Molecular Interactions and Implications of Aldose Reductase Inhibition by PGA1 and Clinically Used Prostaglandins
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’S Biological Network Based on Systematic Pharmacology
ORIGINAL RESEARCH published: 25 June 2021 doi: 10.3389/fphar.2021.601846 Exploring the Regulatory Mechanism of Hedysarum Multijugum Maxim.-Chuanxiong Rhizoma Compound on HIF-VEGF Pathway and Cerebral Ischemia-Reperfusion Injury’s Biological Network Based on Systematic Pharmacology Kailin Yang 1†, Liuting Zeng 1†, Anqi Ge 2†, Yi Chen 1†, Shanshan Wang 1†, Xiaofei Zhu 1,3† and Jinwen Ge 1,4* Edited by: 1 Takashi Sato, Key Laboratory of Hunan Province for Integrated Traditional Chinese and Western Medicine on Prevention and Treatment of 2 Tokyo University of Pharmacy and Life Cardio-Cerebral Diseases, Hunan University of Chinese Medicine, Changsha, China, Galactophore Department, The First 3 Sciences, Japan Hospital of Hunan University of Chinese Medicine, Changsha, China, School of Graduate, Central South University, Changsha, China, 4Shaoyang University, Shaoyang, China Reviewed by: Hui Zhao, Capital Medical University, China Background: Clinical research found that Hedysarum Multijugum Maxim.-Chuanxiong Maria Luisa Del Moral, fi University of Jaén, Spain Rhizoma Compound (HCC) has de nite curative effect on cerebral ischemic diseases, *Correspondence: such as ischemic stroke and cerebral ischemia-reperfusion injury (CIR). However, its Jinwen Ge mechanism for treating cerebral ischemia is still not fully explained. [email protected] †These authors share first authorship Methods: The traditional Chinese medicine related database were utilized to obtain the components of HCC. The Pharmmapper were used to predict HCC’s potential targets. Specialty section: The CIR genes were obtained from Genecards and OMIM and the protein-protein This article was submitted to interaction (PPI) data of HCC’s targets and IS genes were obtained from String Ethnopharmacology, a section of the journal database. -

Identification and Characterization of Genes That Control Fat
Claire D’Andre et al. Journal of Animal Science and Biotechnology 2013, 4:43 http://www.jasbsci.com/content/4/1/43 JOURNAL OF ANIMAL SCIENCE AND BIOTECHNOLOGY RESEARCH Open Access Identification and characterization of genes that control fat deposition in chickens Hirwa Claire D’Andre1,3*, Wallace Paul2, Xu Shen3, Xinzheng Jia3, Rong Zhang3, Liang Sun3 and Xiquan Zhang3 Abstract Background: Fat deposits in chickens contribute significantly to meat quality attributes such as juiciness, flavor, taste and other organoleptic properties. The quantity of fat deposited increases faster and earlier in the fast- growing chickens than in slow-growing chickens. In this study, Affymetrix Genechip® Chicken Genome Arrays 32773 transcripts were used to compare gene expression profiles in liver and hypothalamus tissues of fast-growing and slow-growing chicken at 8 wk of age. Real-time RT-PCR was used to validate the differential expression of genes selected from the microarray analysis. The mRNA expression of the genes was further examined in fat tissues. The association of single nucleotide polymorphisms of four lipid-related genes with fat traits was examined in a F2 resource population. Results: Four hundred genes in the liver tissues and 220 genes hypothalamus tissues, respectively, were identified to be differentially expressed in fast-growing chickens and slow-growing chickens. Expression levels of genes for lipid metabolism (SULT1B1, ACSBG2, PNPLA3, LPL, AOAH) carbohydrate metabolism (MGAT4B, XYLB, GBE1, PGM1, HKDC1)cholesttrol biosynthesis (FDPS, LSS, HMGCR, NSDHL, DHCR24, IDI1, ME1) HSD17B7 and other reaction or pro- cesses (CYP1A4, CYP1A1, AKR1B1, CYP4V2, DDO) were higher in the fast-growing White Recessive Rock chickens than in the slow-growing Xinghua chickens. -

MMP-25 Metalloprotease Regulates Innate Immune Response Through NF- Κb Signaling Clara Soria-Valles, Ana Gutiérrez-Fernández, Fernando G
MMP-25 Metalloprotease Regulates Innate Immune Response through NF- κB Signaling Clara Soria-Valles, Ana Gutiérrez-Fernández, Fernando G. Osorio, Dido Carrero, Adolfo A. Ferrando, Enrique Colado, This information is current as M. Soledad Fernández-García, Elena Bonzon-Kulichenko, of September 27, 2021. Jesús Vázquez, Antonio Fueyo and Carlos López-Otín J Immunol 2016; 197:296-302; Prepublished online 3 June 2016; doi: 10.4049/jimmunol.1600094 Downloaded from http://www.jimmunol.org/content/197/1/296 Supplementary http://www.jimmunol.org/content/suppl/2016/06/01/jimmunol.160009 Material 4.DCSupplemental http://www.jimmunol.org/ References This article cites 41 articles, 15 of which you can access for free at: http://www.jimmunol.org/content/197/1/296.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 27, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology MMP-25 Metalloprotease Regulates Innate Immune Response through NF-kB Signaling Clara Soria-Valles,* Ana Gutie´rrez-Ferna´ndez,* Fernando G. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Clostridium Difficile Exploits a Host Metabolite Produced During Toxin-Mediated Infection
bioRxiv preprint doi: https://doi.org/10.1101/2021.01.14.426744; this version posted January 15, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Clostridium difficile exploits a host metabolite produced during toxin-mediated infection 2 3 Kali M. Pruss and Justin L. Sonnenburg* 4 5 Department of Microbiology & Immunology, Stanford University School of Medicine, Stanford 6 CA, USA 94305 7 *Corresponding author address: [email protected] 8 9 10 11 12 13 14 15 16 17 18 19 20 21 22 bioRxiv preprint doi: https://doi.org/10.1101/2021.01.14.426744; this version posted January 15, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 23 Several enteric pathogens can gain specific metabolic advantages over other members of 24 the microbiota by inducing host pathology and inflammation. The pathogen Clostridium 25 difficile (Cd) is responsible for a toxin-mediated colitis that causes 15,000 deaths in the U.S. 26 yearly1, yet the molecular mechanisms by which Cd benefits from toxin-induced colitis 27 remain understudied. Up to 21% of healthy adults are asymptomatic carriers of toxigenic 28 Cd2, indicating that Cd can persist as part of a healthy microbiota; antibiotic-induced 29 perturbation of the gut ecosystem is associated with transition to toxin-mediated disease. -

Development of Potent and Selective Inhibitors of Aldo−Keto Reductase
Article pubs.acs.org/jmc Development of Potent and Selective Inhibitors of Aldo−Keto Reductase 1C3 (Type 5 17β-Hydroxysteroid Dehydrogenase) Based on N-Phenyl-Aminobenzoates and Their Structure−Activity Relationships † § ‡ § † † † ‡ Adegoke O. Adeniji, , Barry M. Twenter, , Michael C. Byrns, Yi Jin, Mo Chen, Jeffrey D. Winkler,*, † and Trevor M. Penning*, † Department of Pharmacology and Center of Excellence in Environmental Toxicology, Perelman School of Medicine, University of Pennsylvania, 130C John Morgan Building, 3620 Hamilton Walk, Philadelphia, Pennsylvania 19104-6084, United States ‡ Department of Chemistry, University of Pennsylvania, 231 South 34th Street, Philadelphia, Pennsylvania 19104, United States *S Supporting Information ABSTRACT: Aldo−keto reductase 1C3 (AKR1C3; type 5 17β-hydroxy- steroid dehydrogenase) is overexpressed in castration resistant prostate cancer (CRPC) and is implicated in the intratumoral biosynthesis of testosterone and 5α-dihydrotestosterone. Selective AKR1C3 inhibitors are required because compounds should not inhibit the highly related AKR1C1 and AKR1C2 isoforms which are involved in the inactivation of 5α-dihydrotestosterone. NSAIDs, N-phenylanthranilates in particular, are potent but nonselective AKR1C3 inhibitors. Using flufenamic acid, 2-{[3-(trifluoromethyl)phenyl]amino}- benzoic acid, as lead compound, five classes of structural analogues were synthesized and evaluated for AKR1C3 inhibitory potency and selectivity. Structure−activity relationship (SAR) studies revealed that a meta-carboxylic acid group relative to the amine conferred pronounced AKR1C3 selectivity without loss of potency, while electron withdrawing groups on the phenylamino B-ring were optimal for AKR1C3 inhibition. Lead compounds did not inhibit COX-1 or COX-2 but blocked the AKR1C3 mediated production of testosterone in LNCaP-AKR1C3 cells. These compounds offer promising leads toward new therapeutics for CRPC. -

Ular Basis of KAISER, S
Heredity 74 (1995) 267—273 Received6May 1994 OThe Genetical Society of Great Britain 1981. The molecular basis of KAISER, S. 1935. The factorsSOMMER, controlling H. 1991. Genetic shape control ofand flower sizedevelopment in Genetic differentiation among populations of by homeotic genes in Antirrhinum majus. Science, 250, Myopus schisticolor (the wood lemming) — LANDE, R. 1981. The minimumSHORE, number J. S. AND BARRETF, of genes S. C. H. 1990. contributing Quantitative genetics of floral characters in homostylous Turnera ulmifolia var, isozyme variation angustifolia Willd. (Turneraceae). Heredity, 64, LANDE, R, 1983. The responseSINNO1F, to e. w.selection 1935. Evidence onfor the major existence of and genes VADIM B. FEDOROVif, RICKARD FREDRIKSSON1: & KARLFREDGA* SINNO'rr, E. w. 1936. A developmental analysis of inherited t/nstitute of Plant and Animal Ecology, Russian Academy of Sciences, 8 March Street 202, Jekaterinburg 620 008, tion on correlated characters.shape Evolution, differences in Cucurbit37, 1210—1226. fruits. Am. Nat., 70, Russia and Department of Genetics, Uppsa/a University, Box 7003, S-75007Uppsala, Sweden LORD, C. M. AND HILL, .i. i'. 1987. Evidence for heterochrony in Toinvestigate genetic differentiation among populations of the wood lemming Myopus schistico!or (eds) Development as an Evolutiona,ySMITH, H. H. 1950. DevelopmentalProcess, restrictions pp. 47—70. on recombina- (Muridae, Rodentia) isozyme variation in 10 populations from three regions, Fennoscandia, Western and Eastern Siberia, was examined. From 20 loci examined, the two most polymorphic MACNAIR, M. R. AND CUMBES,TAKHTAJAN, o. .i. 1989. A. 1976. The Neoteny genetic and the architectureorigin of flowering ones, Idh-1 and Pgi-1, were used in the complete survey. -

Transcriptome Analysis of Different Multidrug-Resistant Gastric Carcinoma Cells
in vivo 19: 583-590 (2005) Transcriptome Analysis of Different Multidrug-resistant Gastric Carcinoma Cells STEFFEN HEIM1 and HERMANN LAGE2 1Europroteome AG, Berlin-Hennigsdorf; 2Institute of Pathology, Charité Campus Mitte, Schumannstr. 20/21, D-10117 Berlin, Germany Abstract. Multidrug resistance (MDR) of human cancers is advanced cancer. These therapeutic protocols produce an the major cause of failure of chemotherapy. To better overall response rate of about 20% at best using a single- understand the molecular events associated with the agent, and up to 50% using combination chemotherapy. development of different types of MDR, two different multidrug- Thus, a large number of malignancies are incurable. resistant gastric carcinoma cell lines, the MDR1/P-glycoprotein- Generally, gastrointestinal cancers, including gastric expressing cell line EPG85-257RDB and the MDR1/P- carcinoma, are naturally resistant to many anticancer drugs. glycoprotein-negative cell variant EPG85-257RNOV, as well as Additionally, these tumors are able to develop acquired the corresponding drug-sensitive parental cell line EPG85-257P, drug resistance phenotypes, which include the multidrug were used for analyses of the mRNA expression profiles by resistance (MDR) phenomenon. cDNA array hybridization. Of more than 12,000 genes spotted The MDR phenotype is characterized by simultaneous on the arrays, 156 genes were detected as being significantly resistance of tumor cells to various antineoplastic agents regulated in the cell line EPG85-257RDB in comparison to the which are structurally and functionally unrelated. Besides non-resistant cell variant, and 61 genes were found to be the classical MDR phenotype, mediated by the enhanced differentially expressed in the cell line EPG85-257RNOV. -
Data Supplement
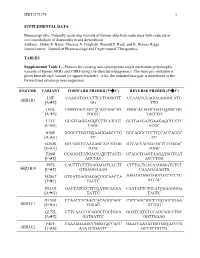
JPET#173179 1 SUPPLEMENTAL DATA Manuscript title: Naturally occurring variants of human aldo-keto reductases with reduced in vitro metabolism of daunorubicin and doxorubicin Authors: Onkar S. Bains, Thomas A. Grigliatti, Ronald E. Reid, and K. Wayne Riggs Journal name: Journal of Pharmacology and Experimental Therapeutics TABLES Supplemental Table 1—Primers for creating non-synonymous single nucleotide polymorphic variants of human AKRs and CBR4 using site-directed mutagenesis. The base pair mutation is given beneath each variant [in square brackets]. Also, the mutated base pair is underlined in the forward and reverse primer sequences. ENZYME VARIANT FORWARD PRIMER (5'3') REVERSE PRIMER (5'3') I15F CAAGATGCCCTTCCTGGGGTT CCAACCCCAGGAAGGGCATC AKR1B1 [AT] GG TTG H42L CGGGTACCGCCTCATCGACTG GGGCACAGTCGATGAGGCGG [AT] TGCCC TACCCG L73V GCGTGAGGAGGTCTTCATCGT GCTGACGATGAAGACCTCCTC [CG] CAGC ACGC K90E GGGCCTGGTGGAAGGAGCCTG GGCAGGCTCCTTCCACCAGGC [AG] CC CC G204S GCCAGTCCAAAAGCATCGTGG GTCACCACGATGCTTTTGGAC [GA] TGAC TGGC T288I CCAGGATATGACCATCTTACTC GTAGCTGAGTAAGATGGTCAT [CT] AGCTAC ATCCTGG P87S CACTTTCTTTGAGAGATCCCTT CTTTCCTCACAAGGGATCTCT AKR1B10 [CT] GTGAGGAAAG CAAAGAAAGTG M286T GTGATGAGGAGACGGCAACCA GAGTATGGTTGCCGTCTCCTC [TC] TACTC ATCAC N313D GACTATCCCTTCGATGCAGAA CAATATTCTGCATCGAAGGGA [AG] TATTG TAGTC R170H CCAACTTCAACCACAGGCAGC CTCCAGCTGCCTGTGGTTGAA AKR1C1 [GA] TGGAG GTTGG Q172L CTTCAACCGCAGGCTGCTGGA GGATCATCTCCAGCAGCCTGC [AT] GATGATCC GGTTGAAG F46Y CAATAGAAGCCGGGTACCACC GAATCAATATGGTGGTACCCG AKR1C2 [TA] ATATTTGATTC GCTTCTATTG JPET#173179 -

Global Profiling of Metabolic Adaptation to Hypoxic Stress in Human Glioblastoma Cells
Global profiling of metabolic adaptation to hypoxic stress in human glioblastoma cells. Kucharzewska, Paulina; Christianson, Helena; Belting, Mattias Published in: PLoS ONE DOI: 10.1371/journal.pone.0116740 2015 Link to publication Citation for published version (APA): Kucharzewska, P., Christianson, H., & Belting, M. (2015). Global profiling of metabolic adaptation to hypoxic stress in human glioblastoma cells. PLoS ONE, 10(1), [e0116740]. https://doi.org/10.1371/journal.pone.0116740 Total number of authors: 3 General rights Unless other specific re-use rights are stated the following general rights apply: Copyright and moral rights for the publications made accessible in the public portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognise and abide by the legal requirements associated with these rights. • Users may download and print one copy of any publication from the public portal for the purpose of private study or research. • You may not further distribute the material or use it for any profit-making activity or commercial gain • You may freely distribute the URL identifying the publication in the public portal Read more about Creative commons licenses: https://creativecommons.org/licenses/ Take down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. LUND UNIVERSITY PO Box 117 221 00 Lund +46 46-222 00 00 Download date: 07. Oct. 2021 RESEARCH ARTICLE Global Profiling of Metabolic Adaptation to Hypoxic Stress in Human Glioblastoma Cells Paulina Kucharzewska1, Helena C. -

Orlistat, a Novel Potent Antitumor Agent for Ovarian Cancer: Proteomic Analysis of Ovarian Cancer Cells Treated with Orlistat
INTERNATIONAL JOURNAL OF ONCOLOGY 41: 523-532, 2012 Orlistat, a novel potent antitumor agent for ovarian cancer: proteomic analysis of ovarian cancer cells treated with Orlistat HUI-QIONG HUANG1*, JING TANG1*, SHENG-TAO ZHOU1, TAO YI1, HONG-LING PENG1, GUO-BO SHEN2, NA XIE2, KAI HUANG2, TAO YANG2, JIN-HUA WU2, CAN-HUA HUANG2, YU-QUAN WEI2 and XIA ZHAO1,2 1Gynecological Oncology of Biotherapy Laboratory, Department of Gynecology and Obstetrics, West China Second Hospital, Sichuan University, Chengdu, Sichuan; 2State Key Laboratory of Biotherapy and Cancer Center, West China Hospital, Sichuan University, Chengdu, Sichuan, P.R. China Received February 9, 2012; Accepted March 19, 2012 DOI: 10.3892/ijo.2012.1465 Abstract. Orlistat is an orally administered anti-obesity drug larly PKM2. These changes confirmed our hypothesis that that has shown significant antitumor activity in a variety of Orlistat is a potential inhibitor of ovarian cancer and can be tumor cells. To identify the proteins involved in its antitumor used as a novel adjuvant antitumor agent. activity, we employed a proteomic approach to reveal protein expression changes in the human ovarian cancer cell line Introduction SKOV3, following Orlistat treatment. Protein expression profiles were analyzed by 2-dimensional polyacrylamide In the 1920s, the Nobel Prize winner Otto Warburg observed gel electrophoresis (2-DE) and protein identification was a marked increase in glycolysis and enhanced lactate produc- performed on a MALDI-Q-TOF MS/MS instrument. More tion in tumor cells even when maintained in conditions of high than 110 differentially expressed proteins were visualized oxygen tension (termed Warburg effect), leading to widespread by 2-DE and Coomassie brilliant blue staining. -

Integrative Analysis of Promising Molecular Biomarkers and Pathways for Coronary Artery Disease Using WGCNA and Metade Methods
MOLECULAR MEDICINE REPORTS 18: 2789-2797, 2018 Integrative analysis of promising molecular biomarkers and pathways for coronary artery disease using WGCNA and MetaDE methods SHILIN YAN Department of Cardiology, Yangling Demonstration Zone Hospital, Xianyang, Shaanxi 712100, P.R. China Received January 8, 2018; Accepted May 31, 2018 DOI: 10.3892/mmr.2018.9277 Abstract. The present study aimed to examine the molecular ‘Fc gamma R‑mediated phagocytosis’ pathways may serve mechanisms of coronary artery disease (CAD). A total of four important roles in CAD. microarray datasets (training dataset no. GSE12288; valida- tion dataset nos. GSE20680, GSE20681 and GSE42148) were Introduction downloaded from the Gene Expression Omnibus database, which included CAD and healthy samples. Weighted gene Coronary artery disease (CAD) remains one of the most co-expression network analysis was applied to identify highly common causes of morbidity and mortality globally, with preserved modules across the four datasets. Differentially an increased prevalence predicted in the near future (1). expressed genes (DEGs) with significant consistency in the The occurrence of CAD events is primarily due to crosstalk four datasets were selected using the MetaDE method. The between genetic and environmental factors (2). A number of overlapping genes amongst the DEGs with significant consis- risk factors of CAD have been identified, including smoking, tency and in the preserved modules were used to construct obesity and family history (3). Available treatment options a protein-protein interaction (PPI) network, followed by primarily include lifestyle alterations, medical treatment functional enrichment analysis. A total of 11 modules were and surgical interventions, which are chosen depending on established in the training dataset, and five of them were highly comorbidities and the preferences of the individual patient (4).