Expression of Cancer-Testis Antigens MAGEA1, MAGEA3, ACRBP, PRAME, SSX2, and CTAG2 in Myxoid and Round Cell Liposarcoma
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Cancer Immunity (1 December 2006) Vol
Cancer Immun 1424-9634Academy of Cancer Immunology Cancer Immunity (1 December 2006) Vol. 6, p. 12 Submitted: 26 September 2006. Accepted: 10 October 2006. Copyright © 2006 by Andrew J. G. Simpson 061012 Article Physical interaction of two cancer-testis antigens, MAGE-C1 (CT7) and NY-ESO-1 (CT6) Hearn J. Cho1*,**, Otavia L. Caballero2*, Sacha Gnjatic2, Valéria C. C. Andrade3, Gisele W. Colleoni3, Andre L. Vettore4, Hasina H. Outtz1, Sheila Fortunato2, Nasser Altorki1, Cathy A. Ferrera1, Ramon Chua2, Achim A. Jungbluth2, Yao-Tseng Chen1, Lloyd J. Old2 and Andrew J. G. Simpson2 1Weill Medical College of Cornell University, 1300 York Avenue, New York, NY 10021, USA 2Ludwig Institute for Cancer Research, New York Branch at Memorial Sloan-Kettering Cancer Center, 1275 York Avenue, New York, NY 10021, USA 3Escola Paulista de Medicina, Universidade Federal de Sao Paulo, Sao Paulo, SP, Brazil 4Ludwig Institute for Cancer Research, Sao Paulo Branch, Sao Paulo, SP, Brazil *These authors contributed equally to this work **Present address: NYU Cancer Institute, New York University School of Medicine, 550 First Avenue, New York, NY 10016, USA Contributed by: LJ Old Cancer/testis (CT) antigens are the protein products of germ line- encoded on the X chromosome (CT-X antigens) and those that associated genes that are activated in a wide variety of tumors and can are not (non-X CT antigens) (1). elicit autologous cellular and humoral immune responses. CT It is estimated that 10% of the genes on the X-chromosome antigens can be divided between those that are encoded on the X belong to CT-X families (5). -
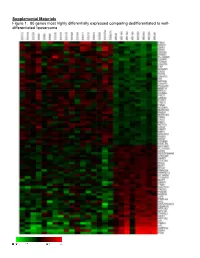
Supplemental Materials Figure 1. 80 Genes Most Highly Differentially Expressed Comparing Dedifferentiated to Well- Differentiated Liposarcoma
Supplemental Materials Figure 1. 80 genes most highly differentially expressed comparing dedifferentiated to well- differentiated liposarcoma Figure 2. Validation of microa CDC2, RACGAP1, FGFR-2, MAD2 and CITED1 200 CDK4 by Liposarcoma Subtype 16 0 12 0 80 Expression40 Level 0 Nl Fat rray data by quantitative RT-P Well Diff 16 0 p16 by Liposarcoma Subtype Dediff 12 0 80 Myxoid Expression40 Level Round Cell 0 12 0 MDM2 by Liposarcoma Subtype 10 0 Pleomorphic Nl Fat 80 60 40 Expression Level 20 RACGAP1 by LiposarcomaWell Diff Subtype 0 70 CR for genes CDK4, MDM2, p16, 60 Dediff 50 Nl Fat 40 30 Myxoid Expression20 Level Well Diff 10 50 0 Round Cell CDC2 by Liposarcoma Subtype Dediff 40 Nl Fat Pleomorphic 30 Myxoid 20 Expression Level Well Diff 10 MAD2L1 by Liposarcoma Subtype Round Cell 60 0 50 Dediff Pleomorphic 40 Nl Fat 30 Myxoid 20 Expression Level Well Diff 10 FGFR-2 by Liposarcoma Subtype 0 Round Cell 200 16 0 Dediff Nl Fat Pleomorphic 12 0 Myxoid 80 Expression Level Well Diff 40 Round Cell 0 Dediff Pleomorphic Nl Fat Myxoid Well Diff CITED1 by Liposarcoma Subtype Round Cell 10 0 80 Dediff Pleomorphic 60 40 Myxoid Expression Level 20 0 Round Cell Nl Fat Pleomorphic Well Diff Dediff Myxoid Round Cell Pleomorphic Table 1. 142 Gene classifier for liposarcoma subtype The genes used for each pair wise subtype comparison are grouped together. The flag column indicates which genes are unique to each subtype comparison. The values show the mean expression levels (actually the mean of the log expression levels was computed and than transformed back to absolute expression level). -

Alterations of Genetic Variants and Transcriptomic Features of Response to Tamoxifen in the Breast Cancer Cell Line
Alterations of Genetic Variants and Transcriptomic Features of Response to Tamoxifen in the Breast Cancer Cell Line Mahnaz Nezamivand-Chegini Shiraz University Hamed Kharrati-Koopaee Shiraz University https://orcid.org/0000-0003-2345-6919 seyed taghi Heydari ( [email protected] ) Shiraz University of Medical Sciences https://orcid.org/0000-0001-7711-1137 Hasan Giahi Shiraz University Ali Dehshahri Shiraz University of Medical Sciences Mehdi Dianatpour Shiraz University of Medical Sciences Kamran Bagheri Lankarani Shiraz University of Medical Sciences Research Keywords: Tamoxifen, breast cancer, genetic variants, RNA-seq. Posted Date: August 17th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-783422/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/33 Abstract Background Breast cancer is one of the most important causes of mortality in the world, and Tamoxifen therapy is known as a medication strategy for estrogen receptor-positive breast cancer. In current study, two hypotheses of Tamoxifen consumption in breast cancer cell line (MCF7) were investigated. First, the effect of Tamoxifen on genes expression prole at transcriptome level was evaluated between the control and treated samples. Second, due to the fact that Tamoxifen is known as a mutagenic factor, there may be an association between the alterations of genetic variants and Tamoxifen treatment, which can impact on the drug response. Methods In current study, the whole-transcriptome (RNA-seq) dataset of four investigations (19 samples) were derived from European Bioinformatics Institute (EBI). At transcriptome level, the effect of Tamoxifen was investigated on gene expression prole between control and treatment samples. -
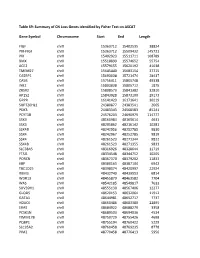
Table S9: Summary of CN Loss Genes Identified by Fisher Test on ASCAT
Table S9: Summary of CN Loss Genes identified by Fisher Test on ASCAT Gene Symbol Chromosome Start End Length FIGF chrX 15363712 15402535 38824 PIR-FIGF chrX 15363712 15509432 145721 PIR chrX 15402923 15511711 108789 BMX chrX 15518899 15574652 55754 ACE2 chrX 15579155 15620192 41038 TMEM27 chrX 15645440 15683154 37715 CA5BP1 chrX 15693038 15721474 28437 CA5B chrX 15756411 15805748 49338 INE2 chrX 15803838 15805712 1875 ZRSR2 chrX 15808573 15841382 32810 AP1S2 chrX 15843928 15873100 29173 GRPR chrX 16141423 16171641 30219 SUPT20HL1 chrX 24380877 24383541 2665 PDK3 chrX 24483343 24568583 85241 PCYT1B chrX 24576203 24690979 114777 SSX9 chrX 48160984 48165614 4631 SSX3 chrX 48205862 48216142 10281 SSX4B chrX 48242956 48252785 9830 SSX4 chrX 48242967 48252785 9819 SSX4 chrX 48261523 48271344 9822 SSX4B chrX 48261523 48271355 9833 SLC38A5 chrX 48316926 48328644 11719 FTSJ1 chrX 48334548 48344752 10205 PORCN chrX 48367370 48379202 11833 EBP chrX 48380163 48387104 6942 TBC1D25 chrX 48398074 48420997 22924 RBM3 chrX 48432740 48439553 6814 WDR13 chrX 48455879 48463582 7704 WAS chrX 48542185 48549817 7633 SUV39H1 chrX 48555130 48567406 12277 GLOD5 chrX 48620153 48632064 11912 GATA1 chrX 48644981 48652717 7737 HDAC6 chrX 48660486 48683380 22895 ERAS chrX 48684922 48688279 3358 PCSK1N chrX 48689503 48694036 4534 TIMM17B chrX 48750729 48755426 4698 PQBP1 chrX 48755194 48760422 5229 SLC35A2 chrX 48760458 48769235 8778 PIM2 chrX 48770458 48776413 5956 OTUD5 chrX 48779302 48815648 36347 KCND1 chrX 48818638 48828251 9614 GRIPAP1 chrX 48830133 48858675 28543 -

NY-ESO-1 (CTAG1B) Expression in Mesenchymal Tumors
Modern Pathology (2015) 28, 587–595 & 2015 USCAP, Inc. All rights reserved 0893-3952/15 $32.00 587 NY-ESO-1 (CTAG1B) expression in mesenchymal tumors Makoto Endo1,2,7, Marieke A de Graaff3,7, Davis R Ingram4, Simin Lim1, Dina C Lev4, Inge H Briaire-de Bruijn3, Neeta Somaiah5, Judith VMG Bove´e3, Alexander J Lazar6 and Torsten O Nielsen1 1Department of Pathology and Laboratory Medicine, University of British Columbia, Vancouver, British Columbia, Canada; 2Department of Orthopaedic Surgery, Kyushu University, Fukuoka, Japan; 3Department of Pathology, Leiden University Medical Center, Leiden, The Netherlands; 4Department of Surgical Oncology, The University of Texas MD Anderson Cancer Center, Houston, TX, USA; 5Department of Sarcoma Medical Oncology, The University of Texas MD Anderson Cancer Center, Houston, TX, USA and 6Department of Pathology, The University of Texas MD Anderson Cancer Center, Houston, TX, USA New York esophageal squamous cell carcinoma 1 (NY-ESO-1, CTAG1B) is a cancer-testis antigen and currently a focus of several targeted immunotherapeutic strategies. We performed a large-scale immunohistochemical expression study of NY-ESO-1 using tissue microarrays of mesenchymal tumors from three institutions in an international collaboration. A total of 1132 intermediate and malignant and 175 benign mesenchymal lesions were enrolled in this study. Immunohistochemical staining was performed on tissue microarrays using a monoclonal antibody for NY-ESO-1. Among mesenchymal tumors, myxoid liposarcomas showed the highest positivity for NY-ESO-1 (88%), followed by synovial sarcomas (49%), myxofibrosarcomas (35%), and conventional chondrosarcomas (28%). Positivity of NY-ESO-1 in the remaining mesenchymal tumors was consistently low, and no immunoreactivity was observed in benign mesenchymal lesions. -

Gene Expression Profiling Using Nanostring Digital RNA Counting to Identify Potential Target Antigens for Melanoma Immunotherapy
Published OnlineFirst September 10, 2013; DOI: 10.1158/1078-0432.CCR-13-1253 Clinical Cancer Human Cancer Biology Research Gene Expression Profiling using Nanostring Digital RNA Counting to Identify Potential Target Antigens for Melanoma Immunotherapy Rachel E. Beard, Daniel Abate-Daga, Shannon F. Rosati, Zhili Zheng, John R. Wunderlich, Steven A. Rosenberg, and Richard A. Morgan Abstract Purpose: The success of immunotherapy for the treatment of metastatic cancer is contingent on the identification of appropriate target antigens. Potential targets must be expressed on tumors but show restricted expression on normal tissues. To maximize patient eligibility, ideal target antigens should be expressed on a high percentage of tumors within a histology and, potentially, in multiple different malignancies. Design: A Nanostring probeset was designed containing 97 genes, 72 of which are considered potential candidate genes for immunotherapy. Five established melanoma cell lines, 59 resected metastatic mela- noma tumors, and 31 normal tissue samples were profiled and analyzed using Nanostring technology. Results: Of the 72 potential target genes, 33 were overexpressed in more than 20% of studied melanoma tumor samples. Twenty of those genes were identified as differentially expressed between normal tissues and tumor samples by ANOVA analysis. Analysis of normal tissue gene expression identified seven genes with limited normal tissue expression that warrant further consideration as potential immunotherapy target antigens: CSAG2, MAGEA3, MAGEC2, IL13RA2, PRAME, CSPG4, and SOX10. These genes were highly overexpressed on a large percentage of the studied tumor samples, with expression in a limited number of normal tissue samples at much lower levels. Conclusion: The application of Nanostring RNA counting technology was used to directly quantitate the gene expression levels of multiple potential tumor antigens. -

Seromic Profiling of Ovarian and Pancreatic Cancer
Seromic profiling of ovarian and pancreatic cancer Sacha Gnjatica,1, Erika Rittera, Markus W. Büchlerb, Nathalia A. Gieseb, Benedikt Brorsc, Claudia Freid, Anne Murraya, Niels Halamad, Inka Zörnigd, Yao-Tseng Chene, Christopher Andrewsf, Gerd Rittera, Lloyd J. Olda,1, Kunle Odunsig,2, and Dirk Jägerd,2 aLudwig Institute for Cancer Research Ltd, Memorial-Sloan Kettering Cancer Center, New York, NY 10065; bDepartment of General Surgery, cDepartment of Theoretical Bioinformatics, and dMedizinische Onkologie, Nationales Centrum für Tumorerkrankungen, University Hospital Heidelberg, Heidelberg D-69120, Germany; eDepartment of Pathology, Weill Medical College of Cornell University, New York, NY 10065; and fDepartment of Biostatistics and gDepartment of Gynecologic Oncology, Roswell Park Cancer Institute, Buffalo, NY 14263 Contributed by Lloyd J. Old, December 10, 2009 (sent for review August 20, 2009) Autoantibodies, a hallmark of both autoimmunity and cancer, analyzing a series of lung cancer and healthy control sera on a represent an easily accessible surrogate for measuring adaptive small array (329 proteins) for antigen reactivity using this anti- immune responses to cancer. Sera can now be assayed for re- body profiling method, referred to here as “seromics,” we were activity against thousands of proteins using microarrays, but there able to detect known antigens with sensitivity and specificity is no agreed-upon standard to analyze results. We developed a set comparable to ELISA, as well as new antigens that are now of tailored quality control and normalization procedures based on under further investigation. Contrary to gene microarrays where ELISA validation to allow patient comparisons and determination changes in the pattern of gene expression are detected in clus- of individual cutoffs for specificity and sensitivity. -

(12) United States Patent (10) Patent No.: US 7,662,561 B2 Godfrey Et Al
USOO7662561 B2 (12) United States Patent (10) Patent No.: US 7,662,561 B2 Godfrey et al. (45) Date of Patent: Feb. 16, 2010 (54) IDENTIFICATION OF MARKERS IN 6,440,725 B1 8/2002 Pourahmadi et al. ESOPHAGEAL CANCER, COLON CANCER, 7,101,663 B2 9/2006 Godfrey et al. HEAD AND NECK CANCER, AND 2001/005.1344 A1* 12/2001 Shalon et al. .................. 435/6 MELANOMA 2006/0068418 A1* 3/2006 Godfrey et al. ................ 435/6 (75) Inventors: Tony E. Godfrey, Bronxville, NY (US); FOREIGN PATENT DOCUMENTS Liqiang Xi, Plainsboro, NJ (US); Siva EP 105O587 11, 2000 Raja, Jamaica Plain, MA (US); Steven WO WO95/11687 5, 1995 J. Hughes, Blawnox, PA (US); William WO WO98,0897O 3, 1998 E. Gooding, Pittsburgh, PA (US) WO WO99, 13104 3, 1999 WO WOOOf 44774 8, 2000 (73) Assignee: University of Pittsburgh-Of the WO WOOOf 72970 12/2000 commonwealth System of Higher WO WOOOf73412 12/2000 WO WOOOf73413 12/2000 Education, Pittsburgh, PA (US) WO WOO1/O1129 1, 2001 WO WOO1? 45845 6, 2001 (*) Notice: Subject to any disclaimer, the term of this WO WO O1, 57253 8, 2001 patent is extended or adjusted under 35 WO WOO1 (84.463 11, 2001 U.S.C. 154(b) by 159 days. WO WO O2, 18902 3, 2002 WO WO O2/O52030 T 2002 (21) Appl. No.: 11/178,134 WO WO O2/O70751 9, 2002 WO WOO3/O55973 T 2003 (22) Filed: Jul. 8, 2005 WO WOO3,O72253 9, 2003 WO WOO3,O77O55 9, 2003 (65) Prior Publication Data WO WO 2004/048931 6, 2004 US 2006/OO1929.0 A1 Jan. -

Genome-Wide Analysis of DNA Methylation, Copy Number Variation, and Gene Expression in Monozygotic Twins Discordant for Primary Biliary Cirrhosis
UC Davis UC Davis Previously Published Works Title Genome-wide analysis of DNA methylation, copy number variation, and gene expression in monozygotic twins discordant for primary biliary cirrhosis. Permalink https://escholarship.org/uc/item/34d4m5nk Journal Frontiers in immunology, 5(MAR) ISSN 1664-3224 Authors Selmi, Carlo Cavaciocchi, Francesca Lleo, Ana et al. Publication Date 2014 DOI 10.3389/fimmu.2014.00128 Peer reviewed eScholarship.org Powered by the California Digital Library University of California ORIGINAL RESEARCH ARTICLE published: 28 March 2014 doi: 10.3389/fimmu.2014.00128 Genome-wide analysis of DNA methylation, copy number variation, and gene expression in monozygotic twins discordant for primary biliary cirrhosis Carlo Selmi 1,2*, Francesca Cavaciocchi 1,3, Ana Lleo4, Cristina Cheroni 5, Raffaele De Francesco5, Simone A. Lombardi 1, Maria De Santis 1,3, Francesca Meda1, Maria Gabriella Raimondo1, Chiara Crotti 1, Marco Folci 1, Luca Zammataro1, Marlyn J. Mayo6, Nancy Bach7, Shinji Shimoda8, Stuart C. Gordon9, Monica Miozzo10,11, Pietro Invernizzi 4, Mauro Podda1, Rossana Scavelli 5, Michelle R. Martin12, Michael F. Seldin13,14, Janine M. LaSalle 12 and M. Eric Gershwin2 1 Division of Rheumatology and Clinical Immunology, Humanitas Clinical and Research Center, Milan, Italy 2 Division of Rheumatology, Allergy, and Clinical Immunology, University of California at Davis, Davis, CA, USA 3 BIOMETRA Department, University of Milan, Milan, Italy 4 Liver Unit and Center for Autoimmune Liver Diseases, Humanitas Clinical and Research Center, Milan, Italy 5 National Institute of Molecular Genetics (INGM), Milan, Italy 6 University of Texas Southwestern, Dallas, TX, USA 7 Mt. Sinai University, NewYork, NY, USA 8 Clinical Research Center, National Nagasaki Medical Center, Nagasaki, Japan 9 Henry Ford Hospital, Detroit, MI, USA 10 Department of Pathophysiology and Transplantation, University of Milan, Milan, Italy 11 Division of Pathology, Fondazione IRCCS Cà Granda Ospedale Maggiore Policlinico, Milan, Italy 12 Genome Center and M.I.N.D. -

Supporting Information Combination of a Novel Gene Expression Signature with a Clinical Nomogram Improves the Prediction of Survival in High-Risk Bladder Cancer
Supporting Information Combination of a novel gene expression signature with a clinical nomogram improves the prediction of survival in high-risk bladder cancer Markus Riester*,1 Jennifer M. Taylor*,2 Andrew Feifer,2 Theresa Koppie,3 Jonathan E. Rosenberg,4 Robert J. Downey,5 Bernard H. Bochner,2 and Franziska Michor1† 1Department of Biostatistics and Computational Biology, Dana-Farber Cancer Institute, and Department of Biostatistics, Harvard School of Public Health, Boston, MA 02215, USA. 2Urology Service, Department of Surgery, Memorial Sloan- Kettering Cancer Center, New York, NY 10065, USA. 3Department of Urology, University of California-Davis, Sacramento, CA. 4Bladder Cancer Center, Dana-Farber Cancer Institute, Boston, MA 02215, USA. 5Thoracic Service, Department of Surgery, Memorial Sloan-Kettering Cancer Center, New York, NY 10065, USA. *These authors contributed equally. †Author for correspondence. Dana-Farber Cancer Institute, 450 Brookline Avenue, Boston, MA 02215, USA. Tel: 671 632 5045. Fax: 617 632 4222. Email: [email protected]. Table of Contents Supplemental Materials and Methods ....................................................................................................................... 2 Datasets .......................................................................................................................................................................................................... 2 Published Signatures ............................................................................................................................................................................... -

NSDHL-Containing Duplication at Xq28 in a Male Patient with Autism
Hu et al. BMC Medical Genetics (2018) 19:192 https://doi.org/10.1186/s12881-018-0705-7 CASEREPORT Open Access NSDHL-containing duplication at Xq28 in a male patient with autism spectrum disorder: a case report Chun-Chun Hu1, Yun-Jun Sun2*, Chun-xue Liu1, Bing-rui Zhou1, Chun-yang Li1, Qiong Xu1 and Xiu Xu1* Abstract Background: Autism spectrum disorder (ASD) is a neurodevelopmental disorder in which genetics plays a key aetiological role. The gene encoding NAD(P)H steroid dehydrogenase-like protein (NSDHL) is expressed in developing cortical neurons and glia, and its mutation may result in intellectual disability or congenital hemidysplasia. Case presentation: An 8-year-old boy presented with a 260-kb NSDHL-containing duplication at Xq28 (151,868,909 – 152,129,300) inherited from his mother. His clinical features included defects in social communication and interaction, restricted interests, attention deficit, impulsive behaviour, minor facial anomalies and serum free fatty acid abnormality. Conclusion: This is the first report of an ASD patient with a related NSDHL-containing duplication at Xq28. Further studies and case reports are required for genetic research to demonstrate that duplication as well as mutation can cause neurodevelopmental diseases. Keywords: Xq28 duplication, NSDHL, Autism, CNV Background 300005), located at Xq28, causing severe X-linked intellec- Autism spectrum disorder (ASD) is a neurodevelopmental tual disability (XLID) [10], and its loss of function causes disorder that is defined in DSM-5 [1] as persistent deficits Rett syndrome (OMIM: 613454) [11, 12]. Duplication of in social communication and social interaction across chromosome 15q11–13, which includes a series of im- multiple contexts in conjunction with restricted, repetitive printing and non-imprinting genes, results in the recur- patterns, interests, or activities as manifested by at least rent cytogenetic abnormalities associated with ASD and two prototypically inflexible behaviours [2]. -

Genome-Wide Analysis of Cancer/Testis Gene Expression
Genome-wide analysis of cancer/testis gene expression Oliver Hofmanna,b,1, Otavia L. Caballeroc, Brian J. Stevensond,e, Yao-Tseng Chenf, Tzeela Cohenc, Ramon Chuac, Christopher A. Maherb, Sumir Panjib, Ulf Schaeferb, Adele Krugerb, Minna Lehvaslaihob, Piero Carnincig,h, Yoshihide Hayashizakig,h, C. Victor Jongeneeld,e, Andrew J. G. Simpsonc, Lloyd J. Oldc,1, and Winston Hidea,b aDepartment of Biostatistics, Harvard School of Public Health, 655 Huntington Avenue, SPH2, 4th Floor, Boston, MA 02115; bSouth African National Bioinformatics Institute, University of the Western Cape, Private Bag X17, Bellville 7535, South Africa; cLudwig Institute for Cancer Research, New York Branch at Memorial Sloan-Kettering Cancer Center, 1275 York Avenue, New York, NY 10021; dLudwig Institute for Cancer Research, Lausanne Branch, 1015 Lausanne, Switzerland; eSwiss Institute of Bioinformatics, 1015 Lausanne, Switzerland; fWeill Medical College of Cornell University, 1300 York Avenue, New York, NY 10021; gGenome Exploration Research Group (Genome Network Project Core Group), RIKEN Genomic Sciences Center (GSC), RIKEN Yokohama Institute, 1-7-22 Suehiro-cho, Tsurumi-ku, Yokohama, Kanagawa, 230-0045, Japan; and hGenome Science Laboratory, Discovery Research Institute, RIKEN Wako Institute, 2-1 Hirosawa, Wako, Saitama, 3510198, Japan Contributed by Lloyd J. Old, October 28, 2008 (sent for review June 6, 2008) Cancer/Testis (CT) genes, normally expressed in germ line cells but expression profile information frequently limited to the original also activated in a wide range of cancer types, often encode defining articles. In some cases, e.g., ACRBP, the original antigens that are immunogenic in cancer patients, and present CT-restricted expression in normal tissues could not be con- potential for use as biomarkers and targets for immunotherapy.