be repaired, the p53 protein prevents the cell from dividing and signals it to undergo apoptosis (programmed cell death). The name p53 is due to protein’s 53 kilo-Dalton molecular mass. The gene which codes for this protein is located on the short (p) arm of chromosome 17 at position 13.1 (17p13.1). The gene begins at base pair 7,571,719 and ends at base pair 7, 590,862 making it 19,143 base pairs long. (1, 2) What Does the p53 Gene Look Like When Translated Into Protein? The TP53 gene has 11 exons and a very large 10 kb intron between exons 1 and 2. In humans, exon 1 is non-coding and it has been shown that this region could form a stable stem-loop structure which binds tightly to normal p53 but not to mutant p53 proteins. -

Biomarker Testing in Non- Small Cell Lung Cancer (NSCLC)
The biopharma business of Merck KGaA, Darmstadt, Germany operates as EMD Serono in the U.S. and Canada. Biomarker testing in non- small cell lung cancer (NSCLC) Copyright © 2020 EMD Serono, Inc. All rights reserved. US/TEP/1119/0018(1) Lung cancer in the US: Incidence, mortality, and survival Lung cancer is the second most common cancer diagnosed annually and the leading cause of mortality in the US.2 228,820 20.5% 57% Estimated newly 5-year Advanced or 1 survival rate1 metastatic at diagnosed cases in 2020 diagnosis1 5.8% 5-year relative 80-85% 2 135,720 survival with NSCLC distant disease1 Estimated deaths in 20201 2 NSCLC, non-small cell lung cancer; US, United States. 1. National Institutes of Health (NIH), National Cancer Institute. Cancer Stat Facts: Lung and Bronchus Cancer website. www.seer.cancer.gov/statfacts/html/lungb.html. Accessed May 20, 2020. 2. American Cancer Society. What is Lung Cancer? website. https://www.cancer.org/cancer/non-small-cell-lung-cancer/about/what-is-non-small-cell-lung-cancer.html. Accessed May 20, 2020. NSCLC is both histologically and genetically diverse 1-3 NSCLC distribution by histology Prevalence of genetic alterations in NSCLC4 PTEN 10% DDR2 3% OTHER 25% PIK3CA 12% LARGE CELL CARCINOMA 10% FGFR1 20% SQUAMOUS CELL CARCINOMA 25% Oncogenic drivers in adenocarcinoma Other or ADENOCARCINOMA HER2 1.9% 40% KRAS 25.5% wild type RET 0.7% 55% NTRK1 1.7% ROS1 1.7% Oncogenic drivers in 0% 20% 40% 60% RIT1 2.2% squamous cell carcinoma Adenocarcinoma DDR2 2.9% Squamous cell carcinoma NRG1 3.2% Large cell carcinoma -

Identification of Transcriptional Mechanisms Downstream of Nf1 Gene Defeciency in Malignant Peripheral Nerve Sheath Tumors Daochun Sun Wayne State University
Wayne State University DigitalCommons@WayneState Wayne State University Dissertations 1-1-2012 Identification of transcriptional mechanisms downstream of nf1 gene defeciency in malignant peripheral nerve sheath tumors Daochun Sun Wayne State University, Follow this and additional works at: http://digitalcommons.wayne.edu/oa_dissertations Recommended Citation Sun, Daochun, "Identification of transcriptional mechanisms downstream of nf1 gene defeciency in malignant peripheral nerve sheath tumors" (2012). Wayne State University Dissertations. Paper 558. This Open Access Dissertation is brought to you for free and open access by DigitalCommons@WayneState. It has been accepted for inclusion in Wayne State University Dissertations by an authorized administrator of DigitalCommons@WayneState. IDENTIFICATION OF TRANSCRIPTIONAL MECHANISMS DOWNSTREAM OF NF1 GENE DEFECIENCY IN MALIGNANT PERIPHERAL NERVE SHEATH TUMORS by DAOCHUN SUN DISSERTATION Submitted to the Graduate School of Wayne State University, Detroit, Michigan in partial fulfillment of the requirements for the degree of DOCTOR OF PHILOSOPHY 2012 MAJOR: MOLECULAR BIOLOGY AND GENETICS Approved by: _______________________________________ Advisor Date _______________________________________ _______________________________________ _______________________________________ © COPYRIGHT BY DAOCHUN SUN 2012 All Rights Reserved DEDICATION This work is dedicated to my parents and my wife Ze Zheng for their continuous support and understanding during the years of my education. I could not achieve my goal without them. ii ACKNOWLEDGMENTS I would like to express tremendous appreciation to my mentor, Dr. Michael Tainsky. His guidance and encouragement throughout this project made this dissertation come true. I would also like to thank my committee members, Dr. Raymond Mattingly and Dr. John Reiners Jr. for their sustained attention to this project during the monthly NF1 group meetings and committee meetings, Dr. -
Beyond Traditional Morphological Characterization of Lung
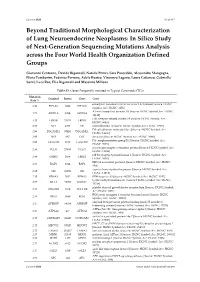
Cancers 2020 S1 of S15 Beyond Traditional Morphological Characterization of Lung Neuroendocrine Neoplasms: In Silico Study of Next-Generation Sequencing Mutations Analysis across the Four World Health Organization Defined Groups Giovanni Centonze, Davide Biganzoli, Natalie Prinzi, Sara Pusceddu, Alessandro Mangogna, Elena Tamborini, Federica Perrone, Adele Busico, Vincenzo Lagano, Laura Cattaneo, Gabriella Sozzi, Luca Roz, Elia Biganzoli and Massimo Milione Table S1. Genes Frequently mutated in Typical Carcinoids (TCs). Mutation Original Entrez Gene Gene Rate % eukaryotic translation initiation factor 1A X-linked [Source: HGNC 4.84 EIF1AX 1964 EIF1AX Symbol; Acc: HGNC: 3250] AT-rich interaction domain 1A [Source: HGNC Symbol;Acc: HGNC: 4.71 ARID1A 8289 ARID1A 11110] LDL receptor related protein 1B [Source: HGNC Symbol; Acc: 4.35 LRP1B 53353 LRP1B HGNC: 6693] 3.53 NF1 4763 NF1 neurofibromin 1 [Source: HGNC Symbol;Acc: HGNC: 7765] DS cell adhesion molecule like 1 [Source: HGNC Symbol; Acc: 2.90 DSCAML1 57453 DSCAML1 HGNC: 14656] 2.90 DST 667 DST dystonin [Source: HGNC Symbol;Acc: HGNC: 1090] FA complementation group D2 [Source: HGNC Symbol; Acc: 2.90 FANCD2 2177 FANCD2 HGNC: 3585] piccolo presynaptic cytomatrix protein [Source: HGNC Symbol; Acc: 2.90 PCLO 27445 PCLO HGNC: 13406] erb-b2 receptor tyrosine kinase 2 [Source: HGNC Symbol; Acc: 2.44 ERBB2 2064 ERBB2 HGNC: 3430] BRCA1 associated protein 1 [Source: HGNC Symbol; Acc: HGNC: 2.35 BAP1 8314 BAP1 950] capicua transcriptional repressor [Source: HGNC Symbol; Acc: 2.35 CIC 23152 CIC HGNC: -

Cancer Biology Introduction Proto-Oncogenes Tumor
Introduction • Tissue homeostasis depends on the regulated cell division and self-elimination (programmed cell Cancer Biology death) of each of its constituent members except its stem cells • A tumor arises as a result of uncontrolled cell division and failure for self-elimination Chapter 18 • Alterations in three groups of genes are responsible Eric J. Hall., Amato Giaccia, for the deregulated control mechanisms that are the hallmarks of cancer cells: proto-oncogenes, tumor- Radiobiology for the Radiologist supressor genes, and DNA stability genes Proto-oncogenes Tumor-suppressor genes • Proto-oncogenes are components of signaling • Tumor-suppressor genes are also components of networks that act as positive growth the same signaling networks as proto-oncogenes, except that they act as negative growth regulators regulators in response to mitogens, cytokines, • They modulate proliferation and survival by and cell-to-cell contact antagonizing the biochemical functions of proto- • A gain-of-function mutation in only one copy oncogenes or responding to unchecked growth signals of a protooncogene results in a dominantly • In contrast to oncogenes, inactivation of both acting oncogene that often fails to respond to copies of tumor-suppressor genes is required for extracellular signals loss of function in most cases DNA stability genes Mechanisms of carcinogenesis • DNA stability genes form a class of genes • A single genetic alteration that leads to the involved in both monitoring and activation of an oncogene or loss of a tumor maintaining