UK Standards for Microbiology Investigations
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Number Genera
Table 4. Location of the various genes Modified April 28, 2021 Originally modified from AAC 1999 43:2823-30 with permission from ASM Journals Gene Number Genera METHYLASES e erm(A) 11 AggregatibacterL,1, Bacteroides, Enterococcus , Haemophilusr, Helcococcus, Klebsiella, a a Listeria, Peptostreptococcus , Prevotella , Staphylococcus, Streptococcus erm(B) 39 Aggregatibacter1, Aerococcus, Bacillus, Bacteroidesa, Campylobacteraf, a Citrobacter, Corynebacterium, Clostridium , Clostridioides, Enterobacter, Escherichia, a a Eubacterium , Enterococcus, Fusobacterium , Gallibacteriumcp, Gemella, Haemophilus, Klebsiella, a Lactobacillus, Listeriacc, Micrococcus, Neisseria, Pantoea, Pediococcus, Peptostreptococcus , a a Porphyromonas , Proteus, Pseudomonas, Rothia, Ruminococcus , Salmonellacn, Serratia, Shigella ag, Staphylococcus, Streptococcus, UreaplasmaO, Wollinellaa, Treponemab, Trueperella1 a erm(C) 38 Aeromonasy, AggregatibacterL, Actinomyces, Bacillus, Bacteroides , Brevundimonas y, Burkholderia y, Campylobacter, Capnocytophagaa,ca ,Chryseomonasy, ,Corynebacterium, n a Clostridiuma,n Escherichia , Eubacterium , Enterococcus, Fusobacteriuma,br, Haemophilusr, y y Lactobacillus, Listeria, Macrococcus, Micrococcus, Neisseria, Paenibacillus , Pasteurella , a a Prevotella , Peptostreptococcus , Pseudomonasy, Pseudoramibactera,br, Rhizobiumy, Salmonella, y y y Serratia, Sinorhizobium , Sphingomonas , Stenotrophomans , Staphylococcus, Streptococcus, a Trueperella n, Wolinella erm(D) 2 Bacillus, Salmonella a a a a n erm(E) 7 Bacteroides , Eubacterium -

Early Photosynthetic Eukaryotes Inhabited Low-Salinity Habitats
Early photosynthetic eukaryotes inhabited PNAS PLUS low-salinity habitats Patricia Sánchez-Baracaldoa,1, John A. Ravenb,c, Davide Pisanid,e, and Andrew H. Knollf aSchool of Geographical Sciences, University of Bristol, Bristol BS8 1SS, United Kingdom; bDivision of Plant Science, University of Dundee at the James Hutton Institute, Dundee DD2 5DA, United Kingdom; cPlant Functional Biology and Climate Change Cluster, University of Technology Sydney, Ultimo, NSW 2007, Australia; dSchool of Biological Sciences, University of Bristol, Bristol BS8 1TH, United Kingdom; eSchool of Earth Sciences, University of Bristol, Bristol BS8 1TH, United Kingdom; and fDepartment of Organismic and Evolutionary Biology, Harvard University, Cambridge, MA 02138 Edited by Peter R. Crane, Oak Spring Garden Foundation, Upperville, Virginia, and approved July 7, 2017 (received for review December 7, 2016) The early evolutionary history of the chloroplast lineage remains estimates for the origin of plastids ranging over 800 My (7). At the an open question. It is widely accepted that the endosymbiosis that same time, the ecological setting in which this endosymbiotic event established the chloroplast lineage in eukaryotes can be traced occurred has not been fully explored (8), partly because of phy- back to a single event, in which a cyanobacterium was incorpo- logenetic uncertainties and preservational biases of the fossil re- rated into a protistan host. It is still unclear, however, which cord. Phylogenomics and trait evolution analysis have pointed to a Cyanobacteria are most closely related to the chloroplast, when the freshwater origin for Cyanobacteria (9–11), providing an approach plastid lineage first evolved, and in what habitats this endosym- to address the early diversification of terrestrial biota for which the biotic event occurred. -

Genomics 98 (2011) 370–375
Genomics 98 (2011) 370–375 Contents lists available at ScienceDirect Genomics journal homepage: www.elsevier.com/locate/ygeno Whole-genome comparison clarifies close phylogenetic relationships between the phyla Dictyoglomi and Thermotogae Hiromi Nishida a,⁎, Teruhiko Beppu b, Kenji Ueda b a Agricultural Bioinformatics Research Unit, Graduate School of Agricultural and Life Sciences, University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo 113-8657, Japan b Life Science Research Center, College of Bioresource Sciences, Nihon University, Fujisawa, Japan article info abstract Article history: The anaerobic thermophilic bacterial genus Dictyoglomus is characterized by the ability to produce useful Received 2 June 2011 enzymes such as amylase, mannanase, and xylanase. Despite the significance, the phylogenetic position of Accepted 1 August 2011 Dictyoglomus has not yet been clarified, since it exhibits ambiguous phylogenetic positions in a single gene Available online 7 August 2011 sequence comparison-based analysis. The number of substitutions at the diverging point of Dictyoglomus is insufficient to show the relationships in a single gene comparison-based analysis. Hence, we studied its Keywords: evolutionary trait based on whole-genome comparison. Both gene content and orthologous protein sequence Whole-genome comparison Dictyoglomus comparisons indicated that Dictyoglomus is most closely related to the phylum Thermotogae and it forms a Bacterial systematics monophyletic group with Coprothermobacter proteolyticus (a constituent of the phylum Firmicutes) and Coprothermobacter proteolyticus Thermotogae. Our findings indicate that C. proteolyticus does not belong to the phylum Firmicutes and that the Thermotogae phylum Dictyoglomi is not closely related to either the phylum Firmicutes or Synergistetes but to the phylum Thermotogae. © 2011 Elsevier Inc. -

Growth and Grazing Rates of the Herbivorous Dinoflagellate Gymnodinium Sp
MARINE ECOLOGY PROGRESS SERIES Published December 16 Mar. Ecol. Prog. Ser. Growth and grazing rates of the herbivorous dinoflagellate Gymnodinium sp. from the open subarctic Pacific Ocean Suzanne L. Strom' School of Oceanography WB-10, University of Washington. Seattle. Washington 98195, USA ABSTRACT: Growth, grazing and cell volume of the small heterotroph~cdinoflagellate Gyrnnodin~um sp. Isolated from the open subarctic Pacific Ocean were measured as a funct~onof food concentration using 2 phytoplankton food species. Growth and lngestlon rates increased asymptotically with Increas- ing phytoplankon food levels, as did grazer cell volume; rates at representative oceanic food levels were high but below maxima. Clearance rates decreased with lncreaslng food levels when Isochrysis galbana was the food source; they increased ~vithlncreaslng food levels when Synechococcus sp. was the food source. There was apparently a grazlng threshold for Ingestion of Synechococcus: below an initial Synechococcus concentration of 20 pgC 1.' ingestion rates on this alga were very low, while above this initial concentratlon Synechococcus was grazed preferent~ally Gross growth efficiency varied between 0.03 and 0.53 (mean 0.21) and was highest at low food concentrations. Results support the hypothesis that heterotrophic d~noflagellatesmay contribute to controlling population increases of small, rap~dly-grow~ngphytoplankton specles even at low oceanic phytoplankton concentrations. INTRODUCTION as Gymnodinium and Gyrodinium is difficult or impos- sible using older preservation and microscopy tech- Heterotrophic dinoflagellates can be a significant niques; experimental emphasis has been on more component of the microzooplankton in marine waters. easily recognizable and collectable microzooplankton In the oceanic realm, Lessard (1984) and Shapiro et al. -

Metatranscriptomic Analysis Reveals Active Bacterial Communities in Diabetic Foot Infections
fmicb-11-01688 July 20, 2020 Time: 12:22 # 1 ORIGINAL RESEARCH published: 22 July 2020 doi: 10.3389/fmicb.2020.01688 Metatranscriptomic Analysis Reveals Active Bacterial Communities in Diabetic Foot Infections Fatemah Sadeghpour Heravi1, Martha Zakrzewski2, Karen Vickery1, Matthew Malone3,4,5 and Honghua Hu1* 1 Surgical Infection Research Group, Faculty of Medicine and Health Sciences, Macquarie University, Sydney, NSW, Australia, 2 QIMR Berghofer Medical Research Institute, Brisbane, QLD, Australia, 3 Infectious Diseases and Microbiology, School of Medicine, Western Sydney University, Sydney, NSW, Australia, 4 Liverpool Hospital, South Western Sydney LHD, Sydney, NSW, Australia, 5 Liverpool Diabetes Collaborative Research Unit, Ingham Institute for Applied Medical Research, Sydney, NSW, Australia Despite the extended view of the composition of diabetic foot infections (DFIs), little is known about which transcriptionally active bacterial communities are pertinent to infection, and if any differences are associated with increased infection severity. We applied a RNA sequencing approach to analyze the composition, function, and pathogenicity of the active bacterial communities in DFIs. Taxonomic profiling of bacterial transcripts revealed the presence of 14 bacterial phyla in DFIs. The abundance Edited by: Qi Zhao, of the Spiroplasma, Vibrio, and Mycoplasma were significantly different in different Liaoning University, China infection severities (P < 0.05). Mild and severe stages of infections were dominated Reviewed by: by Staphylococcus aureus and Porphyromonas asaccharolytica, respectively. A total of Patrick R. M. Harnarayan, The University of the West Indies 132 metabolic pathways were identified of which ribosome and thiamin being among the at St. Augustine, Trinidad and Tobago most highly transcribed pathways. Moreover, a total of 131 antibiotic resistance genes, Maria José Saavedra, primarily involved in the multidrug efflux pumps/exporters, were identified. -

Table S4. Phylogenetic Distribution of Bacterial and Archaea Genomes in Groups A, B, C, D, and X
Table S4. Phylogenetic distribution of bacterial and archaea genomes in groups A, B, C, D, and X. Group A a: Total number of genomes in the taxon b: Number of group A genomes in the taxon c: Percentage of group A genomes in the taxon a b c cellular organisms 5007 2974 59.4 |__ Bacteria 4769 2935 61.5 | |__ Proteobacteria 1854 1570 84.7 | | |__ Gammaproteobacteria 711 631 88.7 | | | |__ Enterobacterales 112 97 86.6 | | | | |__ Enterobacteriaceae 41 32 78.0 | | | | | |__ unclassified Enterobacteriaceae 13 7 53.8 | | | | |__ Erwiniaceae 30 28 93.3 | | | | | |__ Erwinia 10 10 100.0 | | | | | |__ Buchnera 8 8 100.0 | | | | | | |__ Buchnera aphidicola 8 8 100.0 | | | | | |__ Pantoea 8 8 100.0 | | | | |__ Yersiniaceae 14 14 100.0 | | | | | |__ Serratia 8 8 100.0 | | | | |__ Morganellaceae 13 10 76.9 | | | | |__ Pectobacteriaceae 8 8 100.0 | | | |__ Alteromonadales 94 94 100.0 | | | | |__ Alteromonadaceae 34 34 100.0 | | | | | |__ Marinobacter 12 12 100.0 | | | | |__ Shewanellaceae 17 17 100.0 | | | | | |__ Shewanella 17 17 100.0 | | | | |__ Pseudoalteromonadaceae 16 16 100.0 | | | | | |__ Pseudoalteromonas 15 15 100.0 | | | | |__ Idiomarinaceae 9 9 100.0 | | | | | |__ Idiomarina 9 9 100.0 | | | | |__ Colwelliaceae 6 6 100.0 | | | |__ Pseudomonadales 81 81 100.0 | | | | |__ Moraxellaceae 41 41 100.0 | | | | | |__ Acinetobacter 25 25 100.0 | | | | | |__ Psychrobacter 8 8 100.0 | | | | | |__ Moraxella 6 6 100.0 | | | | |__ Pseudomonadaceae 40 40 100.0 | | | | | |__ Pseudomonas 38 38 100.0 | | | |__ Oceanospirillales 73 72 98.6 | | | | |__ Oceanospirillaceae -

21 Pathogens of Harmful Microalgae
21 Pathogens of Harmful Microalgae RS. Salomon and I. Imai 2L1 Introduction Pathogens are any organisms that cause disease to other living organisms. Parasitism is an interspecific interaction where one species (the parasite) spends the whole or part of its life on or inside cells and tissues of another living organism (the host), from where it derives most of its food. Parasites that cause disease to their hosts are, by definition, pathogens. Although infection of metazoans by other metazoans and protists are the more fre quently studied, there are interactions where both host and parasite are sin gle-celled organisms. Here we describe such interactions involving microal gae as hosts. The aim of this chapter is to review the current status of research on pathogens of harmful microalgae and present future perspec tives within the field. Pathogens with the ability to impair and kill micro algae include viruses, bacteria, fungi and a number of protists (see reviews by Elbrachter and Schnepf 1998; Brussaard 2004; Park et al. 2004; Mayali and Azam 2004; Ibelings et al. 2004). Valuable information exists from non-harm ful microalgal hosts, and these studies will be referred to throughout the text. Nevertheless, emphasis is given to cases where hosts are recognizable harmful microalgae. 21.2 Viruses Viruses and virus-like particles (VLPs) have been found in more than 50 species of eukaryotic microalgae, and several of them have been isolated in laboratory cultures (Brussaard 2004; Nagasaki et al. 2005). These viruses are diverse both in size and genome type, and some of them infect harmful algal bloom (HAB)-causing species (Table 21.1). -

Infective Endocarditis Caused by C. Sordellii: the First Case Report from India
Published online: 2021-05-19 THIEME 74 C.Case sordellii Report in Endocarditis Chaudhry et al. Infective Endocarditis Caused by C. sordellii: The First Case Report from India Rama Chaudhry1 Tej Bahadur1 Tanu Sagar1 Sonu Kumari Agrawal1 Nazneen Arif1 Shiv K. Choudhary2 Nishant Verma1 1Department of Microbiology, All India Institute of Medical Address for correspondence Dr. Rama Chaudhry, MD, Department Sciences, New Delhi, India of Microbiology, All India Institute of Medical Sciences, Ansari Nagar, 2Department of Cardiothoracic and Vascular Surgery, All India New Delhi 110029, India (e-mail: [email protected]). Institute of Medical Sciences, New Delhi, India J Lab Physicians 2021;13:74–76. Abstract Clostridium sordellii is a gram-positive anaerobic bacteria most commonly isolated Keywords from skin and soft tissue infection, penetrating injurious and intravenous drug abus- ► infective endocarditis ers. The exotoxins produced by the bacteria are associated with toxic shock syndrome. ► Clostridium sordellii We report here a first case of infective endocarditis due to C. sordellii from a female ► diagnosis patient with ventricular septal defect from India. Introduction admitted to the cardiothoracic vascular surgery ward of All India Institute of Medical Sciences (AIIMS), New Delhi, India Anaerobes are major components of the normal micro- with complaints of worsening shortness of breath and pal- bial flora present on human skin and mucosa. Infections pitations. Patient reported having an episode of IE 3 months due to anaerobic bacteria are common, but they are dif- back for which she had been admitted to AIIMS and was ficult to isolate from infected sites and are often over- discharged after treatment. -

Responses of the Picoprasinophyte Micromonas Commoda to Light and Ultraviolet Stress
RESEARCH ARTICLE Responses of the picoprasinophyte Micromonas commoda to light and ultraviolet stress Marie L. Cuvelier1☯¤a³, Jian Guo1☯¤b³, Alejandra C. Ortiz1¤c, Marijke J. van Baren1, Muhammad Akram Tariq2¤d, FreÂdeÂric Partensky3, Alexandra Z. Worden1,4,5* 1 Monterey Bay Aquarium Research Institute (MBARI), Moss Landing, CA, United States of America, 2 Department of Biomolecular Engineering, University of California Santa Cruz, Santa Cruz, CA, United States of America, 3 Sorbonne UniversiteÂsÐUPMC Universite Paris 06, CNRS UMR, Station Biologique, CS, a1111111111 Roscoff, France, 4 Department of Ocean Sciences, University of California Santa Cruz, Santa Cruz, CA, a1111111111 United States of America, 5 Integrated Microbial Biodiversity Program, Canadian Institute for Advanced a1111111111 Research, Toronto, Canada a1111111111 a1111111111 ☯ These authors contributed equally to this work. ¤a Current address: Department of Biological Sciences, Nova Southeastern University, Fort Lauderdale, FL, United States of America ¤b Current address: Department of Molecular, Cell and Developmental Biology, University of California Santa Cruz, Santa Cruz, CA, United States of America ¤c Current address: Department of Geological Sciences, Indiana University Bloomington, Bloomington, IN, OPEN ACCESS United States of America ¤d Current address: School of Health Sciences, University of Management and Technology, Lahore, Citation: Cuvelier ML, Guo J, Ortiz AC, van Baren Pakistan MJ, Tariq MA, Partensky F, et al. (2017) ³ These authors are co-first authors on this work. Responses of the picoprasinophyte Micromonas * [email protected] commoda to light and ultraviolet stress. PLoS ONE 12(3): e0172135. doi:10.1371/journal. pone.0172135 Abstract Editor: Amanda M. Cockshutt, Mount Allison University, CANADA Micromonas is a unicellular marine green alga that thrives from tropical to polar ecosystems. -

Intraspecies Comparative Genomics of Rickettsia
AIX ͲMARSEILLE UNIVERSITÉ FACULTÉ DE MÉDECINE DE MARSEILLE ÉCOLE DOCTORALE DES SCIENCES DE LA VIE ET DE LA SANTÉ T H È S E Présentée et publiquement soutenue devant LA FACULTÉ DE MÉDECINE DE MARSEILLE Le 13 décembre 2013 Par M. Erwin SENTAUSA Né le 16 décembre 1979 àMalang, Indonésie INTRASPECIES COMPARATIVE GENOMICS OF RICKETTSIA Pour obtenir le grade de DOCTORAT d’AIX ͲMARSEILLE UNIVERSITÉ SPÉCIALITÉ :PATHOLOGIE HUMAINE Ͳ MALADIES INFECTIEUSES Membres du Jury de la Thèse : Dr. Patricia RENESTO Rapporteur Pr. Max MAURIN Rapporteur Dr. Florence FENOLLAR Membre du Jury Pr. Pierre ͲEdouard FOURNIER Directeur de thèse Unité de Recherche sur les Maladies Infectieuses et Tropicales Émergentes UM63, CNRS 7278, IRD 198, Inserm 1095 Avant Propos Le format de présentation de cette thèse correspond à une recommandation de la spécialité Maladies Infectieuses et Microbiologie, à l’intérieur du Master de Sciences de la Vie et de la Santé qui dépend de l’Ecole Doctorale des Sciences de la Vie de Marseille. Le candidat est amené àrespecter des règles qui lui sont imposées et qui comportent un format de thèse utilisé dans le Nord de l’Europe permettant un meilleur rangement que les thèses traditionnelles. Par ailleurs, la partie introduction et bibliographie est remplacée par une revue envoyée dans un journal afin de permettre une évaluation extérieure de la qualité de la revue et de permettre àl’étudiant de le commencer le plus tôt possible une bibliographie exhaustive sur le domaine de cette thèse. Par ailleurs, la thèse est présentée sur article publié, accepté ou soumis associé d’un bref commentaire donnant le sens général du travail. -
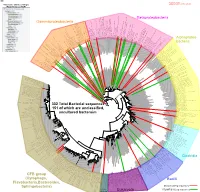
Bacteria Clostridia Bacilli Eukaryota CFB Group
AM935842.1.1361 uncultured Burkholderiales bacterium Class Betaproteobacteria AY283260.1.1552 Alcaligenes sp. PCNB−2 Class Betaproteobacteria AM934953.1.1374 uncultured Burkholderiales bacterium Class Betaproteobacteria AJ581593.1.1460 uncultured betaAM936569.1.1351 proteobacterium uncultured Class Betaproteobacteria Derxia sp. Class Betaproteobacteria AJ581621.1.1418 uncultured beta proteobacterium Class Betaproteobacteria DQ248272.1.1498 uncultured soil bacterium soil uncultured DQ248272.1.1498 DQ248235.1.1498 uncultured soil bacterium RS49 DQ248270.1.1496 uncultured soil bacterium DQ256489.1.1211 Variovorax paradoxus Class Betaproteobacteria Class paradoxus Variovorax DQ256489.1.1211 AF523053.1.1486 uncultured Comamonadaceae bacterium Class Betaproteobacteria AY706442.1.1396 uncultured bacterium uncultured AY706442.1.1396 AJ536763.1.1422 uncultured bacterium CS000359.1.1530 Variovorax paradoxus Class Betaproteobacteria Class paradoxus Variovorax CS000359.1.1530 AY168733.1.1411 uncultured bacterium AJ009470.1.1526 uncultured bacterium SJA−62 Class Betaproteobacteria Class SJA−62 bacterium uncultured AJ009470.1.1526 AY212561.1.1433 uncultured bacterium D16212.1.1457 Rhodoferax fermentans Class Betaproteobacteria Class fermentans Rhodoferax D16212.1.1457 AY957894.1.1546 uncultured bacterium AJ581620.1.1452 uncultured beta proteobacterium Class Betaproteobacteria RS76 AY625146.1.1498 uncultured bacterium RS65 DQ316832.1.1269 uncultured beta proteobacterium Class Betaproteobacteria DQ404909.1.1513 uncultured bacterium uncultured DQ404909.1.1513 AB021341.1.1466 bacterium rM6 AJ487020.1.1500 uncultured bacterium uncultured AJ487020.1.1500 RS7 RS86RC AF364862.1.1425 bacterium BA128 Class Betaproteobacteria AY957931.1.1529 uncultured bacterium uncultured AY957931.1.1529 CP000884.723807.725332 Delftia acidovorans SPH−1 Class Betaproteobacteria AY957923.1.1520 uncultured bacterium uncultured AY957923.1.1520 RS18 AY957918.1.1527 uncultured bacterium uncultured AY957918.1.1527 AY945883.1.1500 uncultured bacterium AF526940.1.1489 uncultured Ralstonia sp. -

Use of the Diagnostic Bacteriology Laboratory: a Practical Review for the Clinician
148 Postgrad Med J 2001;77:148–156 REVIEWS Postgrad Med J: first published as 10.1136/pmj.77.905.148 on 1 March 2001. Downloaded from Use of the diagnostic bacteriology laboratory: a practical review for the clinician W J Steinbach, A K Shetty Lucile Salter Packard Children’s Hospital at EVective utilisation and understanding of the Stanford, Stanford Box 1: Gram stain technique University School of clinical bacteriology laboratory can greatly aid Medicine, 725 Welch in the diagnosis of infectious diseases. Al- (1) Air dry specimen and fix with Road, Palo Alto, though described more than a century ago, the methanol or heat. California, USA 94304, Gram stain remains the most frequently used (2) Add crystal violet stain. USA rapid diagnostic test, and in conjunction with W J Steinbach various biochemical tests is the cornerstone of (3) Rinse with water to wash unbound A K Shetty the clinical laboratory. First described by Dan- dye, add mordant (for example, iodine: 12 potassium iodide). Correspondence to: ish pathologist Christian Gram in 1884 and Dr Steinbach later slightly modified, the Gram stain easily (4) After waiting 30–60 seconds, rinse with [email protected] divides bacteria into two groups, Gram positive water. Submitted 27 March 2000 and Gram negative, on the basis of their cell (5) Add decolorising solvent (ethanol or Accepted 5 June 2000 wall and cell membrane permeability to acetone) to remove unbound dye. Growth on artificial medium Obligate intracellular (6) Counterstain with safranin. Chlamydia Legionella Gram positive bacteria stain blue Coxiella Ehrlichia Rickettsia (retained crystal violet).