Bremer Et Al. 2001
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Toward a Resolution of Campanulid Phylogeny, with Special Reference to the Placement of Dipsacales
TAXON 57 (1) • February 2008: 53–65 Winkworth & al. • Campanulid phylogeny MOLECULAR PHYLOGENETICS Toward a resolution of Campanulid phylogeny, with special reference to the placement of Dipsacales Richard C. Winkworth1,2, Johannes Lundberg3 & Michael J. Donoghue4 1 Departamento de Botânica, Instituto de Biociências, Universidade de São Paulo, Caixa Postal 11461–CEP 05422-970, São Paulo, SP, Brazil. [email protected] (author for correspondence) 2 Current address: School of Biology, Chemistry, and Environmental Sciences, University of the South Pacific, Private Bag, Laucala Campus, Suva, Fiji 3 Department of Phanerogamic Botany, The Swedish Museum of Natural History, Box 50007, 104 05 Stockholm, Sweden 4 Department of Ecology & Evolutionary Biology and Peabody Museum of Natural History, Yale University, P.O. Box 208106, New Haven, Connecticut 06520-8106, U.S.A. Broad-scale phylogenetic analyses of the angiosperms and of the Asteridae have failed to confidently resolve relationships among the major lineages of the campanulid Asteridae (i.e., the euasterid II of APG II, 2003). To address this problem we assembled presently available sequences for a core set of 50 taxa, representing the diver- sity of the four largest lineages (Apiales, Aquifoliales, Asterales, Dipsacales) as well as the smaller “unplaced” groups (e.g., Bruniaceae, Paracryphiaceae, Columelliaceae). We constructed four data matrices for phylogenetic analysis: a chloroplast coding matrix (atpB, matK, ndhF, rbcL), a chloroplast non-coding matrix (rps16 intron, trnT-F region, trnV-atpE IGS), a combined chloroplast dataset (all seven chloroplast regions), and a combined genome matrix (seven chloroplast regions plus 18S and 26S rDNA). Bayesian analyses of these datasets using mixed substitution models produced often well-resolved and supported trees. -

The New Zealand Rain Forest: a Comparison with Tropical Rain Forest! J
The New Zealand Rain Forest: A Comparison with Tropical Rain Forest! J. W. DAWSON2 and B. V. SNEDDON2 ABSTRACT: The structure of and growth forms and habits exhibited by the New Zealand rain forest are described and compared with those of lowland tropical rain forest. Theories relating to the frequent regeneration failure of the forest dominants are outlined. The floristic affinities of the forest type are discussed and it is suggested that two main elements can be recognized-lowland tropical and montane tropical. It is concluded that the New Zealand rain forest is comparable to lowland tropical rain forest in structure and in range of special growth forms and habits. It chiefly differs in its lower stature, fewer species, and smaller leaves. The floristic similarity between the present forest and forest floras of the Tertiary in New Zealand suggest that the former may be a floristically reduced derivative of the latter. PART 1 OF THIS PAPER describes the structure The approximate number of species of seed and growth forms of the New Zealand rain plants in these forests is 240. From north to forest as exemplified by a forest in the far north. south there is an overall decrease in number of In Part 2, theories relating to the regeneration species. At about 38°S a number of species, of the dominant trees in the New Zealand rain mostly trees and shrubs, drop out or become forest generally are reviewed briefly, and their restricted to coastal sites, but it is not until about relevance to the situation in the study forest is 42°S, in the South Island, that many of the con considered. -

Flowering Plants Eudicots Apiales, Gentianales (Except Rubiaceae)
Edited by K. Kubitzki Volume XV Flowering Plants Eudicots Apiales, Gentianales (except Rubiaceae) Joachim W. Kadereit · Volker Bittrich (Eds.) THE FAMILIES AND GENERA OF VASCULAR PLANTS Edited by K. Kubitzki For further volumes see list at the end of the book and: http://www.springer.com/series/1306 The Families and Genera of Vascular Plants Edited by K. Kubitzki Flowering Plants Á Eudicots XV Apiales, Gentianales (except Rubiaceae) Volume Editors: Joachim W. Kadereit • Volker Bittrich With 85 Figures Editors Joachim W. Kadereit Volker Bittrich Johannes Gutenberg Campinas Universita¨t Mainz Brazil Mainz Germany Series Editor Prof. Dr. Klaus Kubitzki Universita¨t Hamburg Biozentrum Klein-Flottbek und Botanischer Garten 22609 Hamburg Germany The Families and Genera of Vascular Plants ISBN 978-3-319-93604-8 ISBN 978-3-319-93605-5 (eBook) https://doi.org/10.1007/978-3-319-93605-5 Library of Congress Control Number: 2018961008 # Springer International Publishing AG, part of Springer Nature 2018 This work is subject to copyright. All rights are reserved by the Publisher, whether the whole or part of the material is concerned, specifically the rights of translation, reprinting, reuse of illustrations, recitation, broadcasting, reproduction on microfilms or in any other physical way, and transmission or information storage and retrieval, electronic adaptation, computer software, or by similar or dissimilar methodology now known or hereafter developed. The use of general descriptive names, registered names, trademarks, service marks, etc. in this publication does not imply, even in the absence of a specific statement, that such names are exempt from the relevant protective laws and regulations and therefore free for general use. -

The Species of Alseuosmia (Alseuosmiaceae)
New Zealand Journal of Botany, 1978, Vol. 16: 271-7. 271 The species of Alseuosmia (Alseuosmiaceae) RHYS O. GARDNER Department of Botany, University of Auckland, Private Bag, Auckland, New Zealand (Received 15 September 1977) ABSTRACT A new species Alseuosmia turneri R. 0. Gardner (Alseuosmiaceae Airy Shaw) from the Volcanic Plateau, North Island, New Zealand is described and illustrated. A. linariifolia A. Cunn. is reduced to a variety of A. banksii A. Cunn. and A. quercifolia A. Cunn. is given hybrid status (=^4. banksii A. Cunn. x A. macrophylla A. Cunn.). A key to the four Alseuosmia species, synonymy, and a generalised distribution map are given. A. pusilla Col. is illustrated for the first time. INTRODUCTION judged to be frequent between only one pair of Allan Cunningham (1839) described eight species and the "excessive variability" lies mostly species of a new flowering plant genus Alseuosmia there. from material collected in the Bay of Islands region by Banks and Solander in 1769, by himself in 1826 and 1838, and by his brother Richard in 1833-4. The A NEW SPECIES OF ALSEUOSMIA species, all shrubs of the lowland forest, were sup- A. CUNN. posed to differ from one another chiefly in the shape and toothing of their leaves. An undescribed species of Alseuosmia from the Hooker (1852-5, 1864), with the benefit of Waimarino region of the Volcanic Plateau has been additional Colenso and Sinclair material, reduced known for some time (Cockayne 1928, p. 179; A. P. Cunningham's species to four, but stated that these Druce in Atkinson 1971). The following description four species were "excessively variable". -

Arthur Monrad Johnson Colletion of Botanical Drawings
http://oac.cdlib.org/findaid/ark:/13030/kt7489r5rb No online items Arthur Monrad Johnson colletion of botanical drawings 1914-1941 Processed by Pat L. Walter. Louise M. Darling Biomedical Library History and Special Collections Division History and Special Collections Division UCLA 12-077 Center for Health Sciences Box 951798 Los Angeles, CA 90095-1798 Phone: 310/825-6940 Fax: 310/825-0465 Email: [email protected] URL: http://www.library.ucla.edu/libraries/biomed/his/ ©2008 The Regents of the University of California. All rights reserved. Arthur Monrad Johnson colletion 48 1 of botanical drawings 1914-1941 Descriptive Summary Title: Arthur Monrad Johnson colletion of botanical drawings, Date (inclusive): 1914-1941 Collection number: 48 Creator: Johnson, Arthur Monrad 1878-1943 Extent: 3 boxes (2.5 linear feet) Repository: University of California, Los Angeles. Library. Louise M. Darling Biomedical Library History and Special Collections Division Los Angeles, California 90095-1490 Abstract: Approximately 1000 botanical drawings, most in pen and black ink on paper, of the structural parts of angiosperms and some gymnosperms, by Arthur Monrad Johnson. Many of the illustrations have been published in the author's scientific publications, such as his "Taxonomy of the Flowering Plants" and articles on the genus Saxifraga. Dr. Johnson was both a respected botanist and an accomplished artist beyond his botanical subjects. Physical location: Collection stored off-site (Southern Regional Library Facility): Advance notice required for access. Language of Material: Collection materials in English Preferred Citation [Identification of item], Arthur Monrad Johnson colletion of botanical drawings (Manuscript collection 48). Louise M. Darling Biomedical Library History and Special Collections Division, University of California, Los Angeles. -

The Origin of the Bifurcating Style in Asteraceae (Compositae)
Annals of Botany 117: 1009–1021, 2016 doi:10.1093/aob/mcw033, available online at www.aob.oxfordjournals.org The origin of the bifurcating style in Asteraceae (Compositae) Liliana Katinas1,2,*, Marcelo P. Hernandez 2, Ana M. Arambarri2 and Vicki A. Funk3 1Division Plantas Vasculares, Museo de La Plata, La Plata, Argentina, 2Laboratorio de Morfologıa Comparada de Espermatofitas (LAMCE), Facultad de Ciencias Agrarias y Forestales, Universidad Nacional de La Plata, La Plata, Argentina and 3Department of Botany, NMNH, Smithsonian Institution, Washington D.C., USA *For correspondence. E-mail [email protected] Received: 20 November 2015 Returned for revision: 22 December 2015 Accepted: 8 January 2016 Published electronically: 20 April 2016 Background and Aims The plant family Asteraceae (Compositae) exhibits remarkable morphological variation in the styles of its members. Lack of studies on the styles of the sister families to Asteraceae, Goodeniaceae and Calyceraceae, obscures our understanding of the origin and evolution of this reproductive feature in these groups. The aim of this work was to perform a comparative study of style morphology and to discuss the relevance of im- portant features in the evolution of Asteraceae and its sister families. Methods The histochemistry, venation and general morphology of the styles of members of Goodeniaceae, Calyceraceae and early branching lineages of Asteraceae were analysed and put in a phylogenetic framework to dis- cuss the relevance of style features in the evolution of these families. Key Results The location of lipophilic substances allowed differentiation of receptive from non-receptive style papillae, and the style venation in Goodeniaceae and Calyceraceae proved to be distinctive. -

Oral Allergy Syndrome: a Confuence of Immunology and Phylogeny by Merle K
NATIONAL CENTER FOR CASE STUDY TEACHING IN SCIENCE NATIONAL CENTER FOR CASE STUDY TEACHING IN SCIENCE Oral Allergy Syndrome: A Confuence of Immunology and Phylogeny by Merle K. Heidemann, Mike S. Taylor, Amanda Storm, Cassie Dresser-Briggs, Alexa Warwick, and Peter J.T. White Objectives Upon completion of this case study, you should be able to: • Explain the symptoms of allergic reactions in terms of cell biology (immune system cells). • Describe the molecular features of cross reactivity. • Build phylogenetic trees using morphological data. • Build phylogenetic trees using molecular data and interpret two kinds of phylogenetic trees. Part I – Immunology Sam had a history of allergic reactions, including reactions to various plant pollens. Birch pollen elicited a particularly strong reaction, causing him annoying sneezing fts and a sore throat. As he grew to adulthood, he discovered that he was also variably allergic to common foods, including some raw fruits and vegetables, as well as most nuts. Sam was puzzled by his recent food allergies. Also puzzling was the variability in his reactions. He reacted strongly to some foods; others resulted in only a mild itchy throat. He wondered, What exactly causes allergic reactions? Sam was determined to fgure out how and why he reacted to birch pollen and later became allergic to other plant- based food. He also hoped this information might help him avoid other foods that might cause him allergies. First, Sam decided he needed to know something about the cellular bases of allergic reactions. He knew it had some- thing to do with the immune system. He did some basic research on the Internet and here’s what he found: Allergens are bits of protein from an innocuous foreign substance, such as pollen or food. -

Evolutionary History of Floral Key Innovations in Angiosperms Elisabeth Reyes
Evolutionary history of floral key innovations in angiosperms Elisabeth Reyes To cite this version: Elisabeth Reyes. Evolutionary history of floral key innovations in angiosperms. Botanics. Université Paris Saclay (COmUE), 2016. English. NNT : 2016SACLS489. tel-01443353 HAL Id: tel-01443353 https://tel.archives-ouvertes.fr/tel-01443353 Submitted on 23 Jan 2017 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. NNT : 2016SACLS489 THESE DE DOCTORAT DE L’UNIVERSITE PARIS-SACLAY, préparée à l’Université Paris-Sud ÉCOLE DOCTORALE N° 567 Sciences du Végétal : du Gène à l’Ecosystème Spécialité de Doctorat : Biologie Par Mme Elisabeth Reyes Evolutionary history of floral key innovations in angiosperms Thèse présentée et soutenue à Orsay, le 13 décembre 2016 : Composition du Jury : M. Ronse de Craene, Louis Directeur de recherche aux Jardins Rapporteur Botaniques Royaux d’Édimbourg M. Forest, Félix Directeur de recherche aux Jardins Rapporteur Botaniques Royaux de Kew Mme. Damerval, Catherine Directrice de recherche au Moulon Président du jury M. Lowry, Porter Curateur en chef aux Jardins Examinateur Botaniques du Missouri M. Haevermans, Thomas Maître de conférences au MNHN Examinateur Mme. Nadot, Sophie Professeur à l’Université Paris-Sud Directeur de thèse M. -

Phylogeny and Phylogenetic Taxonomy of Dipsacales, with Special Reference to Sinadoxa and Tetradoxa (Adoxaceae)
PHYLOGENY AND PHYLOGENETIC TAXONOMY OF DIPSACALES, WITH SPECIAL REFERENCE TO SINADOXA AND TETRADOXA (ADOXACEAE) MICHAEL J. DONOGHUE,1 TORSTEN ERIKSSON,2 PATRICK A. REEVES,3 AND RICHARD G. OLMSTEAD 3 Abstract. To further clarify phylogenetic relationships within Dipsacales,we analyzed new and previously pub- lished rbcL sequences, alone and in combination with morphological data. We also examined relationships within Adoxaceae using rbcL and nuclear ribosomal internal transcribed spacer (ITS) sequences. We conclude from these analyses that Dipsacales comprise two major lineages:Adoxaceae and Caprifoliaceae (sensu Judd et al.,1994), which both contain elements of traditional Caprifoliaceae.Within Adoxaceae, the following relation- ships are strongly supported: (Viburnum (Sambucus (Sinadoxa (Tetradoxa, Adoxa)))). Combined analyses of C ap ri foliaceae yield the fo l l ow i n g : ( C ap ri folieae (Diervilleae (Linnaeeae (Morinaceae (Dipsacaceae (Triplostegia,Valerianaceae)))))). On the basis of these results we provide phylogenetic definitions for the names of several major clades. Within Adoxaceae, Adoxina refers to the clade including Sinadoxa, Tetradoxa, and Adoxa.This lineage is marked by herbaceous habit, reduction in the number of perianth parts,nectaries of mul- ticellular hairs on the perianth,and bifid stamens. The clade including Morinaceae,Valerianaceae, Triplostegia, and Dipsacaceae is here named Valerina. Probable synapomorphies include herbaceousness,presence of an epi- calyx (lost or modified in Valerianaceae), reduced endosperm,and distinctive chemistry, including production of monoterpenoids. The clade containing Valerina plus Linnaeeae we name Linnina. This lineage is distinguished by reduction to four (or fewer) stamens, by abortion of two of the three carpels,and possibly by supernumerary inflorescences bracts. Keywords: Adoxaceae, Caprifoliaceae, Dipsacales, ITS, morphological characters, phylogeny, phylogenetic taxonomy, phylogenetic nomenclature, rbcL, Sinadoxa, Tetradoxa. -

Phylogenomic Approach
Toward the ultimate phylogeny of Magnoliaceae: phylogenomic approach Sangtae Kim*1, Suhyeon Park1, and Jongsun Park2 1 Sungshin University, Korea 2 InfoBoss Co., Korea Mr. Carl Ferris Miller Founder of Chollipo Arboretum in Korea Chollipo Arboretum Famous for its magnolia collection 2020. Annual Meeting of Magnolia Society International Cholliop Arboretum in Korea. April 13th~22th, 2020 http://WWW.Chollipo.org Sungshin University, Seoul, Korea Dr. Hans Nooteboom Dr. Liu Yu-Hu Twenty-one years ago... in 1998 The 1st International Symposium on the Family Magnoliaceae, Gwangzhow Dr. Hiroshi Azuma Mr. Richard Figlar Dr. Hans Nooteboom Dr. Qing-wen Zeng Dr. Weibang Sun Handsome young boy Dr. Yong-kang Sima Dr. Yu-wu Law Presented ITS study on Magnoliaceae - never published Ten years ago... in 2009 Presented nine cp genome region study (9.2 kbp) on Magnoliaceae – published in 2013 2015 1st International Sympodium on Neotropical Magnoliaceae Gadalajara, 2019 3rd International Sympodium and Workshop on Neotropical Magnoliaceae Asterales Dipsacales Apiales Why magnolia study is Aquifoliales Campanulids (Euasterids II) Garryales Gentianales Laminales Solanales Lamiids important in botany? Ericales Asterids (Euasterids I) Cornales Sapindales Malvales Brassicales Malvids Fagales (Eurosids II) • As a member of early-diverging Cucurbitales Rosales Fabales Zygophyllales Celestrales Fabids (Eurosid I) angiosperms, reconstruction of the Oxalidales Malpighiales Vitales Geraniales Myrtales Rosids phylogeny of Magnoliaceae will Saxifragales Caryphyllales -

Anatomy and Affinities of Penthorum Melanie Lynn Haskins
University of Richmond UR Scholarship Repository Biology Faculty Publications Biology 2-1987 Anatomy and Affinities of Penthorum Melanie Lynn Haskins W. John Hayden University of Richmond, [email protected] Follow this and additional works at: http://scholarship.richmond.edu/biology-faculty-publications Part of the Botany Commons, Other Plant Sciences Commons, and the Plant Biology Commons Recommended Citation Haskins, Melanie Lynn, and W. John Hayden. "Anatomy and Affinities of Penthorum." American Journal of Botany 74, no. 2 (February 1987): 164-77. This Article is brought to you for free and open access by the Biology at UR Scholarship Repository. It has been accepted for inclusion in Biology Faculty Publications by an authorized administrator of UR Scholarship Repository. For more information, please contact [email protected]. Amer. J. Bot. 74(2): 164-177. 1987. ANATOMY AND AFFINITIES OF PENTHORUM' MELANIE L. HASKINS AND W. JOHN HAYDEN Department of Biology, University of Richmond, Richmond, Virginia 23173 ABSTRACT The genus Penthorum L. consists of two species of perennial herbs, P. sedoides of eastern North America and P. chinense ofeastern Asia. Pentho rum has long been considered intermediate between Crassulaceae and Saxifragaceae. An anatomical study of both species was undertaken to contribute to a better understanding of the relationships ofthese plants. Prominent anatomical features of Penthorum include: an aerenchymatous cortex and closely-spaced collateral vascular bundles of stems; one-trace unilacunar nodes; brochidodromous venation, rosoid teeth bearing hydathodes, and anomocytic stomata of leaves; angular vessel elements with many-barred scalariform perforation plates and alternate to scattered intervascular pits; thin-walled non septate fiber-tracheids; abundant homocellular erect uniseriate and biseriate rays; and absence of axial xylem parenchyma. -
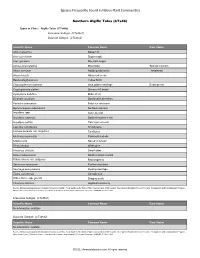
Algific Talus (Cts46)
Species Frequently Found in Native Plant Communities Southern Algific Talus (CTs46) Types in Class: Algific Talus (CTs46a) Limestone Subtype (CTs46a1) Dolomite Subtype (CTs46a2) Scientific Name Column1 Common Name Rare Status Abies balsamea Balsam fir Acer saccharum Sugar maple Acer spicatum Mountain maple Adoxa moschatellina Moschatel Special Concern Allium cernuum Nodding wild onion Threatened Arabis hirsuta Hairy rock cress Betula alleghaniensis Yellow birch Chrysosplenium iowense Iowa golden saxifrage Endangered Cryptogramma stelleri Slender cliff brake Cystopteris bulbifera Bulblet fern Dicentra cucullaria Dutchman's breeches Enemion biternatum False rue anemone Gymnocarpium robertianum Northern oak fern Impatiens spp. touch-me-not Impatiens capensis Spotted touch-me-not Impatiens pallida Pale touch-me-not Laportea canadensis Wood nettle Linnaea borealis var. longiflora Twinflower Mertensia paniculata Panicled bluebells Mitella nuda Naked miterwort Pinus strobus White pine Rhamnus alnifolia Dwarf alder Ribes hudsonianum Northern black currant Rubus idaeus var. strigosus Red raspberry Sambucus racemosa Red-berried elder Saxifraga pensylvanica Swamp saxifrage Taxus canadensis Canada yew Urtica dioica ssp. gracilis Stinging nettle Viburnum trilobum Highbush cranberry Source: Minnesota Department of Natural Resources (2005). Field Guide to the Native Plant Communities of Minnesota: The Eastern Broadleaf Forest Province. Ecological Land Classification Program, Minnesota County Biological Survey, and Natural Heritage and Nongame Research Program. MNDNR St. Paul, MN. Limestone Subtype (CTs46a1) Scientific Name Column1 Common Name Rare Status No information available Dolomite Subtype (CTs46a2) Scientific Name Column1 Common Name Rare Status No information available Source: Minnesota Department of Natural Resources (2005). Field Guide to the Native Plant Communities of Minnesota: The Eastern Broadleaf Forest Province. Ecological Land Classification Program, Minnesota County Biological Survey, and Natural Heritage and Nongame Research Program.