Wo 2007/103515 A2
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supporting Online Material
1 2 3 4 5 6 7 Supplementary Information for 8 9 Fractalkine-induced microglial vasoregulation occurs within the retina and is altered early in diabetic 10 retinopathy 11 12 *Samuel A. Mills, *Andrew I. Jobling, *Michael A. Dixon, Bang V. Bui, Kirstan A. Vessey, Joanna A. Phipps, 13 Ursula Greferath, Gene Venables, Vickie H.Y. Wong, Connie H.Y. Wong, Zheng He, Flora Hui, James C. 14 Young, Josh Tonc, Elena Ivanova, Botir T. Sagdullaev, Erica L. Fletcher 15 * Joint first authors 16 17 Corresponding author: 18 Prof. Erica L. Fletcher. Department of Anatomy & Neuroscience. The University of Melbourne, Grattan St, 19 Parkville 3010, Victoria, Australia. 20 Email: [email protected] ; Tel: +61-3-8344-3218; Fax: +61-3-9347-5219 21 22 This PDF file includes: 23 24 Supplementary text 25 Figures S1 to S10 26 Tables S1 to S7 27 Legends for Movies S1 to S2 28 SI References 29 30 Other supplementary materials for this manuscript include the following: 31 32 Movies S1 to S2 33 34 35 36 1 1 Supplementary Information Text 2 Materials and Methods 3 Microglial process movement on retinal vessels 4 Dark agouti rats were anaesthetized, injected intraperitoneally with rhodamine B (Sigma-Aldrich) to label blood 5 vessels and retinal explants established as described in the main text. Retinal microglia were labelled with Iba-1 6 and imaging performed on an inverted confocal microscope (Leica SP5). Baseline images were taken for 10 7 minutes, followed by the addition of PBS (10 minutes) and then either fractalkine or fractalkine + candesartan 8 (10 minutes) using concentrations outlined in the main text. -

1 Metabolic Dysfunction Is Restricted to the Sciatic Nerve in Experimental
Page 1 of 255 Diabetes Metabolic dysfunction is restricted to the sciatic nerve in experimental diabetic neuropathy Oliver J. Freeman1,2, Richard D. Unwin2,3, Andrew W. Dowsey2,3, Paul Begley2,3, Sumia Ali1, Katherine A. Hollywood2,3, Nitin Rustogi2,3, Rasmus S. Petersen1, Warwick B. Dunn2,3†, Garth J.S. Cooper2,3,4,5* & Natalie J. Gardiner1* 1 Faculty of Life Sciences, University of Manchester, UK 2 Centre for Advanced Discovery and Experimental Therapeutics (CADET), Central Manchester University Hospitals NHS Foundation Trust, Manchester Academic Health Sciences Centre, Manchester, UK 3 Centre for Endocrinology and Diabetes, Institute of Human Development, Faculty of Medical and Human Sciences, University of Manchester, UK 4 School of Biological Sciences, University of Auckland, New Zealand 5 Department of Pharmacology, Medical Sciences Division, University of Oxford, UK † Present address: School of Biosciences, University of Birmingham, UK *Joint corresponding authors: Natalie J. Gardiner and Garth J.S. Cooper Email: [email protected]; [email protected] Address: University of Manchester, AV Hill Building, Oxford Road, Manchester, M13 9PT, United Kingdom Telephone: +44 161 275 5768; +44 161 701 0240 Word count: 4,490 Number of tables: 1, Number of figures: 6 Running title: Metabolic dysfunction in diabetic neuropathy 1 Diabetes Publish Ahead of Print, published online October 15, 2015 Diabetes Page 2 of 255 Abstract High glucose levels in the peripheral nervous system (PNS) have been implicated in the pathogenesis of diabetic neuropathy (DN). However our understanding of the molecular mechanisms which cause the marked distal pathology is incomplete. Here we performed a comprehensive, system-wide analysis of the PNS of a rodent model of DN. -
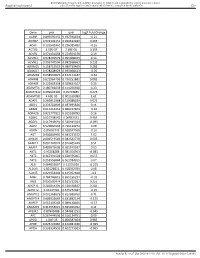
Gene Pval Qval Log2 Fold Change AAMP 0.895690332 0.952598834
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) Gut Gene pval qval Log2 Fold Change AAMP 0.895690332 0.952598834 -0.21 ABI3BP 0.002302151 0.020612283 0.465 ACHE 0.103542461 0.296385483 -0.16 ACTG2 2.99E-07 7.68E-05 3.195 ACVR1 0.071431098 0.224504378 0.19 ACVR1C 0.978209579 0.995008423 0.14 ACVRL1 0.006747504 0.042938663 0.235 ADAM15 0.158715519 0.380719469 0.285 ADAM17 0.978208929 0.995008423 -0.05 ADAM28 0.038932876 0.152174187 -0.62 ADAM8 0.622964796 0.790251882 0.085 ADAM9 0.122003358 0.329623107 0.25 ADAMTS1 0.180766659 0.414256926 0.23 ADAMTS12 0.009902195 0.05703885 0.425 ADAMTS8 4.60E-05 0.001169089 1.61 ADAP1 0.269811968 0.519388039 0.075 ADD1 0.233702809 0.487695826 0.11 ADM2 0.012213453 0.066227879 -0.36 ADRA2B 0.822777921 0.915518785 0.16 AEBP1 0.010738542 0.06035531 0.465 AGGF1 0.117946691 0.320915024 -0.095 AGR2 0.529860903 0.736120272 0.08 AGRN 0.85693743 0.928047568 -0.16 AGT 0.006849995 0.043233572 1.02 AHNAK 0.006519543 0.042542779 0.605 AKAP12 0.001747074 0.016405449 0.51 AKAP2 0.409929603 0.665919397 0.05 AKT1 0.95208288 0.985354963 -0.085 AKT2 0.367391504 0.620376005 0.055 AKT3 0.253556844 0.501934205 0.07 ALB 0.064833867 0.21195036 -0.315 ALDOA 0.83128831 0.918352939 0.08 ALOX5 0.029954404 0.125352668 -0.3 AMH 0.784746815 0.895196237 -0.03 ANG 0.050500474 0.181732067 0.255 ANGPT1 0.281853305 0.538528647 0.285 ANGPT2 0.43147281 0.675272487 -0.15 ANGPTL2 0.001368876 0.013688762 0.71 ANGPTL4 0.686032669 0.831882134 -0.175 ANPEP 0.019103243 0.089148466 -0.57 ANXA2P2 0.412553021 0.665966092 0.11 AP1M2 0.87843088 0.944681253 -0.045 APC 0.267444505 0.516134751 0.09 APOD 1.04E-05 0.000587404 0.985 APOE 0.023722987 0.104981036 -0.395 APOH 0.336334555 0.602273505 -0.065 Sundar R, et al. -

Serine Proteases with Altered Sensitivity to Activity-Modulating
(19) & (11) EP 2 045 321 A2 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 08.04.2009 Bulletin 2009/15 C12N 9/00 (2006.01) C12N 15/00 (2006.01) C12Q 1/37 (2006.01) (21) Application number: 09150549.5 (22) Date of filing: 26.05.2006 (84) Designated Contracting States: • Haupts, Ulrich AT BE BG CH CY CZ DE DK EE ES FI FR GB GR 51519 Odenthal (DE) HU IE IS IT LI LT LU LV MC NL PL PT RO SE SI • Coco, Wayne SK TR 50737 Köln (DE) •Tebbe, Jan (30) Priority: 27.05.2005 EP 05104543 50733 Köln (DE) • Votsmeier, Christian (62) Document number(s) of the earlier application(s) in 50259 Pulheim (DE) accordance with Art. 76 EPC: • Scheidig, Andreas 06763303.2 / 1 883 696 50823 Köln (DE) (71) Applicant: Direvo Biotech AG (74) Representative: von Kreisler Selting Werner 50829 Köln (DE) Patentanwälte P.O. Box 10 22 41 (72) Inventors: 50462 Köln (DE) • Koltermann, André 82057 Icking (DE) Remarks: • Kettling, Ulrich This application was filed on 14-01-2009 as a 81477 München (DE) divisional application to the application mentioned under INID code 62. (54) Serine proteases with altered sensitivity to activity-modulating substances (57) The present invention provides variants of ser- screening of the library in the presence of one or several ine proteases of the S1 class with altered sensitivity to activity-modulating substances, selection of variants with one or more activity-modulating substances. A method altered sensitivity to one or several activity-modulating for the generation of such proteases is disclosed, com- substances and isolation of those polynucleotide se- prising the provision of a protease library encoding poly- quences that encode for the selected variants. -

R&D Assay for Alzheimer's Disease
R&DR&D assayassay forfor Alzheimer’sAlzheimer’s diseasedisease Target screening⳼ Ⲽ㬔 antibody array, ᢜ⭉㬔 ⸽ἐⴐ Amyloid β-peptide Alzheimer’s disease⯸ ኸᷠ᧔ ᆹ⸽ inhibitor, antibody, ELISA kit Surwhrph#Surilohu#Dqwlerg|#Duud| 6OUSFBUFE 1."5SFBUFE )41 $3&# &3, &3, )41 $3&# &3, &3, 壤伡庰䋸TBNQMF ɅH 侴䋸嵄䍴䋸BOBMZUFT䋸䬱娴哜塵 1$ 1$ 1$ 1$ 5IFNPTUSFGFSFODFEBSSBZT 1$ 1$ QQ α 34, .4, 503 Q α 34, .4, 503 %SVHTDSFFOJOH0òUBSHFUFòFDUT0ATHWAY涭廐 6OUSFBUFE 堄币䋸4BNQMF侴䋸8FTUFSOPS&-*4"䍘䧽 1."5SFBUFE P 8FTUFSOCMPU廽喜儤应侴䋸0, Z 4VCTUSBUF -JHIU )31DPOKVHBUFE1BO "OUJQIPTQIPUZSPTJOF .FBO1JYFM%FOTJUZ Y $BQUVSF"OUJCPEZ 5BSHFU"OBMZUF "SSBZ.FNCSBOF $3&# &3, &3, )41 .4, Q α 34, 503 Human XL Cytokine Array kit (ARY022, 102 analytes) Adiponectin,Aggrecan,Angiogenin,Angiopoietin-1,Angiopoietin-2,BAFF,BDNF,Complement,Component C5/C5a,CD14,CD30,CD40L, Chitinase 3-like 1,Complement Factor D,C-Reactive Protein,Cripto-1,Cystatin C,Dkk-1,DPPIV,EGF,EMMPRIN,ENA-78,Endoglin, Fas L,FGF basic,FGF- 7,FGF-19,Flt-3 L,G-CSF,GDF-15,GM-CSF,GRO-α,Grow th Hormone,HGF,ICAM-1,IFN-γ,IGFBP-2,IGFBP-3, IL-1α,IL-1β, IL-1ra,IL-2,IL-3,IL-4,IL- 5,IL-6,IL-8, IL-10,IL-11,IL-12, IL-13,IL-15,IL-16,IL-17A,IL-18 BPa,IL-19,IL-22, IL-23,IL-24,IL-27, IL-31,IL-32α/β/γ,IL-33,IL-34,IP-10,I-TAC,Kallikrein 3,Leptin,LIF,Lipocalin-2,MCP-1,MCP-3,M-CSF,MIF,MIG,MIP-1α/MIP-1β,MIP-3α,MIP-3β,MMP-9, Myeloperoxidase,Osteopontin, p70, PDGF-AA, PDGF-AB/BB,Pentraxin-3, PF4, RAGE, RANTES,RBP4,Relaxin-2, Resistin,SDF-1α,Serpin E1, SHBG, ST2, TARC,TFF3,TfR,TGF- ,Thrombospondin-1,TNF-α, uPAR, VEGF, Vitamin D BP Human Protease (34 analytes) / -

Mitochondrial Protein Quality Control Mechanisms
G C A T T A C G G C A T genes Review Mitochondrial Protein Quality Control Mechanisms Pooja Jadiya * and Dhanendra Tomar * Center for Translational Medicine, Lewis Katz School of Medicine, Temple University, Philadelphia, PA 19140, USA * Correspondence: [email protected] (P.J.); [email protected] (D.T.); Tel.: +1-215-707-9144 (D.T.) Received: 29 April 2020; Accepted: 15 May 2020; Published: 18 May 2020 Abstract: Mitochondria serve as a hub for many cellular processes, including bioenergetics, metabolism, cellular signaling, redox balance, calcium homeostasis, and cell death. The mitochondrial proteome includes over a thousand proteins, encoded by both the mitochondrial and nuclear genomes. The majority (~99%) of proteins are nuclear encoded that are synthesized in the cytosol and subsequently imported into the mitochondria. Within the mitochondria, polypeptides fold and assemble into their native functional form. Mitochondria health and integrity depend on correct protein import, folding, and regulated turnover termed as mitochondrial protein quality control (MPQC). Failure to maintain these processes can cause mitochondrial dysfunction that leads to various pathophysiological outcomes and the commencement of diseases. Here, we summarize the current knowledge about the role of different MPQC regulatory systems such as mitochondrial chaperones, proteases, the ubiquitin-proteasome system, mitochondrial unfolded protein response, mitophagy, and mitochondria-derived vesicles in the maintenance of mitochondrial proteome and health. The proper understanding of mitochondrial protein quality control mechanisms will provide relevant insights to treat multiple human diseases. Keywords: mitochondria; proteome; ubiquitin; proteasome; chaperones; protease; mitophagy; mitochondrial protein quality control; mitochondria-associated degradation; mitochondrial unfolded protein response 1. Introduction Mitochondria are double membrane, dynamic, and semiautonomous organelles which have several critical cellular functions. -

Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association Between BMI and Adult-Onset Non- Atopic
Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association between BMI and Adult-Onset Non- Atopic Asthma Ayoung Jeong 1,2, Medea Imboden 1,2, Akram Ghantous 3, Alexei Novoloaca 3, Anne-Elie Carsin 4,5,6, Manolis Kogevinas 4,5,6, Christian Schindler 1,2, Gianfranco Lovison 7, Zdenko Herceg 3, Cyrille Cuenin 3, Roel Vermeulen 8, Deborah Jarvis 9, André F. S. Amaral 9, Florian Kronenberg 10, Paolo Vineis 11,12 and Nicole Probst-Hensch 1,2,* 1 Swiss Tropical and Public Health Institute, 4051 Basel, Switzerland; [email protected] (A.J.); [email protected] (M.I.); [email protected] (C.S.) 2 Department of Public Health, University of Basel, 4001 Basel, Switzerland 3 International Agency for Research on Cancer, 69372 Lyon, France; [email protected] (A.G.); [email protected] (A.N.); [email protected] (Z.H.); [email protected] (C.C.) 4 ISGlobal, Barcelona Institute for Global Health, 08003 Barcelona, Spain; [email protected] (A.-E.C.); [email protected] (M.K.) 5 Universitat Pompeu Fabra (UPF), 08002 Barcelona, Spain 6 CIBER Epidemiología y Salud Pública (CIBERESP), 08005 Barcelona, Spain 7 Department of Economics, Business and Statistics, University of Palermo, 90128 Palermo, Italy; [email protected] 8 Environmental Epidemiology Division, Utrecht University, Institute for Risk Assessment Sciences, 3584CM Utrecht, Netherlands; [email protected] 9 Population Health and Occupational Disease, National Heart and Lung Institute, Imperial College, SW3 6LR London, UK; [email protected] (D.J.); [email protected] (A.F.S.A.) 10 Division of Genetic Epidemiology, Medical University of Innsbruck, 6020 Innsbruck, Austria; [email protected] 11 MRC-PHE Centre for Environment and Health, School of Public Health, Imperial College London, W2 1PG London, UK; [email protected] 12 Italian Institute for Genomic Medicine (IIGM), 10126 Turin, Italy * Correspondence: [email protected]; Tel.: +41-61-284-8378 Int. -

Structure of Neurolysin Reveals a Deep Channel That Limits Substrate Access
Structure of neurolysin reveals a deep channel that limits substrate access C. Kent Brown*†, Kevin Madauss*, Wei Lian‡, Moriah R. Beck§, W. David Tolbert¶, and David W. Rodgersʈ Department of Molecular and Cellular Biochemistry and Center for Structural Biology, University of Kentucky, Lexington, KY 40536 Communicated by Stephen C. Harrison, Harvard University, Cambridge, MA, December 29, 2000 (received for review November 14, 2000) The zinc metallopeptidase neurolysin is shown by x-ray crystallog- cytosolic, but it also can be secreted or associated with the raphy to have large structural elements erected over the active site plasma membrane (11), and some of the enzyme is made with a region that allow substrate access only through a deep narrow mitochondrial targeting sequence by initiation at an alternative channel. This architecture accounts for specialization of this neu- transcription start site (12). ropeptidase to small bioactive peptide substrates without bulky Although neurolysin cleaves a number of neuropeptides in secondary and tertiary structures. In addition, modeling studies vitro, its most established (5, 13, 14) role in vivo (along with indicate that the length of a substrate N-terminal to the site of thimet oligopeptidase) is in metabolism of neurotensin, a 13- hydrolysis is restricted to approximately 10 residues by the limited residue neuropeptide. It hydrolyzes this peptide between resi- size of the active site cavity. Some structural elements of neuro- dues 10 and 11, creating shorter fragments that are believed to  lysin, including a five-stranded -sheet and the two active site be inactive. helices, are conserved with other metallopeptidases. The connect- Neurotensin (pGlu-Leu-Tyr-Gln-Asn-Lys-Pro-Arg-Arg- ing loop regions of these elements, however, are much extended Pro s Tyr-Ile-Leu) is found in a variety of peripheral and in neurolysin, and they, together with other open coil elements, central tissues where it is involved in a number of effects, line the active site cavity. -

ADAM8 Expression in Invasive Breast Cancer Promotes Tumor Dissemination and Metastasis
ADAM8 expression in invasive breast cancer promotes tumor dissemination and metastasis Mathilde Romagnoli, Nora Mineva, Michael Polmear, Catharina Conrad, Srimathi Srinivasan, Delphine Loussouarn, Sophie Barillé-Nion, Irene Georgakoudi, Aine Dagg, Enda Mcdermott, et al. To cite this version: Mathilde Romagnoli, Nora Mineva, Michael Polmear, Catharina Conrad, Srimathi Srinivasan, et al.. ADAM8 expression in invasive breast cancer promotes tumor dissemination and metastasis. EMBO Molecular Medicine, Wiley Open Access, 2014, 6 (2), pp.278-294. 10.1002/emmm.201303373. inserm-02447040 HAL Id: inserm-02447040 https://www.hal.inserm.fr/inserm-02447040 Submitted on 21 Jan 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Research Article ADAM8 expression in invasive breast cancer promotes tumor dissemination and metastasis Mathilde Romagnoli1, Nora D Mineva1, Michael Polmear2, Catharina Conrad3, Srimathi Srinivasan1, Delphine Loussouarn4, Sophie Barille-Nion4, Irene Georgakoudi2, Aine Dagg5,6, Enda W McDermott6, Michael J Duffy6, Patricia M. McGowan5,6, Uwe Schlomann3, Maddy Parsons7,Jorg€ W Bartsch3 & Gail E Sonenshein1,* Abstract Introduction The transmembrane metalloprotease-disintegrin ADAM8 mediates Cancer metastasis results from a multistep process that selects for cell adhesion and shedding of ligands, receptors and extracellular invasive tumor cells capable of escaping from the primary site and matrix components. -
VPS10P-Domain Receptor Family
R&D Systems Tools for Cell Biology Research™ NEUROSCIENCE FOCUS: NEUROTROPHIC FACTORS VPS10P-domain Receptor Family FEATURED DATA: APP · BDNF · b-NGF · NGF R · Phospho-APP · Phospho-TrkA · Phospho-TrkB · Phospho-TrkC · SorCS2 · SorLA · Sortilin · TrkA VPS10P-domain Receptors Vacuolar protein sorting 10 protein (VPS10P)-domain receptors are type I transmembrane proteins that bind a range of ligands including neu- rotrophins, neuropeptides, and other transmembrane proteins. Additional studies suggest novel roles for VPS10P-domain receptors in ciliary neurotrophic factor (CNTF) signaling, and in the trafficking of Lipoprotein Lipase (LPL) and Cholesterol. In vertebrates, there are five members of the family, Sortilin, sorting protein-related receptor with A-type repeats (SorLA), Sortilin-related receptor CNS expressed 1 (SorCS1), SorCS2, and SorCS3. These multifunctional molecules have been shown to affect neuronal viability and function by regulating protein transport and signal transduction. Each receptor is expressed in distinct neuronal populations, suggesting discrete functions in different cell types. Variable function between family members is supported by the subcellular expression of each receptor. Sortilin and SorLA are predominantly found intracellularly, in the trans-Golgi network (TGN), with less than 10% at the cell surface. Subcellular trafficking of SorCS1 is dependent on the splice variant. SorCS1a is predominantly intracellular, SorCS1b is expressed at the cell surface, and SorCS1c is evenly divided between the two. SorCS2 -

Airway Inflammation in Mice Adam8 Limits the Development of Allergic
Adam8 Limits the Development of Allergic Airway Inflammation in Mice Martin D. Knolle, Takahiro Nakajima, Anja Hergrueter, Kushagra Gupta, Francesca Polverino, Vanessa J. Craig, This information is current as Susanne E. Fyfe, Muhammad Zahid, Perdita Permaul, of October 2, 2021. Manuela Cernadas, Gilbert Montano, Yohannes Tesfaigzi, Lynette Sholl, Lester Kobzik, Elliot Israel and Caroline A. Owen J Immunol 2013; 190:6434-6449; Prepublished online 13 May 2013; Downloaded from doi: 10.4049/jimmunol.1202329 http://www.jimmunol.org/content/190/12/6434 http://www.jimmunol.org/ Supplementary http://www.jimmunol.org/content/suppl/2013/05/13/jimmunol.120232 Material 9.DC1 References This article cites 77 articles, 18 of which you can access for free at: http://www.jimmunol.org/content/190/12/6434.full#ref-list-1 Why The JI? Submit online. by guest on October 2, 2021 • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Adam8 Limits the Development of Allergic Airway Inflammation in Mice Martin D. -

ADAM10 Controls Collagen Signaling and Cell Migration on Collagen by Shedding the Ectodomain of Discoidin Domain Receptor 1 (DDR1)
M BoC | ARTICLE ADAM10 controls collagen signaling and cell migration on collagen by shedding the ectodomain of discoidin domain receptor 1 (DDR1) Yasuyuki Shitomia, Ida B. Thøgersenb, Noriko Itoa, Birgit Leitingerc, Jan J. Enghildb, and Yoshifumi Itoha aKennedy Institute of Rheumatology, Nuffield Department of Orthopaedics, Rheumatology and Musculoskeletal Sciences, University of Oxford, Oxford OX3 7FY, United Kingdom; bDepartment of Molecular Biology and Genetics, University of Aarhus, DK-8000 Aarhus C, Denmark; cNational Heart and Lung Institute, Imperial College London, London SW7 2AZ, United Kingdom ABSTRACT Discoidin domain receptor 1 (DDR1) is a receptor tyrosine kinase that binds and Monitoring Editor transmits signals from various collagens in epithelial cells. However, how DDR1–dependent Jean E. Schwarzbauer signaling is regulated has not been understood. Here we report that collagen binding in- Princeton University duces ADAM10-dependent ectodomain shedding of DDR1. DDR1 shedding is not a result of Received: Oct 21, 2014 an activation of its signaling pathway, since DDR1 mutants defective in signaling were shed Revised: Dec 4, 2014 in an efficient manner. DDR1 and ADAM10 were found to be in a complex on the cell surface, Accepted: Dec 16, 2014 but shedding did not occur unless collagen bound to DDR1. Using a shedding-resistant DDR1 mutant, we found that ADAM10-dependent DDR1 shedding regulates the half-life of colla- gen-induced phosphorylation of the receptor. Our data also revealed that ADAM10 plays an important role in regulating DDR1-mediated cell adhesion to achieve efficient cell migration on collagen matrices. INTRODUCTION Extracellular matrix (ECM) is essential in multicellular organisms to discoidin domain receptors (DDRs), glycoprotein VI, leukocyte-asso- maintain functional tissue structures; it acts as scaffolding to support ciated, immunoglobulin-like receptors, and mannose receptors such cell migration and as a reservoir for growth factors.