Presentation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

Hidden Targets in RAF Signalling Pathways to Block Oncogenic RAS Signalling
G C A T T A C G G C A T genes Review Hidden Targets in RAF Signalling Pathways to Block Oncogenic RAS Signalling Aoife A. Nolan 1, Nourhan K. Aboud 1, Walter Kolch 1,2,* and David Matallanas 1,* 1 Systems Biology Ireland, School of Medicine, University College Dublin, Belfield, Dublin 4, Ireland; [email protected] (A.A.N.); [email protected] (N.K.A.) 2 Conway Institute of Biomolecular & Biomedical Research, University College Dublin, Belfield, Dublin 4, Ireland * Correspondence: [email protected] (W.K.); [email protected] (D.M.) Abstract: Oncogenic RAS (Rat sarcoma) mutations drive more than half of human cancers, and RAS inhibition is the holy grail of oncology. Thirty years of relentless efforts and harsh disappointments have taught us about the intricacies of oncogenic RAS signalling that allow us to now get a pharma- cological grip on this elusive protein. The inhibition of effector pathways, such as the RAF-MEK-ERK pathway, has largely proven disappointing. Thus far, most of these efforts were aimed at blocking the activation of ERK. Here, we discuss RAF-dependent pathways that are regulated through RAF functions independent of catalytic activity and their potential role as targets to block oncogenic RAS signalling. We focus on the now well documented roles of RAF kinase-independent functions in apoptosis, cell cycle progression and cell migration. Keywords: RAF kinase-independent; RAS; MST2; ASK; PLK; RHO-α; apoptosis; cell cycle; cancer therapy Citation: Nolan, A.A.; Aboud, N.K.; Kolch, W.; Matallanas, D. Hidden Targets in RAF Signalling Pathways to Block Oncogenic RAS Signalling. -

CDK4/6 Inhibitors in Melanoma: a Comprehensive Review
cells Review CDK4/6 Inhibitors in Melanoma: A Comprehensive Review Mattia Garutti 1,*, Giada Targato 2 , Silvia Buriolla 2 , Lorenza Palmero 1,2 , Alessandro Marco Minisini 2 and Fabio Puglisi 1,2 1 CRO Aviano National Cancer Institute IRCCS, 33081 Aviano, Italy; [email protected] (L.P.); [email protected] (F.P.) 2 Department of Medicine (DAME), University of Udine, 33100 Udine, Italy; [email protected] (G.T.); [email protected] (S.B.); [email protected] (A.M.M.) * Correspondence: [email protected] Abstract: Historically, metastatic melanoma was considered a highly lethal disease. However, recent advances in drug development have allowed a significative improvement in prognosis. In particular, BRAF/MEK inhibitors and anti-PD1 antibodies have completely revolutionized the management of this disease. Nonetheless, not all patients derive a benefit or a durable benefit from these therapies. To overtake this challenges, new clinically active compounds are being tested in the context of clinical trials. CDK4/6 inhibitors are drugs already available in clinical practice and preliminary evidence showed a promising activity also in melanoma. Herein we review the available literature to depict a comprehensive landscape about CDK4/6 inhibitors in melanoma. We present the molecular and genetic background that might justify the usage of these drugs, the preclinical evidence, the clinical available data, and the most promising ongoing clinical trials. Keywords: CDK4/6; CDK4; CDK6; melanoma; Palbociclib; Ribociclib; Abemaciclib Citation: Garutti, M.; Targato, G.; Buriolla, S.; Palmero, L.; Minisini, A.M.; Puglisi, F. CDK4/6 Inhibitors in Melanoma: A Comprehensive 1. Introduction Review. Cells 2021, 10, 1334. -

In Vitro Pharmacological Profiling of R406 Identifies Molecular Targets
ORIGINAL ARTICLE In vitro pharmacological profiling of R406 identifies molecular targets underlying the clinical effects of fostamatinib Michael G. Rolf, Jon O. Curwen, Margaret Veldman-Jones, Cath Eberlein, Jianyan Wang, Alex Harmer, Caroline J. Hellawell & Martin Braddock AstraZeneca R&D Alderley Park, Macclesfield, Cheshire SK10 4TG, United Kingdom Keywords Abstract Blood pressure elevation, fostamatinib, in vitro pharmacological profiling, R406, SYK Off-target pharmacology may contribute to both adverse and beneficial effects of a new drug. In vitro pharmacological profiling is often applied early in drug Correspondence discovery; there are fewer reports addressing the relevance of broad profiles to Michael G. Rolf, AstraZeneca R&D Molndal,€ clinical adverse effects. Here, we have characterized the pharmacological profile Pepparedsleden 1, 431 83 Mo¨ lndal, Sweden. of the active metabolite of fostamatinib, R406, linking an understanding of drug Tel: +46 31 776 60 40; Fax: +46 31 776 37 selectivity to the increase in blood pressure observed in clinical studies. R406 60; E-mail: [email protected] was profiled in a broad range of in vitro assays to generate a comprehensive Funding Information pharmacological profile and key targets were further investigated using func- No funding information provided. tional and cellular assay systems. A combination of traditional literature searches and text-mining approaches established potential mechanistic links Received: 9 April 2015; Accepted: 14 July between the profile of R406 and clinical side effects. R406 was selective outside 2015 the kinase domain, with only antagonist activity at the adenosine A3 receptor in the range relevant to clinical effects. R406 was less selective in the kinase Pharma Res Per, 3(5), 2015, e00175, doi: 10.1002/prp2.175 domain, having activity at many protein kinases at therapeutically relevant con- centrations when tested in multiple in vitro systems. -

Endothelial Cell Injury in Atherosclerosis Is Regulated by Glycolysis (Review)
INTERNATIONAL JOURNAL OF MOleCular meDICine 47: 65-76, 2021 Mechanism overview and target mining of atherosclerosis: Endothelial cell injury in atherosclerosis is regulated by glycolysis (Review) RUIYING WANG1-5, MIN WANG1-5, JINGXUE YE1-5, GUIBO SUN1-5 and XIAOBO SUN1-5 1Institute of Medicinal Plant Development, Peking Union Medical College and Chinese Academy of Medical Sciences; 2Beijing Key Laboratory of Innovative Drug Discovery of Traditional Chinese Medicine (Natural Medicine) and Translational Medicine, Institute of Medicinal Plant Development, Peking Union Medical College and Chinese Academy of Medical Sciences; 3Key Laboratory of Bioactive Substances and Resources Utilization of Chinese Herbal Medicine, Ministry of Education, Institute of Medicinal Plant Development, Peking Union Medical College and Chinese Academy of Medical Sciences; 4Key Laboratory of Efficacy Evaluation of Chinese Medicine Against Glycolipid Metabolic Disorders, State Administration of Traditional Chinese Medicine, Institute of Medicinal Plant Development, Peking Union Medical College and Chinese Academy of Medical Sciences; 5Key Laboratory of New Drug Discovery Based on Classic Chinese Medicine Prescription, Chinese Academy of Medical Sciences, Beijing 100193, P.R. China Received August 14, 2020; Accepted November 5, 2020 DOI: 10.3892/ijmm.2020.4798 Abstract. Atherosclerosis (AS) is a chronic disease with a In conclusion, endothelial cell injury in AS may be alleviated by complex pathology that may lead to several cardiovascular and glycolysis and is a potential clinical treatment strategy for AS. cerebrovascular diseases; however, further research is neces- sary to fully elucidate its pathogenesis. The main risk factors for AS include lipid metabolism disorders, endothelial cell Contents injury, inflammation and immune dysfunction, among which vascular endothelial cell damage is considered as the main 1. -

Phospho-RPS6KA1(T359) Antibody Peptide Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # Ap3497a
9765 Clairemont Mesa Blvd, Suite C San Diego, CA 92124 Tel: 858.875.1900 Fax: 858.622.0609 Phospho-RPS6KA1(T359) Antibody Peptide Affinity Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP3497a Specification Phospho-RPS6KA1(T359) Antibody - Product Information Application WB, DB,E Primary Accession Q15418 Other Accession P10666, P10665, P18652 Reactivity Human Predicted Chicken, Xenopus Host Rabbit Clonality Polyclonal Isotype Rabbit Ig Calculated MW 82723 Phospho-RPS6KA1(T359) Antibody - Additional Information Gene ID 6195 Other Names Western blot analysis of extracts from Hela Ribosomal protein S6 kinase alpha-1, cells,untreated or treated with S6K-alpha-1, 90 kDa ribosomal protein S6 TPA,200nM,using phospho RPS6KA1-T359 (left) kinase 1, p90-RSK 1, p90RSK1, p90S6K, MAP or RPS6KA1 antibody(right) kinase-activated protein kinase 1a, MAPK-activated protein kinase 1a, MAPKAP kinase 1a, MAPKAPK-1a, Ribosomal S6 kinase 1, RSK-1, RPS6KA1, MAPKAPK1A, RSK1 Target/Specificity This RPS6KA1 Antibody is generated from rabbits immunized with a KLH conjugated synthetic phosphopeptide corresponding to amino acid residues surrounding T359 of human RPS6KA1. Dilution WB~~1:2000 DB~~1:500 Format Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity purification. Dot blot analysis of anti-RPS6KA1-pT359 Pab (RB13385) on nitrocellulose membrane. 50ng Storage of Phospho-peptide or Non Phospho-peptide per Maintain refrigerated at 2-8°C for up to 6 dot were adsorbed. Antibody working months. For long term storage store at -20°C concentrations are 0.5ug per ml. in small aliquots to prevent freeze-thaw cycles. -

CDK4/6 Inhibition Reprograms Mitochondrial Metabolism in BRAFV600 Melanoma Via a P53 Dependent Pathway
cancers Article CDK4/6 Inhibition Reprograms Mitochondrial Metabolism in BRAFV600 Melanoma via a p53 Dependent Pathway Nancy T. Santiappillai 1,2, Shatha Abuhammad 1 , Alison Slater 1, Laura Kirby 1, Grant A. McArthur 1,3,†, Karen E. Sheppard 1,2,3,*,† and Lorey K. Smith 1,3,*,† 1 Cancer Research Division, Peter MacCallum Cancer Centre, Melbourne 3052, Australia; [email protected] (N.T.S.); [email protected] (S.A.); [email protected] (A.S.); [email protected] (L.K.); [email protected] (G.A.M.) 2 Department of Biochemistry and Molecular Biology, University of Melbourne, Melbourne 3052, Australia 3 Sir Peter MacCallum Department of Oncology, University of Melbourne, Melbourne 3052, Australia * Correspondence: [email protected] (K.E.S.); [email protected] (L.K.S.) † These authors contributed equally to the work. Simple Summary: Cyclin-dependent kinases 4 and 6 (CDK4/6) are key enzymes controlling the cell cycle. CDK4/6 inhibitors are being tested in multiple clinical trials for a range of cancers including melanoma, and a deeper understanding of how they interact with other therapies is vital for their clinical development. Beyond the cell cycle, CDK4/6 regulates cell metabolism, which is a critical factor determining response to standard-of-care mitogen-activated protein kinase (MAPK) pathway therapies in melanoma. Here, we show that CDK4/6 inhibitors increase glutamine and fatty acid-oxidation-dependent mitochondrial metabolism in melanoma cells, but they do not alter the metabolic response to MAPK inhibitors. These observations shed light on how CDK4/6 inhibitors Citation: Santiappillai, N.T.; impinge on the regulation of metabolism and how they interact with other therapies in the setting of Abuhammad, S.; Slater, A.; Kirby, L.; melanoma. -

Deep Multiomics Profiling of Brain Tumors Identifies Signaling Networks
ARTICLE https://doi.org/10.1038/s41467-019-11661-4 OPEN Deep multiomics profiling of brain tumors identifies signaling networks downstream of cancer driver genes Hong Wang 1,2,3, Alexander K. Diaz3,4, Timothy I. Shaw2,5, Yuxin Li1,2,4, Mingming Niu1,4, Ji-Hoon Cho2, Barbara S. Paugh4, Yang Zhang6, Jeffrey Sifford1,4, Bing Bai1,4,10, Zhiping Wu1,4, Haiyan Tan2, Suiping Zhou2, Laura D. Hover4, Heather S. Tillman 7, Abbas Shirinifard8, Suresh Thiagarajan9, Andras Sablauer 8, Vishwajeeth Pagala2, Anthony A. High2, Xusheng Wang 2, Chunliang Li 6, Suzanne J. Baker4 & Junmin Peng 1,2,4 1234567890():,; High throughput omics approaches provide an unprecedented opportunity for dissecting molecular mechanisms in cancer biology. Here we present deep profiling of whole proteome, phosphoproteome and transcriptome in two high-grade glioma (HGG) mouse models driven by mutated RTK oncogenes, PDGFRA and NTRK1, analyzing 13,860 proteins and 30,431 phosphosites by mass spectrometry. Systems biology approaches identify numerous master regulators, including 41 kinases and 23 transcription factors. Pathway activity computation and mouse survival indicate the NTRK1 mutation induces a higher activation of AKT down- stream targets including MYC and JUN, drives a positive feedback loop to up-regulate multiple other RTKs, and confers higher oncogenic potency than the PDGFRA mutation. A mini-gRNA library CRISPR-Cas9 validation screening shows 56% of tested master regulators are important for the viability of NTRK-driven HGG cells, including TFs (Myc and Jun) and metabolic kinases (AMPKa1 and AMPKa2), confirming the validity of the multiomics inte- grative approaches, and providing novel tumor vulnerabilities. -
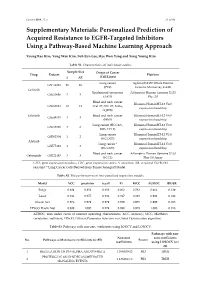
Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach
Cancers 2019, 11, x S1 of S9 Supplementary Materials: Personalized Prediction of Acquired Resistance to EGFR-Targeted Inhibitors Using a Pathway-Based Machine Learning Approach Young Rae Kim, Yong Wan Kim, Suh Eun Lee, Hye Won Yang and Sung Young Kim Table S1. Characteristics of individual studies. Sample Size Origin of Cancer Drug Dataset Platform S AR (Cell Lines) Lung cancer Agilent-014850 Whole Human GSE34228 26 26 (PC9) Genome Microarray 4x44K Gefitinib Epidermoid carcinoma Affymetrix Human Genome U133 GSE10696 3 3 (A431) Plus 2.0 Head and neck cancer Illumina HumanHT-12 V4.0 GSE62061 12 12 (Cal-27, SSC-25, FaDu, expression beadchip SQ20B) Erlotinib Head and neck cancer Illumina HumanHT-12 V4.0 GSE49135 3 3 (HN5) expression beadchip Lung cancer (HCC827, Illumina HumanHT-12 V3.0 GSE38310 3 6 ER3, T15-2) expression beadchip Lung cancer Illumina HumanHT-12 V3.0 GSE62504 1 2 (HCC827) expression beadchip Afatinib Lung cancer * Illumina HumanHT-12 V4.0 GSE75468 1 3 (HCC827) expression beadchip Head and neck cancer Affymetrix Human Genome U133 Cetuximab GSE21483 3 3 (SCC1) Plus 2.0 Array GEO, gene expression omnibus; GSE, gene expression series; S, sensitive; AR, acquired EGFR-TKI resistant; * Lung Cancer Cells Derived from Tumor Xenograft Model. Table S2. The performances of four penalized regression models. Model ACC precision recall F1 MCC AUROC BRIER Ridge 0.889 0.852 0.958 0.902 0.782 0.964 0.129 Lasso 0.944 0.957 0.938 0.947 0.889 0.991 0.042 Elastic Net 0.978 0.979 0.979 0.979 0.955 0.999 0.023 EPSGO Elastic Net 0.989 1.000 0.979 0.989 0.978 1.000 0.018 AUROC, area under curve of receiver operating characteristic; ACC, accuracy; MCC, Matthews correlation coefficient; EPSGO, Efficient Parameter Selection via Global Optimization algorithm. -

The P90 RSK Family Members: Common Functions and Isoform Specificity
Published OnlineFirst August 22, 2013; DOI: 10.1158/0008-5472.CAN-12-4448 Cancer Review Research The p90 RSK Family Members: Common Functions and Isoform Specificity Romain Lara, Michael J. Seckl, and Olivier E. Pardo Abstract The p90 ribosomal S6 kinases (RSK) are implicated in various cellular processes, including cell proliferation, survival, migration, and invasion. In cancer, RSKs modulate cell transformation, tumorigenesis, and metastasis. Indeed, changes in the expression of RSK isoforms have been reported in several malignancies, including breast, prostate, and lung cancers. Four RSK isoforms have been identified in humans on the basis of their high degree of sequence homology. Although this similarity suggests some functional redundancy between these proteins, an increasing body of evidence supports the existence of isoform-based specificity among RSKs in mediating particular cellular processes. This review briefly presents the similarities between RSK family members before focusing on the specific function of each of the isoforms and their involvement in cancer progression. Cancer Res; 73(17); 1–8. Ó2013 AACR. Introduction subsequently cloned throughout the Metazoan kingdom (2). The extracellular signal–regulated kinase (ERK)1/2 pathway The genomic analysis of several cancer types suggests that fi is involved in key cellular processes, including cell prolifera- these genes are not frequently ampli ed or mutated, with some tion, differentiation, survival, metabolism, and migration. notable exceptions (e.g., in the case of hepatocellular carcino- More than 30% of all human cancers harbor mutations within ma; ref. 6). Table 1 summarizes reported genetic changes in this pathway, mostly resulting in gain of function and conse- RSK genes. -

In This Issue
IN THIS ISSUE AACR Project GENIE Facilitates Sharing of Genomic and Clinical Data • Project GENIE launched with data • Project GENIE aims to promote • Data from Project GENIE may guide from 19,000 patients from 8 institu data sharing to enhance precision identification of drug targets and tions with a variety of tumor types . medicine research . biomarkers in patients with cancer . Genomic profi ling of tumors accessible in the cBioPortal for Cancer Genomics. Despite the has become increasingly com- differences in genomic testing at the contributing centers, the mon across cancer types, but genomic data collected so far are largely concordant across the data are not regularly made the centers, and mutation rates are similar to those reported by available to the entire research The Cancer Genome Atlas. Initial results from Project GENIE community. The American Asso- suggest that more than 30% of tumors harbor potentially ciation for Cancer Research clinically actionable mutations. Advantages of this platform (AACR) launched the Genomics, include integration of clinical data from electronic health Evidence, Neoplasia, Informa- records and increased statistical power to potentially facilitate tion, Exchange (GENIE) project understanding of the clinical relevance of somatic mutations in partnership with eight academic institutions to facilitate and improve patient selection for targeted therapies. The large-scale sharing of genomic and clinical data. The AACR establishment of infrastructure for integrating genomic and Project GENIE Consortium released the fi rst of set of data clinical data has the potential to aid identifi cation of thera- from 19,000 patients in January 2017, and the number of sam- peutic targets and biomarkers of treatment and response in ples included is expected to grow to more than 100,000 within patients with cancer to enhance precision medicine research 5 years as more centers join the Consortium. -

Protein Kinase C As a Therapeutic Target in Non-Small Cell Lung Cancer
International Journal of Molecular Sciences Review Protein Kinase C as a Therapeutic Target in Non-Small Cell Lung Cancer Mohammad Mojtaba Sadeghi 1,2, Mohamed F. Salama 2,3,4 and Yusuf A. Hannun 1,2,3,* 1 Department of Biochemistry, Molecular and Cellular Biology, Stony Brook University, Stony Brook, NY 11794, USA; [email protected] 2 Stony Brook Cancer Center, Stony Brook University Hospital, Stony Brook, NY 11794, USA; [email protected] 3 Department of Medicine, Stony Brook University, Stony Brook, NY 11794, USA 4 Department of Biochemistry, Faculty of Veterinary Medicine, Mansoura University, Mansoura 35516, Dakahlia Governorate, Egypt * Correspondence: [email protected] Abstract: Driver-directed therapeutics have revolutionized cancer treatment, presenting similar or better efficacy compared to traditional chemotherapy and substantially improving quality of life. Despite significant advances, targeted therapy is greatly limited by resistance acquisition, which emerges in nearly all patients receiving treatment. As a result, identifying the molecular modulators of resistance is of great interest. Recent work has implicated protein kinase C (PKC) isozymes as mediators of drug resistance in non-small cell lung cancer (NSCLC). Importantly, previous findings on PKC have implicated this family of enzymes in both tumor-promotive and tumor-suppressive biology in various tissues. Here, we review the biological role of PKC isozymes in NSCLC through extensive analysis of cell-line-based studies to better understand the rationale for PKC inhibition. Citation: Sadeghi, M.M.; Salama, PKC isoforms α, ", η, ι, ζ upregulation has been reported in lung cancer, and overexpression correlates M.F.; Hannun, Y.A. Protein Kinase C with worse prognosis in NSCLC patients.