A Splice-Site Mutation in GABRG2 Associated with Childhood Absence Epilepsy and Febrile Convulsions
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genomic Analysis of Bone Marrow Failure and Myelodysplastic Syndromes Reveals Phenotypic and Diagnostic Complexity
Bone Marrow Failure SUPPLEMENTARY APPENDIX Genomic analysis of bone marrow failure and myelodysplastic syndromes reveals phenotypic and diagnostic complexity Michael Y. Zhang,1 Siobán B. Keel,2 Tom Walsh,3 Ming K. Lee,3 Suleyman Gulsuner,3 Amanda C. Watts,3 Colin C. Pritchard,4 Stephen J. Salipante,4 Michael R. Jeng,5 Inga Hofmann,6 David A. Williams,6,7 Mark D. Fleming,8 Janis L. Abkowitz,2 Mary-Claire King,3 and Akiko Shimamura1,9,10 M.Y.Z. and S.B.K. contributed equally to this work. 1Clinical Research Division, Fred Hutchinson Cancer Research Center, Seattle, WA; 2Department of Medicine, Division of Hematology, University of Washington, Seattle, WA; 3Department of Medicine and Department of Genome Sciences, University of Washington, Seattle, WA; 4Department of Laboratory Medicine, University of Washington, Seattle, WA; 5Department of Pediatrics, Stanford Uni- versity School of Medicine, Stanford, CA; 6Division of Hematology/Oncology, Boston Children’s Hospital, Dana Farber Cancer Insti- tute, and Harvard Medical School, Boston, MA; 7Harvard Stem Cell Institute, Boston, MA; 8Department of Pathology, Boston Children’s Hospital, MA; 9Department of Pediatric Hematology/Oncology, Seattle Children’s Hospital, WA; 10Department of Pedi- atrics, University of Washington, Seattle, WA, USA ©2014 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol.2014.113456 Manuscript received on July 22, 2014. Manuscript accepted on September 15, 2014. Correspondence: [email protected] Supplementary Methods Genomics. Libraries were prepared in 96-well format with a Bravo liquid handling robot (Agilent Technologies). One to two micrograms of genomic DNA were sheared to a peak size of 150 bp using a Covaris E series instrument. -
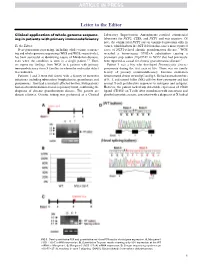
Clinical Application of Whole-Genome Sequencing in Patients with Primary
Letter to the Editor Clinical application of whole-genome sequenc- Laboratory Improvement Amendments–certified commercial ing in patients with primary immunodeficiency laboratory for NCF2, CYBA, and NCF1 and was negative. Of note, the commercial NCF1 screen examined mutations only in To the Editor: exon 2, which harbors the 2GT deletion that causes most reported Next-generation sequencing, including whole-exome sequenc- cases of NCF1-related chronic granulomatous disease.4 WGS ing and whole-genome sequencing (WES and WGS, respectively), revealed a homozygous 579G>A substitution causing a has been successful at identifying causes of Mendelian diseases, premature stop codon (Trp193X) in NCF1 that had previously even when the condition is seen in a single patient.1-3 Here, been reported as causal for chronic granulomatous disease.5 we report our findings from WGS in 6 patients with primary Patient 3 was a boy who developed Pneumocystis jiroveci immunodeficiency from 5 families in whom the molecular defect pneumonia during the first year of life. There was no family was unknown. history of primary immunodeficiency. Immune evaluation Patients 1 and 2 were full sisters with a history of recurrent demonstrated absent serum IgG and IgA. He had normal numbers infections, including tuberculous lymphadenitis, granulomas, and of B, T, and natural killer (NK) cells by flow cytometry and had pneumonias. They had a similarly affected brother. Both patients normal T-cell proliferative responses to mitogens and antigens. had an absent rhodamine-based respiratory burst, confirming the However, the patient lacked any detectable expression of CD40 diagnosis of chronic granulomatous disease. The parents are ligand (CD154) on T cells after stimulation with ionomycin and distant relatives. -

Characterization of Zebrafish GABAA Receptor Subunits
www.nature.com/scientificreports OPEN Characterization of zebrafsh GABAA receptor subunits Kenichiro Sadamitsu, Leona Shigemitsu, Marina Suzuki, Daishi Ito, Makoto Kashima & Hiromi Hirata* γ-Aminobutyric acid (GABA), the major inhibitory neurotransmitter in the central nervous system, exerts its efect through the activation of GABA receptors. GABAA receptors are ligand-gated chloride channels composed of fve subunit proteins. Mammals have 19 diferent GABAA receptor subunits (α1–6, β1–3, γ1–3, δ, ε, π, θ, and ρ1–3), the physiological properties of which have been assayed by electrophysiology. However, the evolutionary conservation of the physiological characteristics of diverged GABAA receptor subunits remains unclear. Zebrafsh have 23 subunits (α1, α2a, α2b, α3–5, α6a, α6b, β1–4, γ1–3, δ, π, ζ, ρ1, ρ2a, ρ2b, ρ3a, and ρ3b), but the electrophysiological properties of these subunits have not been explored. In this study, we cloned the coding sequences for zebrafsh GABAA receptor subunits and investigated their expression patterns in larval zebrafsh by whole- mount in situ hybridization. We also performed electrophysiological recordings of GABA-evoked currents from Xenopus oocytes injected with one or multiple zebrafsh GABAA receptor subunit cRNAs and calculated the half-maximal efective concentrations (EC50s) for each. Our results revealed the spatial expressions and electrophysiological GABA sensitivities of zebrafsh GABAA receptors, suggesting that the properties of GABAA receptor subunits are conserved among vertebrates. γ-Aminobutyric acid (GABA), the major inhibitory neurotransmitter in the central nervous system of vertebrates, 1 controls the excitability of neural networks mainly through GABA A receptors . Te GABAA receptor mediates two types of inhibition, known as phasic and tonic inhibition2. -

In Silico Tools for Splicing Defect Prediction: a Survey from the Viewpoint of End Users
© American College of Medical Genetics and Genomics REVIEW In silico tools for splicing defect prediction: a survey from the viewpoint of end users Xueqiu Jian, MPH1, Eric Boerwinkle, PhD1,2 and Xiaoming Liu, PhD1 RNA splicing is the process during which introns are excised and informaticians in relevant areas who are working on huge data sets exons are spliced. The precise recognition of splicing signals is critical may also benefit from this review. Specifically, we focus on those tools to this process, and mutations affecting splicing comprise a consider- whose primary goal is to predict the impact of mutations within the able proportion of genetic disease etiology. Analysis of RNA samples 5′ and 3′ splicing consensus regions: the algorithms used by different from the patient is the most straightforward and reliable method to tools as well as their major advantages and disadvantages are briefly detect splicing defects. However, currently, the technical limitation introduced; the formats of their input and output are summarized; prohibits its use in routine clinical practice. In silico tools that predict and the interpretation, evaluation, and prospection are also discussed. potential consequences of splicing mutations may be useful in daily Genet Med advance online publication 21 November 2013 diagnostic activities. In this review, we provide medical geneticists with some basic insights into some of the most popular in silico tools Key Words: bioinformatics; end user; in silico prediction tool; for splicing defect prediction, from the viewpoint of end users. Bio- medical genetics; splicing consensus region; splicing mutation INTRODUCTION TO PRE-mRNA SPLICING AND small nuclear ribonucleoproteins and more than 150 proteins, MUTATIONS AFFECTING SPLICING serine/arginine-rich (SR) proteins, heterogeneous nuclear ribo- Sixty years ago, the milestone discovery of the double-helix nucleoproteins, and the regulatory complex (Figure 1). -

Excluded Test List – As of 08/01/2016
Excluded Test List – as of 08/01/2016 CPT As Gene Code 81161 DMD, dup_del 81200 ASPA, cv 81201 APC, fgs 81202 APC, kfv 81203 APC, dup_del 81205 BCKDHA, cv 81205 BCKDHB, cv 81209 BLM, 2281del6ins7 81220 CFTR, cv 81221 CFTR, kfv 81222 CFTR, dup_del 81223 CFTR, fgs 81224 CFTR, intron 8 poly-T 81228 Microarray, CNV 81229 Microarray, SNP 81240 F2, 20210G>A 81241 F5, Leiden (1691G>A) 81242 FANCC, cv 81243 FMR1 81244 FMR1, ma 81250 G6PC, cv 81251 GBA, cv 81252 GJB2, fgs 81253 GJB2, kfv 81254 GJB6, cv 81255 HEXA, cv 81257 HBA1_HBA2, cdel or v 81260 IKBKAP, cv 81271 SRY-FISH 81280 LQT, fgs 12 genes 81281 LQT, kfv 81282 LQT, dup_del 12 genes 81290 MCOLN1, cv 81291 MTHFR, cv 81302 MECP2, fgs 81304 MECP2, dup_del 81324 PMP22, dup_del 81325 PMP22, fgs 1, V14 81326 PMP22, kfv 81330 SMPD1, cv 81331 SNRPN_UBE3A, ma 81350 UGT1A1, cv 81400 ABCC8, F1388del 81400 ACADM, K304E 81400 AGTR1, 1166A>C 81400 BCKDHA, Y438n 81400 CCR5, del 81400 CLRN1, N48K 81400 DPYD, IVS14+1G>A 81400 FGFR1, P252R 81400 FGFR3, P250R 81400 FKTN, retro ins v 81400 GNE, M712T 81400 HPA-6 81400 HPA-9 81400 IVD, A282V 81400 LCT, 13910 C>T 81400 NEB, exon 55 81400 PCDH15, R245X 81400 SHOC2, S2G 81400 SLCO1B1, V174A 81400 SMN1, exon 7 del 81400 SRY, fgs 81400 TOR1A, var 81401 ABCC8, cv 81401 ACADM, cv 81401 ADRB2, cv 81401 AFF2, detect abn 81401 APOB, cv 81401 APOE, cv 81401 AR, alleles 81401 ATN1 81401 ATXN1 81401 ATXN10 81401 ATXN2 81401 ATXN3 81401 ATXN7 81401 ATXN8OS 81401 CACNA1A 2, V14 81401 CBS, cv 81401 CFH_ARMS2, cv 81401 CNBP 81401 CSTB 81401 CYP3A4, cv 81401 CYP3A5, cv 81401 -

Genetic Features of Myelodysplastic Syndrome and Aplastic Anemia in Pediatric and Young Adult Patients
Bone Marrow Failure SUPPLEMENTARY APPENDIX Genetic features of myelodysplastic syndrome and aplastic anemia in pediatric and young adult patients Siobán B. Keel, 1* Angela Scott, 2,3,4 * Marilyn Sanchez-Bonilla, 5 Phoenix A. Ho, 2,3,4 Suleyman Gulsuner, 6 Colin C. Pritchard, 7 Janis L. Abkowitz, 1 Mary-Claire King, 6 Tom Walsh, 6** and Akiko Shimamura 5** 1Department of Medicine, Division of Hematology, University of Washington, Seattle, WA; 2Clinical Research Division, Fred Hutchinson Can - cer Research Center, Seattle, WA; 3Department of Pediatric Hematology/Oncology, Seattle Children’s Hospital, WA; 4Department of Pedi - atrics, University of Washington, Seattle, WA; 5Boston Children’s Hospital, Dana Farber Cancer Institute, and Harvard Medical School, MA; 6Department of Medicine and Department of Genome Sciences, University of Washington, Seattle, WA; and 7Department of Laboratory Medicine, University of Washington, Seattle, WA, USA *SBK and ASc contributed equally to this work **TW and ASh are co-senior authors ©2016 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2016.149476 Received: May 16, 2016. Accepted: July 13, 2016. Pre-published: July 14, 2016. Correspondence: [email protected] or [email protected] Supplementary materials Supplementary methods Retrospective chart review Patient data were collected from medical records by two investigators blinded to the results of genetic testing. The following information was collected: date of birth, transplant, death, and last follow-up, -

Spectrum of Splicing Errors Caused by CHRNE Mutations Affecting Introns and Intron/Exon Boundaries K Ohno, a Tsujino, X-M Shen, M Milone, a G Engel
1of5 ONLINE MUTATION REPORT J Med Genet: first published as 10.1136/jmg.2004.026682 on 1 August 2005. Downloaded from Spectrum of splicing errors caused by CHRNE mutations affecting introns and intron/exon boundaries K Ohno, A Tsujino, X-M Shen, M Milone, A G Engel ............................................................................................................................... J Med Genet 2005;42:e53 (http://www.jmedgenet.com/cgi/content/full/42/8/e53). doi: 10.1136/jmg.2004.026682 Patients Background: Mutations in CHRNE, the gene encoding the Patients 1–5 (respectively a 59 year old woman, a 23 year old muscle nicotinic acetylcholine receptor e subunit, cause man, a 2.5 year old girl, a 6 year old boy, and a 44 year old congenital myasthenic syndromes. Only three of the eight man) have moderate to severe myasthenic symptoms that intronic splice site mutations of CHRNE reported to date have have been present since birth or infancy, decremental EMG had their splicing consequences characterised. responses, and no AChR antibodies. All respond partially to Methods: We analysed four previously reported and five pyridostigmine. Patient 4 underwent an intercostal muscle novel splicing mutations in CHRNE by introducing the entire biopsy for diagnosis, which showed severe endplate AChR normal and mutant genomic CHRNEs into COS cells. deficiency (6% of normal) and compensatory expression of Results and conclusions: We found that short introns (82– the fetal c-AChR at the endplate. 109 nucleotides) favour intron retention, whereas medium to long introns (306–1210 nucleotides) flanking either or both Construction of CHRNE clones for splicing analysis sides of an exon favour exon skipping. Two mutations are of To examine the consequences of the identified splice site particular interest. -

Background Splicing and Genetic Disease
Background splicing and genetic disease Diana Alexieva Imperial College London Yi Long Imperial College London Rupa Sarkar Imperial College London Hansraj Dhayan Imperial College London Emmanuel Bruet Imperial College London Robert Winston Imperial College London Igor Vorechovsky University of Southampton Leandro Castellano University of Sussex Nick Dibb ( [email protected] ) Imperial College London Research Article Keywords: background splicing, splice site mutations, cryptic splice sites, exon skipping, pseudoexons, recursive splicing, spliceosomal mutations, splicing therapy, BRCA1, BRCA1, DMD Posted Date: October 15th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-92665/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/23 Abstract We report that low level background splicing by normal genes can be used to predict the likely effect of splicing mutations upon cryptic splice site activation and exon skipping, with emphasis on the DBASS databases, BRCA1, BRCA2 and DMD. In addition we show that background RNA splice sites are also involved in pseudoexon formation, recursive splicing and aberrant splicing in cancer. We discuss how background splicing information might inform splicing therapy. Introduction We previously established that cryptic splices sites (css) are already active, albeit at very low levels, in normal genes. We did this by using EST data to identify rare splice sites and then compared their positions to known css that are activated in human disease (1). However, this approach was limited to a minority of genes for which there was sucient EST sequence data. Since that time a large amount of RNA-sequencing data has been deposited, which we reasoned would strongly increase the power of css prediction. -

Stem Cells and Ion Channels
Stem Cells International Stem Cells and Ion Channels Guest Editors: Stefan Liebau, Alexander Kleger, Michael Levin, and Shan Ping Yu Stem Cells and Ion Channels Stem Cells International Stem Cells and Ion Channels Guest Editors: Stefan Liebau, Alexander Kleger, Michael Levin, and Shan Ping Yu Copyright © 2013 Hindawi Publishing Corporation. All rights reserved. This is a special issue published in “Stem Cells International.” All articles are open access articles distributed under the Creative Com- mons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Editorial Board Nadire N. Ali, UK Joseph Itskovitz-Eldor, Israel Pranela Rameshwar, USA Anthony Atala, USA Pavla Jendelova, Czech Republic Hannele T. Ruohola-Baker, USA Nissim Benvenisty, Israel Arne Jensen, Germany D. S. Sakaguchi, USA Kenneth Boheler, USA Sue Kimber, UK Paul R. Sanberg, USA Dominique Bonnet, UK Mark D. Kirk, USA Paul T. Sharpe, UK B. Bunnell, USA Gary E. Lyons, USA Ashok Shetty, USA Kevin D. Bunting, USA Athanasios Mantalaris, UK Igor Slukvin, USA Richard K. Burt, USA Pilar Martin-Duque, Spain Ann Steele, USA Gerald A. Colvin, USA EvaMezey,USA Alexander Storch, Germany Stephen Dalton, USA Karim Nayernia, UK Marc Turner, UK Leonard M. Eisenberg, USA K. Sue O’Shea, USA Su-Chun Zhang, USA Marina Emborg, USA J. Parent, USA Weian Zhao, USA Josef Fulka, Czech Republic Bruno Peault, USA Joel C. Glover, Norway Stefan Przyborski, UK Contents Stem Cells and Ion Channels, Stefan Liebau, -

Ion Channels
UC Davis UC Davis Previously Published Works Title THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels. Permalink https://escholarship.org/uc/item/1442g5hg Journal British journal of pharmacology, 176 Suppl 1(S1) ISSN 0007-1188 Authors Alexander, Stephen PH Mathie, Alistair Peters, John A et al. Publication Date 2019-12-01 DOI 10.1111/bph.14749 License https://creativecommons.org/licenses/by/4.0/ 4.0 Peer reviewed eScholarship.org Powered by the California Digital Library University of California S.P.H. Alexander et al. The Concise Guide to PHARMACOLOGY 2019/20: Ion channels. British Journal of Pharmacology (2019) 176, S142–S228 THE CONCISE GUIDE TO PHARMACOLOGY 2019/20: Ion channels Stephen PH Alexander1 , Alistair Mathie2 ,JohnAPeters3 , Emma L Veale2 , Jörg Striessnig4 , Eamonn Kelly5, Jane F Armstrong6 , Elena Faccenda6 ,SimonDHarding6 ,AdamJPawson6 , Joanna L Sharman6 , Christopher Southan6 , Jamie A Davies6 and CGTP Collaborators 1School of Life Sciences, University of Nottingham Medical School, Nottingham, NG7 2UH, UK 2Medway School of Pharmacy, The Universities of Greenwich and Kent at Medway, Anson Building, Central Avenue, Chatham Maritime, Chatham, Kent, ME4 4TB, UK 3Neuroscience Division, Medical Education Institute, Ninewells Hospital and Medical School, University of Dundee, Dundee, DD1 9SY, UK 4Pharmacology and Toxicology, Institute of Pharmacy, University of Innsbruck, A-6020 Innsbruck, Austria 5School of Physiology, Pharmacology and Neuroscience, University of Bristol, Bristol, BS8 1TD, UK 6Centre for Discovery Brain Science, University of Edinburgh, Edinburgh, EH8 9XD, UK Abstract The Concise Guide to PHARMACOLOGY 2019/20 is the fourth in this series of biennial publications. The Concise Guide provides concise overviews of the key properties of nearly 1800 human drug targets with an emphasis on selective pharmacology (where available), plus links to the open access knowledgebase source of drug targets and their ligands (www.guidetopharmacology.org), which provides more detailed views of target and ligand properties. -

Tissue-And Condition-Specific Isoforms of Mammalian Cytochrome
Hindawi Oxidative Medicine and Cellular Longevity Volume 2017, Article ID 1534056, 19 pages https://doi.org/10.1155/2017/1534056 Review Article Tissue- and Condition-Specific Isoforms of Mammalian Cytochrome c Oxidase Subunits: From Function to Human Disease 1 1 1 2 3 Christopher A. Sinkler, Hasini Kalpage, Joseph Shay, Icksoo Lee, Moh H. Malek, 1 1 Lawrence I. Grossman, and Maik Hüttemann 1Center for Molecular Medicine and Genetics, Wayne State University School of Medicine, Detroit, MI 48201, USA 2College of Medicine, Dankook University, Cheonan-si, Chungcheongnam-do 31116, Republic of Korea 3Department of Health Care Sciences, Eugene Applebaum College of Pharmacy and Health Sciences, Wayne State University, Detroit, MI 48201, USA Correspondence should be addressed to Maik Hüttemann; [email protected] Received 2 February 2017; Accepted 29 March 2017; Published 16 May 2017 Academic Editor: Ryuichi Morishita Copyright © 2017 Christopher A. Sinkler et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Cytochrome c oxidase (COX) is the terminal enzyme of the electron transport chain and catalyzes the transfer of electrons from cytochrome c to oxygen. COX consists of 14 subunits, three and eleven encoded, respectively, by the mitochondrial and nuclear DNA. Tissue- and condition-specific isoforms have only been reported for COX but not for the other oxidative phosphorylation complexes, suggesting a fundamental requirement to fine-tune and regulate the essentially irreversible reaction catalyzed by COX. This article briefly discusses the assembly of COX in mammals and then reviews the functions of the six nuclear-encoded COX subunits that are expressed as isoforms in specialized tissues including those of the liver, heart and skeletal muscle, lung, and testes: COX IV-1, COX IV-2, NDUFA4, NDUFA4L2, COX VIaL, COX VIaH, COX VIb-1, COX VIb-2, COX VIIaH, COX VIIaL, COX VIIaR, COX VIIIH/L, and COX VIII-3. -

Mitochondrial Genetics
Mitochondrial genetics Patrick Francis Chinnery and Gavin Hudson* Institute of Genetic Medicine, International Centre for Life, Newcastle University, Central Parkway, Newcastle upon Tyne NE1 3BZ, UK Introduction: In the last 10 years the field of mitochondrial genetics has widened, shifting the focus from rare sporadic, metabolic disease to the effects of mitochondrial DNA (mtDNA) variation in a growing spectrum of human disease. The aim of this review is to guide the reader through some key concepts regarding mitochondria before introducing both classic and emerging mitochondrial disorders. Sources of data: In this article, a review of the current mitochondrial genetics literature was conducted using PubMed (http://www.ncbi.nlm.nih.gov/pubmed/). In addition, this review makes use of a growing number of publically available databases including MITOMAP, a human mitochondrial genome database (www.mitomap.org), the Human DNA polymerase Gamma Mutation Database (http://tools.niehs.nih.gov/polg/) and PhyloTree.org (www.phylotree.org), a repository of global mtDNA variation. Areas of agreement: The disruption in cellular energy, resulting from defects in mtDNA or defects in the nuclear-encoded genes responsible for mitochondrial maintenance, manifests in a growing number of human diseases. Areas of controversy: The exact mechanisms which govern the inheritance of mtDNA are hotly debated. Growing points: Although still in the early stages, the development of in vitro genetic manipulation could see an end to the inheritance of the most severe mtDNA disease. Keywords: mitochondria/genetics/mitochondrial DNA/mitochondrial disease/ mtDNA Accepted: April 16, 2013 Mitochondria *Correspondence address. The mitochondrion is a highly specialized organelle, present in almost all Institute of Genetic Medicine, International eukaryotic cells and principally charged with the production of cellular Centre for Life, Newcastle energy through oxidative phosphorylation (OXPHOS).