Anaerococcus Provenciensis Sp
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genomics 98 (2011) 370–375
Genomics 98 (2011) 370–375 Contents lists available at ScienceDirect Genomics journal homepage: www.elsevier.com/locate/ygeno Whole-genome comparison clarifies close phylogenetic relationships between the phyla Dictyoglomi and Thermotogae Hiromi Nishida a,⁎, Teruhiko Beppu b, Kenji Ueda b a Agricultural Bioinformatics Research Unit, Graduate School of Agricultural and Life Sciences, University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan b Life Science Research Center, College of Bioresource Sciences, Nihon University, Fujisawa, Japan article info abstract Article history: The anaerobic thermophilic bacterial genus Dictyoglomus is characterized by the ability to produce useful Received 2 June 2011 enzymes such as amylase, mannanase, and xylanase. Despite the significance, the phylogenetic position of Accepted 1 August 2011 Dictyoglomus has not yet been clarified, since it exhibits ambiguous phylogenetic positions in a single gene Available online 7 August 2011 sequence comparison-based analysis. The number of substitutions at the diverging point of Dictyoglomus is insufficient to show the relationships in a single gene comparison-based analysis. Hence, we studied its Keywords: evolutionary trait based on whole-genome comparison. Both gene content and orthologous protein sequence Whole-genome comparison Dictyoglomus comparisons indicated that Dictyoglomus is most closely related to the phylum Thermotogae and it forms a Bacterial systematics monophyletic group with Coprothermobacter proteolyticus (a constituent of the phylum Firmicutes) and Coprothermobacter proteolyticus Thermotogae. Our findings indicate that C. proteolyticus does not belong to the phylum Firmicutes and that the Thermotogae phylum Dictyoglomi is not closely related to either the phylum Firmicutes or Synergistetes but to the phylum Thermotogae. © 2011 Elsevier Inc. -

Table S4. Phylogenetic Distribution of Bacterial and Archaea Genomes in Groups A, B, C, D, and X
Table S4. Phylogenetic distribution of bacterial and archaea genomes in groups A, B, C, D, and X. Group A a: Total number of genomes in the taxon b: Number of group A genomes in the taxon c: Percentage of group A genomes in the taxon a b c cellular organisms 5007 2974 59.4 |__ Bacteria 4769 2935 61.5 | |__ Proteobacteria 1854 1570 84.7 | | |__ Gammaproteobacteria 711 631 88.7 | | | |__ Enterobacterales 112 97 86.6 | | | | |__ Enterobacteriaceae 41 32 78.0 | | | | | |__ unclassified Enterobacteriaceae 13 7 53.8 | | | | |__ Erwiniaceae 30 28 93.3 | | | | | |__ Erwinia 10 10 100.0 | | | | | |__ Buchnera 8 8 100.0 | | | | | | |__ Buchnera aphidicola 8 8 100.0 | | | | | |__ Pantoea 8 8 100.0 | | | | |__ Yersiniaceae 14 14 100.0 | | | | | |__ Serratia 8 8 100.0 | | | | |__ Morganellaceae 13 10 76.9 | | | | |__ Pectobacteriaceae 8 8 100.0 | | | |__ Alteromonadales 94 94 100.0 | | | | |__ Alteromonadaceae 34 34 100.0 | | | | | |__ Marinobacter 12 12 100.0 | | | | |__ Shewanellaceae 17 17 100.0 | | | | | |__ Shewanella 17 17 100.0 | | | | |__ Pseudoalteromonadaceae 16 16 100.0 | | | | | |__ Pseudoalteromonas 15 15 100.0 | | | | |__ Idiomarinaceae 9 9 100.0 | | | | | |__ Idiomarina 9 9 100.0 | | | | |__ Colwelliaceae 6 6 100.0 | | | |__ Pseudomonadales 81 81 100.0 | | | | |__ Moraxellaceae 41 41 100.0 | | | | | |__ Acinetobacter 25 25 100.0 | | | | | |__ Psychrobacter 8 8 100.0 | | | | | |__ Moraxella 6 6 100.0 | | | | |__ Pseudomonadaceae 40 40 100.0 | | | | | |__ Pseudomonas 38 38 100.0 | | | |__ Oceanospirillales 73 72 98.6 | | | | |__ Oceanospirillaceae -
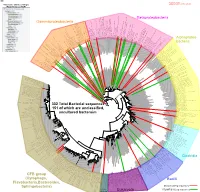
Bacteria Clostridia Bacilli Eukaryota CFB Group
AM935842.1.1361 uncultured Burkholderiales bacterium Class Betaproteobacteria AY283260.1.1552 Alcaligenes sp. PCNB−2 Class Betaproteobacteria AM934953.1.1374 uncultured Burkholderiales bacterium Class Betaproteobacteria AJ581593.1.1460 uncultured betaAM936569.1.1351 proteobacterium uncultured Class Betaproteobacteria Derxia sp. Class Betaproteobacteria AJ581621.1.1418 uncultured beta proteobacterium Class Betaproteobacteria DQ248272.1.1498 uncultured soil bacterium soil uncultured DQ248272.1.1498 DQ248235.1.1498 uncultured soil bacterium RS49 DQ248270.1.1496 uncultured soil bacterium DQ256489.1.1211 Variovorax paradoxus Class Betaproteobacteria Class paradoxus Variovorax DQ256489.1.1211 AF523053.1.1486 uncultured Comamonadaceae bacterium Class Betaproteobacteria AY706442.1.1396 uncultured bacterium uncultured AY706442.1.1396 AJ536763.1.1422 uncultured bacterium CS000359.1.1530 Variovorax paradoxus Class Betaproteobacteria Class paradoxus Variovorax CS000359.1.1530 AY168733.1.1411 uncultured bacterium AJ009470.1.1526 uncultured bacterium SJA−62 Class Betaproteobacteria Class SJA−62 bacterium uncultured AJ009470.1.1526 AY212561.1.1433 uncultured bacterium D16212.1.1457 Rhodoferax fermentans Class Betaproteobacteria Class fermentans Rhodoferax D16212.1.1457 AY957894.1.1546 uncultured bacterium AJ581620.1.1452 uncultured beta proteobacterium Class Betaproteobacteria RS76 AY625146.1.1498 uncultured bacterium RS65 DQ316832.1.1269 uncultured beta proteobacterium Class Betaproteobacteria DQ404909.1.1513 uncultured bacterium uncultured DQ404909.1.1513 AB021341.1.1466 bacterium rM6 AJ487020.1.1500 uncultured bacterium uncultured AJ487020.1.1500 RS7 RS86RC AF364862.1.1425 bacterium BA128 Class Betaproteobacteria AY957931.1.1529 uncultured bacterium uncultured AY957931.1.1529 CP000884.723807.725332 Delftia acidovorans SPH−1 Class Betaproteobacteria AY957923.1.1520 uncultured bacterium uncultured AY957923.1.1520 RS18 AY957918.1.1527 uncultured bacterium uncultured AY957918.1.1527 AY945883.1.1500 uncultured bacterium AF526940.1.1489 uncultured Ralstonia sp. -

A Genomic Journey Through a Genus of Large DNA Viruses
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Virology Papers Virology, Nebraska Center for 2013 Towards defining the chloroviruses: a genomic journey through a genus of large DNA viruses Adrien Jeanniard Aix-Marseille Université David D. Dunigan University of Nebraska-Lincoln, [email protected] James Gurnon University of Nebraska-Lincoln, [email protected] Irina V. Agarkova University of Nebraska-Lincoln, [email protected] Ming Kang University of Nebraska-Lincoln, [email protected] See next page for additional authors Follow this and additional works at: https://digitalcommons.unl.edu/virologypub Part of the Biological Phenomena, Cell Phenomena, and Immunity Commons, Cell and Developmental Biology Commons, Genetics and Genomics Commons, Infectious Disease Commons, Medical Immunology Commons, Medical Pathology Commons, and the Virology Commons Jeanniard, Adrien; Dunigan, David D.; Gurnon, James; Agarkova, Irina V.; Kang, Ming; Vitek, Jason; Duncan, Garry; McClung, O William; Larsen, Megan; Claverie, Jean-Michel; Van Etten, James L.; and Blanc, Guillaume, "Towards defining the chloroviruses: a genomic journey through a genus of large DNA viruses" (2013). Virology Papers. 245. https://digitalcommons.unl.edu/virologypub/245 This Article is brought to you for free and open access by the Virology, Nebraska Center for at DigitalCommons@University of Nebraska - Lincoln. It has been accepted for inclusion in Virology Papers by an authorized administrator of DigitalCommons@University of Nebraska - Lincoln. Authors Adrien Jeanniard, David D. Dunigan, James Gurnon, Irina V. Agarkova, Ming Kang, Jason Vitek, Garry Duncan, O William McClung, Megan Larsen, Jean-Michel Claverie, James L. Van Etten, and Guillaume Blanc This article is available at DigitalCommons@University of Nebraska - Lincoln: https://digitalcommons.unl.edu/ virologypub/245 Jeanniard, Dunigan, Gurnon, Agarkova, Kang, Vitek, Duncan, McClung, Larsen, Claverie, Van Etten & Blanc in BMC Genomics (2013) 14. -

The Influence of Sodium Chloride on the Performance of Gammarus Amphipods and the Community Composition of Microbes Associated with Leaf Detritus
THE INFLUENCE OF SODIUM CHLORIDE ON THE PERFORMANCE OF GAMMARUS AMPHIPODS AND THE COMMUNITY COMPOSITION OF MICROBES ASSOCIATED WITH LEAF DETRITUS By Shelby McIlheran Leaf litter decomposition is a fundamental part of the carbon cycle and helps support aquatic food webs along with being an important assessment of the health of rivers and streams. Disruptions in this organic matter breakdown can signal problems in other parts of ecosystems. One disruption is rising chloride concentrations. Chloride concentrations are increasing in many rivers worldwide due to anthropogenic sources that can harm biota and affect ecosystem processes. Elevated chloride concentrations can lead to lethal or sublethal impacts. While many studies have shown that excessive chloride uptake impacts health (e.g. lowered respiration and growth rates) in a wide variety of aquatic organisms including microbes and benthic invertebrates). The impacts of high chloride concentrations on decomposers are less well understood. My research objective was to assess how increasing chloride concentrations affect the performance and diversity of decomposer organisms in freshwater systems. I experimentally manipulated chloride concentrations in microcosms containing leaves colonized by microbes or containing leaves, microbes and amphipods. Respiration rate, decomposition, and community composition of the microbes were measured along with the amphipod growth rate, egestion rate, and mortality. Elevated chloride concentration did not impact microbial respiration rates or leaf decomposition, but had large impacts on bacteria community composition. It did cause a decrease in instantaneous growth rate, and 100% mortality in the highest amphipod chloride treatment, but amphipod egestion rate was not significantly affected. The results of my research suggest that the widespread increases in chloride concentrations in rivers will have an impact on decomposer communities in these systems. -

UK Standards for Microbiology Investigations
UK Standards for Microbiology Investigations Identification of Anaerobic Cocci Issued by the Standards Unit, Microbiology Services, PHE Bacteriology – Identification | ID 14 | Issue no: 3 | Issue date: 04.02.15 | Page: 1 of 29 © Crown copyright 2015 Identification of Anaerobic Cocci Acknowledgments UK Standards for Microbiology Investigations (SMIs) are developed under the auspices of Public Health England (PHE) working in partnership with the National Health Service (NHS), Public Health Wales and with the professional organisations whose logos are displayed below and listed on the website https://www.gov.uk/uk- standards-for-microbiology-investigations-smi-quality-and-consistency-in-clinical- laboratories. SMIs are developed, reviewed and revised by various working groups which are overseen by a steering committee (see https://www.gov.uk/government/groups/standards-for-microbiology-investigations- steering-committee). The contributions of many individuals in clinical, specialist and reference laboratories who have provided information and comments during the development of this document are acknowledged. We are grateful to the Medical Editors for editing the medical content. For further information please contact us at: Standards Unit Microbiology Services Public Health England 61 Colindale Avenue London NW9 5EQ E-mail: [email protected] Website: https://www.gov.uk/uk-standards-for-microbiology-investigations-smi-quality- and-consistency-in-clinical-laboratories UK Standards for Microbiology Investigations are produced in association with: Logos correct at time of publishing. Bacteriology – Identification | ID 14 | Issue no: 3 | Issue date: 04.02.15 | Page: 2 of 29 UK Standards for Microbiology Investigations | Issued by the Standards Unit, Public Health England Identification of Anaerobic Cocci Contents ACKNOWLEDGMENTS ......................................................................................................... -

Db20-0503.Full.Pdf
Page 1 of 32 Diabetes Analysis of the Composition and Functions of the Microbiome in Diabetic Foot Osteomyelitis based on 16S rRNA and Metagenome Sequencing Technology Zou Mengchen1*; Cai Yulan2*; Hu Ping3*; Cao Yin1; Luo Xiangrong1; Fan Xinzhao1; Zhang Bao4; Wu Xianbo4; Jiang Nan5; Lin Qingrong5; Zhou Hao6; Xue Yaoming1; Gao Fang1# 1Department of Endocrinology and Metabolism, Nanfang Hospital, Southern Medical University, Guangzhou, China 2Department of Endocrinology, Affiliated Hospital of Zunyi Medical College, Zunyi, China 3Department of Geriatric Medicine, Xiaogan Central Hospital, Xiaogan, China 4School of Public Health and Tropic Medicine, Southern Medical University, Guangzhou, China 5Department of Orthopaedics & Traumatology, Nanfang Hospital, Southern Medical University, Guangzhou, China 6Department of Hospital Infection Management of Nanfang Hospital, Southern Medical University, Guangzhou, China *Zou mengchen, Cai yulan and Hu ping contributed equally to this work. Running title: Microbiome of Diabetic Foot Osteomyelitis Word count: 3915 Figures/Tables Count: 4Figures / 3 Tables References: 26 Diabetes Publish Ahead of Print, published online August 14, 2020 Diabetes Page 2 of 32 Keywords: diabetic foot osteomyelitis; microbiome; 16S rRNA sequencing; metagenome sequencing #Corresponding author: Gao Fang, E-mail: [email protected], Tel: 13006871226 Page 3 of 32 Diabetes ABSTRACT Metagenome sequencing has not been used in infected bone specimens. This study aimed to analyze the microbiome and its functions. This prospective observational study explored the microbiome and its functions of DFO (group DM) and posttraumatic foot osteomyelitis (PFO) (group NDM) based on 16S rRNA sequencing and metagenome sequencing technologies. Spearman analysis was used to explore the correlation between dominant species and clinical indicators of patients with DFO. -

Use of Mass Spectometry for the Identification of Anaerobic Bacteria, Change Anaerobic Bacteriology?
Use of mass spectometry for the identification of anaerobic bacteria, change anaerobic bacteriology? University Medical Center Groningen Department Medical Microbiology and Infection Prevention A.C.M. Veloo Expert Center Anaerobic Infections The Netherlands www.mmb-umcg.eu ESCMID eLibrary © by author Workflow MALDI-TOF MS Direct spotting of bacteria on target using a toothpick Add HCCA matrix Data acquisition Data analyses Log score: <1.7 no reliable identification 1.7 – 2.0 identification with low confidence ≥ 2.0 ESCMIDidentification with high confidence eLibrary © by author Anaerobic culture Phenotypic pure culture primary incubation 2 days 2-7 days aerotolerance identification MALDI-TOF MS 1 day 2-14 days primary incubation 2-7 days MALDI-TOF MS testing minutes ESCMID eLibrary © by author ESCMIDThe reliability of the identification eLibrary obtained with MALDI- TOF MS is superior over phenotypic methods © by author Situation 2012/2013 1. ± 60-70 % of the anaerobes can be identified by MALDI-TOF MS 2. Performance differs per species/genus 3. MALDI-TOF MS database needs optimization for the identification of anaerobic bacteria - addition of spectra to existing database increases performance - reference strains - clinical isolates ESCMID eLibrary © by author Example from literature Species identification of clinical Prevotella isolates by Matrix-Assisted Laser Desorption Ionization-Time of Flight mass spectrometry Wybo et al. J. Clin. Microbiol. 2012; 50:1415-1418 Commercial reference database: Commercial database + house made MSPs: - 63 % correct species identification - 83 % correct species identification - 11 % correct genus identification - 6 % correct genus identification - 26 % no identification - 11 % no identification ESCMID14 % due to the addition of MSPs of species eLibrary not yet present in database 6 % due to the addition of MSPs of species already present in database © by author ENRIA European Network for the Rapid Identification of Anaerobes Lead: Prof. -

Foot Ulcers: Characterising the Microbiome of Ulcers Karen Smith1, Andrew Collier2, Eleanor M
Smith et al. BMC Microbiology (2016) 16:54 DOI 10.1186/s12866-016-0665-z RESEARCH ARTICLE Open Access One step closer to understanding the role of bacteria in diabetic foot ulcers: characterising the microbiome of ulcers Karen Smith1, Andrew Collier2, Eleanor M. Townsend1,3, Lindsay E. O’Donnell3, Abhijit M. Bal2, John Butcher1, William G. Mackay1, Gordon Ramage3* and Craig Williams1 Abstract Background: The aim of this study was to characterise the microbiome of new and recurrent diabetic foot ulcers using 16S amplicon sequencing (16S AS), allowing the identification of a wider range of bacterial species that may be important in the development of chronicity in these debilitating wounds. Twenty patients not receiving antibiotics for the past three months were selected, with swabs taken from each individual for culture and 16S AS. DNA was isolated using a combination of bead beating and kit extraction. Samples were sequenced on the Illumina Hiseq 2500 platform. Results: Conventional laboratory culture showed positive growth from only 55 % of the patients, whereas 16S AS was positive for 75 % of the patients (41 unique genera, representing 82 different operational taxonomic units (OTU’s). S. aureus was isolated in 72 % of culture-positive samples, whereas the most commonly detected bacteria in all ulcers were Peptoniphilus spp., Anaerococcus spp. and Corynebacterium spp., with the addition of Staphylococcus spp. in new ulcers. The majority of OTU’s residing in both new and recurrent ulcers (over 67 %) were identified as facultative or strict anaerobic Gram-positive organisms. Principal component analysis (PCA) showed no difference in clustering between the two groups (new and recurrent ulcers). -

Microbial and Mineralogical Characterizations of Soils Collected from the Deep Biosphere of the Former Homestake Gold Mine, South Dakota
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln US Department of Energy Publications U.S. Department of Energy 2010 Microbial and Mineralogical Characterizations of Soils Collected from the Deep Biosphere of the Former Homestake Gold Mine, South Dakota Gurdeep Rastogi South Dakota School of Mines and Technology Shariff Osman Lawrence Berkeley National Laboratory Ravi K. Kukkadapu Pacific Northwest National Laboratory, [email protected] Mark Engelhard Pacific Northwest National Laboratory Parag A. Vaishampayan California Institute of Technology See next page for additional authors Follow this and additional works at: https://digitalcommons.unl.edu/usdoepub Part of the Bioresource and Agricultural Engineering Commons Rastogi, Gurdeep; Osman, Shariff; Kukkadapu, Ravi K.; Engelhard, Mark; Vaishampayan, Parag A.; Andersen, Gary L.; and Sani, Rajesh K., "Microbial and Mineralogical Characterizations of Soils Collected from the Deep Biosphere of the Former Homestake Gold Mine, South Dakota" (2010). US Department of Energy Publications. 170. https://digitalcommons.unl.edu/usdoepub/170 This Article is brought to you for free and open access by the U.S. Department of Energy at DigitalCommons@University of Nebraska - Lincoln. It has been accepted for inclusion in US Department of Energy Publications by an authorized administrator of DigitalCommons@University of Nebraska - Lincoln. Authors Gurdeep Rastogi, Shariff Osman, Ravi K. Kukkadapu, Mark Engelhard, Parag A. Vaishampayan, Gary L. Andersen, and Rajesh K. Sani This article is available at DigitalCommons@University of Nebraska - Lincoln: https://digitalcommons.unl.edu/ usdoepub/170 Microb Ecol (2010) 60:539–550 DOI 10.1007/s00248-010-9657-y SOIL MICROBIOLOGY Microbial and Mineralogical Characterizations of Soils Collected from the Deep Biosphere of the Former Homestake Gold Mine, South Dakota Gurdeep Rastogi & Shariff Osman & Ravi Kukkadapu & Mark Engelhard & Parag A. -

Multicentre Study of the Burden of Multidrug-Resistant Bacteria in the Aetiology of Infected Diabetic Foot Ulcers
African Journal of Laboratory Medicine ISSN: (Online) 2225-2010, (Print) 2225-2002 Page 1 of 10 Original Research Multicentre study of the burden of multidrug-resistant bacteria in the aetiology of infected diabetic foot ulcers Authors: Background: Infected diabetic foot ulcer (IDFU) is a public health issue and the leading cause 1 Adeyemi T. Adeyemo of non-traumatic limb amputation. Very few published data on IDFU exist in most West Babatope Kolawole2 Vincent O. Rotimi3 African countries. Aaron O. Aboderin1,4 Objective: The study investigated the aetiology and antibacterial drug resistance burden of Affiliations: IDFU in tertiary hospitals in Osun state, Nigeria, between July 2016 and April 2017. 1Department of Medical Microbiology and Methods: Isolates were cultured from tissue biopsies or aspirates collected from patients with Parasitology, Obafemi IDFU. Bacterial identification, antibiotic susceptibility testing and phenotypic detection of Awolowo University Teaching extended-spectrum beta-lactamase and carbapenemase production were done by established Hospitals, Ile-Ife, Nigeria protocols. Specific resistance genes were detected by polymerase chain reaction. 2 Department of Medicine, Results: There were 218 microorganisms isolated from 93 IDFUs, comprising 129 (59.2%) Faculty of Clinical Science, Gram-negative bacilli (GNB), 59 (27.1 ) Gram-positive cocci and 29 (13.3 ) anaerobic bacteria. Obafemi Awolowo University, % % Ile-Ife, Nigeria The top five facultative anaerobic bacteria isolated were:Staphylococcus aureus (34; 15.6%), Escherichia coli (23; 10.6%), Pseudomonas aeruginosa (20; 9.2%), Klebsiella pneumoniae (19; 8.7%) 3Department of Microbiology, and Citrobacter spp. (19; 8.7%). The most common anaerobes were Bacteroides spp. (7; 3.2%) Faculty of Medicine, Kuwait and Peptostreptococcus anaerobius (6; 2.8%). -

Grappling Extremes: Molecular Methods Combined with Cultivation
Grappling extremes: Molecular methods combined with cultivation reveal the composition and biology of space- relevant microbial communities Dissertation zur Erlangung des Doktorgrades der Naturwissenschaften (Dr.rer.nat.) der Fakultät für Biologie und Vorklinische Medizin der Universität Regensburg vorgelegt von Alexandra Kristin Perras aus München im Jahr 2017 Das Promotionsgesuch wurde eingereicht am: 13.01.2017 Diese Arbeit wurde angeleitet von: Prof. Dr. Christine Moissl-Eichinger Prüfungsausschuss: Vorsitzender Prof. Dr. Peter Poschlod 1.Gutachter: Prof. Dr. Christine Moissl-Eichinger 2.Gutachter: Prof. Dr. Reinhard Wirth 3. Prüfer: Prof. Dr. Rainer Merkl Unterschrift: Alexandra K. Perras - 2 - Grappling extremes: Molecular methods combined with cultivation reveal the composition and biology of space- relevant microbial communities Dissertation for achieving the doctoral degree of natural sciences (Dr.rer.nat.) at the faculty of Biology and Preclinical Medicine at the University of Regensburg by Alexandra Kristin Perras from Munich 2017 Key words: Mars-analogues, microbial ecology, Archaea, Candidatus Altiarchaeum hamiconexum, hami, extreme environments, the ISS microbiome, MASE - 3 - The dissertation was performed under the supervision of Prof. Dr. Christine Moissl-Eichinger (PhD supervisor) Prof. Dr. Reinhard Wirth (1st mentor) Prof. Dr. Charles Cockell (2nd mentor) At the Naturwissenschaftliche Fakultät 3, Biology and Preclinical Medicine and the Medical University of Graz under fulfilment of all requirements of the Regensburger