The Secreted L-Arabinose Isomerase Displays Anti-Hyperglycemic Effects
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Comparison of Strand-Specific Transcriptomes Of
Landstorfer et al. BMC Genomics 2014, 15:353 http://www.biomedcentral.com/1471-2164/15/353 RESEARCH ARTICLE Open Access Comparison of strand-specific transcriptomes of enterohemorrhagic Escherichia coli O157:H7 EDL933 (EHEC) under eleven different environmental conditions including radish sprouts and cattle feces Richard Landstorfer1, Svenja Simon2, Steffen Schober3, Daniel Keim2, Siegfried Scherer1 and Klaus Neuhaus1* Abstract Background: Multiple infection sources for enterohemorrhagic Escherichia coli O157:H7 (EHEC) are known, including animal products, fruit and vegetables. The ecology of this pathogen outside its human host is largely unknown and one third of its annotated genes are still hypothetical. To identify genetic determinants expressed under a variety of environmental factors, we applied strand-specific RNA-sequencing, comparing the SOLiD and Illumina systems. Results: Transcriptomes of EHEC were sequenced under 11 different biotic and abiotic conditions: LB medium at pH4, pH7, pH9, or at 15°C; LB with nitrite or trimethoprim-sulfamethoxazole; LB-agar surface, M9 minimal medium, spinach leaf juice, surface of living radish sprouts, and cattle feces. Of 5379 annotated genes in strain EDL933 (genome and plasmid), a surprising minority of only 144 had null sequencing reads under all conditions. We therefore developed a statistical method to distinguish weakly transcribed genes from background transcription. We find that 96% of all genes and 91.5% of the hypothetical genes exhibit a significant transcriptional signal under at least one condition. Comparing SOLiD and Illumina systems, we find a high correlation between both approaches for fold-changes of the induced or repressed genes. The pathogenicity island LEE showed highest transcriptional activity in LB medium, minimal medium, and after treatment with antibiotics. -

Enzymatic Synthesis of L-Fucose from L-Fuculose Using a Fucose Isomerase
Kim et al. Biotechnol Biofuels (2019) 12:282 https://doi.org/10.1186/s13068-019-1619-0 Biotechnology for Biofuels RESEARCH Open Access Enzymatic synthesis of L-fucose from L-fuculose using a fucose isomerase from Raoultella sp. and the biochemical and structural analyses of the enzyme In Jung Kim1†, Do Hyoung Kim1†, Ki Hyun Nam1,2 and Kyoung Heon Kim1* Abstract Background: L-Fucose is a rare sugar with potential uses in the pharmaceutical, cosmetic, and food industries. The enzymatic approach using L-fucose isomerase, which interconverts L-fucose and L-fuculose, can be an efcient way of producing L-fucose for industrial applications. Here, we performed biochemical and structural analyses of L-fucose isomerase identifed from a novel species of Raoultella (RdFucI). Results: RdFucI exhibited higher enzymatic activity for L-fuculose than for L-fucose, and the rate for the reverse reaction of converting L-fuculose to L-fucose was higher than that for the forward reaction of converting L-fucose to L-fuculose. In the equilibrium mixture, a much higher proportion of L-fucose (~ ninefold) was achieved at 30 °C and pH 7, indicating that the enzyme-catalyzed reaction favors the formation of L-fucose from L-fuculose. When biochemical analysis was conducted using L-fuculose as the substrate, the optimal conditions for RdFucI activity were determined to be 40 °C and pH 10. However, the equilibrium composition was not afected by reaction temperature in the range 2 of 30 to 50 °C. Furthermore, RdFucI was found to be a metalloenzyme requiring Mn + as a cofactor. The comparative crystal structural analysis of RdFucI revealed the distinct conformation of α7–α8 loop of RdFucI. -
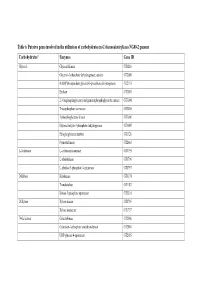
Table 6. Putative Genes Involved in the Utilization of Carbohydrates in G
Table 6. Putative genes involved in the utilization of carbohydrates in G. thermodenitrificans NG80-2 genome Carbohydrates* Enzymes Gene ID Glycerol Glycerol Kinase GT1216 Glycerol-3-phosphate dehydrogenase, aerobic GT2089 NAD(P)H-dependent glycerol-3-phosphate dehydrogenase GT2153 Enolase GT3003 2,3-bisphosphoglycerate-independentphosphoglycerate mutase GT3004 Triosephosphate isomerase GT3005 3-phosphoglycerate kinase GT3006 Glyceraldehyde-3-phosphate dehydrogenase GT3007 Phosphoglycerate mutase GT1326 Pyruvate kinase GT2663 L-Arabinose L-arabinose isomerase GT1795 L-ribulokinase GT1796 L-ribulose 5-phosphate 4-epimerase GT1797 D-Ribose Ribokinase GT3174 Transketolase GT1187 Ribose 5-phosphate epimerase GT3316 D-Xylose Xylose kinase GT1756 Xylose isomerase GT1757 D-Galactose Galactokinase GT2086 Galactose-1-phosphate uridyltransferase GT2084 UDP-glucose 4-epimerase GT2085 Carbohydrates* Enzymes Gene ID D-Fructose 1-phosphofructokinase GT1727 Fructose-1,6-bisphosphate aldolase GT1805 Fructose-1,6-bisphosphate aldolase type II GT3331 Triosephosphate isomerase GT3005 D-Mannose Mannnose-6 phospate isomelase GT3398 6-phospho-1-fructokinase GT2664 D-Mannitol Mannitol-1-phosphate dehydrogenase GT1844 N-Acetylglucosamine N-acetylglucosamine-6-phosphate deacetylase GT2205 N-acetylglucosamine-6-phosphate isomerase GT2204 D-Maltose Alpha-1,4-glucosidase GT0528, GT1643 Sucrose Sucrose phosphorylase GT3215 D-Trehalose Alpha-glucosidase GT1643 Glucose kinase GT2381 Inositol Myo-inositol catabolism protein iolC;5-dehydro-2- GT1807 deoxygluconokinase -

SUPPY Liglucosexlmtdh
US 20100314248A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2010/0314248 A1 Worden et al. (43) Pub. Date: Dec. 16, 2010 (54) RENEWABLE BOELECTRONIC INTERFACE Publication Classification FOR ELECTROBOCATALYTC REACTOR (51) Int. Cl. (76) Inventors: Robert M. Worden, Holt, MI (US); C25B II/06 (2006.01) Brian L. Hassler, Lake Orion, MI C25B II/2 (2006.01) (US); Lawrence T. Drzal, Okemos, GOIN 27/327 (2006.01) MI (US); Ilsoon Lee, Okemo s, MI BSD L/04 (2006.01) (US) C25B 9/00 (2006.01) (52) U.S. Cl. ............... 204/403.14; 204/290.11; 204/400; Correspondence Address: 204/290.07; 427/458; 204/252: 977/734; PRICE HENEVELD COOPER DEWITT & LIT 977/742 TON, LLP 695 KENMOOR, S.E., PO BOX 2567 (57) ABSTRACT GRAND RAPIDS, MI 495.01 (US) An inexpensive, easily renewable bioelectronic device useful for bioreactors, biosensors, and biofuel cells includes an elec (21) Appl. No.: 12/766,169 trically conductive carbon electrode and a bioelectronic inter face bonded to a surface of the electrically conductive carbon (22) Filed: Apr. 23, 2010 electrode, wherein the bioelectronic interface includes cata lytically active material that is electrostatically bound directly Related U.S. Application Data or indirectly to the electrically conductive carbon electrode to (60) Provisional application No. 61/172,337, filed on Apr. facilitate easy removal upon a change in pH, thereby allowing 24, 2009. easy regeneration of the bioelectronic interface. 7\ POWER 1 - SUPPY|- LIGLUCOSEXLMtDH?till pi 6.0 - esses&aaaas-exx-xx-xx-xx-xxxxixax-e- Patent Application Publication Dec. 16, 2010 Sheet 1 of 18 US 2010/0314248 A1 Potential (nV) Patent Application Publication Dec. -

WO 2014/207087 Al 31 December 2014 (31.12.2014) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2014/207087 Al 31 December 2014 (31.12.2014) P O P C T (51) International Patent Classification: Seville (ES). GUTIERREZ GOMEZ, Pablo; Campus C12N 9/00 (2006.01) C12N 9/90 (2006.01) Palmas Altas, Calle Energia Solar 1, E-41014 Seville (ES). C12N 9/02 (2006.01) C12P 5/02 (2006.01) (74) Agent: NEDERLANDSCH OCTROOIBUREAU; J.W. C12N 9/10 (2006.01) C12P 7/76 (2006.01) Frisolaan 13, NL-2517 JS The Hague (NL). C12N 9/12 (2006.01) C12P 7/64 (2006.01) C12N 9/88 (2006.01) C12N 1/1 9 (2006.01) (81) Designated States (unless otherwise indicated, for every kind of national protection available): AE, AG, AL, AM, (21) International Application Number: AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, PCT/EP2014/063484 BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, (22) International Filing Date: DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, 26 June 2014 (26.06.2014) HN, HR, HU, ID, IL, IN, IR, IS, JP, KE, KG, KN, KP, KR, KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, ME, (25) Filing Language: English MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, (26) Publication Language: English OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, (30) Priority Data: TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, 13382241 .1 26 June 2013 (26.06.2013) EP ZW. -

Supplementary Informations SI2. Supplementary Table 1
Supplementary Informations SI2. Supplementary Table 1. M9, soil, and rhizosphere media composition. LB in Compound Name Exchange Reaction LB in soil LBin M9 rhizosphere H2O EX_cpd00001_e0 -15 -15 -10 O2 EX_cpd00007_e0 -15 -15 -10 Phosphate EX_cpd00009_e0 -15 -15 -10 CO2 EX_cpd00011_e0 -15 -15 0 Ammonia EX_cpd00013_e0 -7.5 -7.5 -10 L-glutamate EX_cpd00023_e0 0 -0.0283302 0 D-glucose EX_cpd00027_e0 -0.61972444 -0.04098397 0 Mn2 EX_cpd00030_e0 -15 -15 -10 Glycine EX_cpd00033_e0 -0.0068175 -0.00693094 0 Zn2 EX_cpd00034_e0 -15 -15 -10 L-alanine EX_cpd00035_e0 -0.02780553 -0.00823049 0 Succinate EX_cpd00036_e0 -0.0056245 -0.12240603 0 L-lysine EX_cpd00039_e0 0 -10 0 L-aspartate EX_cpd00041_e0 0 -0.03205557 0 Sulfate EX_cpd00048_e0 -15 -15 -10 L-arginine EX_cpd00051_e0 -0.0068175 -0.00948672 0 L-serine EX_cpd00054_e0 0 -0.01004986 0 Cu2+ EX_cpd00058_e0 -15 -15 -10 Ca2+ EX_cpd00063_e0 -15 -100 -10 L-ornithine EX_cpd00064_e0 -0.0068175 -0.00831712 0 H+ EX_cpd00067_e0 -15 -15 -10 L-tyrosine EX_cpd00069_e0 -0.0068175 -0.00233919 0 Sucrose EX_cpd00076_e0 0 -0.02049199 0 L-cysteine EX_cpd00084_e0 -0.0068175 0 0 Cl- EX_cpd00099_e0 -15 -15 -10 Glycerol EX_cpd00100_e0 0 0 -10 Biotin EX_cpd00104_e0 -15 -15 0 D-ribose EX_cpd00105_e0 -0.01862144 0 0 L-leucine EX_cpd00107_e0 -0.03596182 -0.00303228 0 D-galactose EX_cpd00108_e0 -0.25290619 -0.18317325 0 L-histidine EX_cpd00119_e0 -0.0068175 -0.00506825 0 L-proline EX_cpd00129_e0 -0.01102953 0 0 L-malate EX_cpd00130_e0 -0.03649016 -0.79413596 0 D-mannose EX_cpd00138_e0 -0.2540567 -0.05436649 0 Co2 EX_cpd00149_e0 -

The University of Chicago the Metaproteomic Analysis Of
THE UNIVERSITY OF CHICAGO THE METAPROTEOMIC ANALYSIS OF ARCTIC SOILS WITH NOVEL BIOINFORMATIC METHODS A DISSERTATION SUBMITTED TO THE FACULTY OF THE DIVISION OF THE PHYSICAL SCIENCES IN CANDIDACY FOR THE DEGREE OF DOCTOR OF PHILOSOPHY DEPARTMENT OF THE GEOPHYSICAL SCIENCES BY SAMUEL MILLER CHICAGO, ILLINOIS DECEMBER 2018 COPYRIGHT © 2018 Samuel Edward Miller All Rights Reserved TABLE OF CONTENTS LIST OF FIGURES ................................................................................................................. vii LIST OF TABLES .....................................................................................................................ix ACKNOWLEDGMENTS ........................................................................................................... x ABSTRACT ..............................................................................................................................xi I. INTRODUCTION ................................................................................................................... 1 I.A. IMPETUS FOR RESEARCH ...........................................................................................1 I.B. PROTEOMICS BACKGROUND ....................................................................................2 I.B.1. TANDEM MASS SPECTROMETRY .......................................................................3 I.B.2. METHODS OF AUTOMATIC SEQUENCE ASSIGNMENT ...................................9 I.B.2.i. DE NOVO SEQUENCING BY PEPNOVO+ ................................................... -

O O2 Enzymes Available from Sigma Enzymes Available from Sigma
COO 2.7.1.15 Ribokinase OXIDOREDUCTASES CONH2 COO 2.7.1.16 Ribulokinase 1.1.1.1 Alcohol dehydrogenase BLOOD GROUP + O O + O O 1.1.1.3 Homoserine dehydrogenase HYALURONIC ACID DERMATAN ALGINATES O-ANTIGENS STARCH GLYCOGEN CH COO N COO 2.7.1.17 Xylulokinase P GLYCOPROTEINS SUBSTANCES 2 OH N + COO 1.1.1.8 Glycerol-3-phosphate dehydrogenase Ribose -O - P - O - P - O- Adenosine(P) Ribose - O - P - O - P - O -Adenosine NICOTINATE 2.7.1.19 Phosphoribulokinase GANGLIOSIDES PEPTIDO- CH OH CH OH N 1 + COO 1.1.1.9 D-Xylulose reductase 2 2 NH .2.1 2.7.1.24 Dephospho-CoA kinase O CHITIN CHONDROITIN PECTIN INULIN CELLULOSE O O NH O O O O Ribose- P 2.4 N N RP 1.1.1.10 l-Xylulose reductase MUCINS GLYCAN 6.3.5.1 2.7.7.18 2.7.1.25 Adenylylsulfate kinase CH2OH HO Indoleacetate Indoxyl + 1.1.1.14 l-Iditol dehydrogenase L O O O Desamino-NAD Nicotinate- Quinolinate- A 2.7.1.28 Triokinase O O 1.1.1.132 HO (Auxin) NAD(P) 6.3.1.5 2.4.2.19 1.1.1.19 Glucuronate reductase CHOH - 2.4.1.68 CH3 OH OH OH nucleotide 2.7.1.30 Glycerol kinase Y - COO nucleotide 2.7.1.31 Glycerate kinase 1.1.1.21 Aldehyde reductase AcNH CHOH COO 6.3.2.7-10 2.4.1.69 O 1.2.3.7 2.4.2.19 R OPPT OH OH + 1.1.1.22 UDPglucose dehydrogenase 2.4.99.7 HO O OPPU HO 2.7.1.32 Choline kinase S CH2OH 6.3.2.13 OH OPPU CH HO CH2CH(NH3)COO HO CH CH NH HO CH2CH2NHCOCH3 CH O CH CH NHCOCH COO 1.1.1.23 Histidinol dehydrogenase OPC 2.4.1.17 3 2.4.1.29 CH CHO 2 2 2 3 2 2 3 O 2.7.1.33 Pantothenate kinase CH3CH NHAC OH OH OH LACTOSE 2 COO 1.1.1.25 Shikimate dehydrogenase A HO HO OPPG CH OH 2.7.1.34 Pantetheine kinase UDP- TDP-Rhamnose 2 NH NH NH NH N M 2.7.1.36 Mevalonate kinase 1.1.1.27 Lactate dehydrogenase HO COO- GDP- 2.4.1.21 O NH NH 4.1.1.28 2.3.1.5 2.1.1.4 1.1.1.29 Glycerate dehydrogenase C UDP-N-Ac-Muramate Iduronate OH 2.4.1.1 2.4.1.11 HO 5-Hydroxy- 5-Hydroxytryptamine N-Acetyl-serotonin N-Acetyl-5-O-methyl-serotonin Quinolinate 2.7.1.39 Homoserine kinase Mannuronate CH3 etc. -

12) United States Patent (10
US007635572B2 (12) UnitedO States Patent (10) Patent No.: US 7,635,572 B2 Zhou et al. (45) Date of Patent: Dec. 22, 2009 (54) METHODS FOR CONDUCTING ASSAYS FOR 5,506,121 A 4/1996 Skerra et al. ENZYME ACTIVITY ON PROTEIN 5,510,270 A 4/1996 Fodor et al. MICROARRAYS 5,512,492 A 4/1996 Herron et al. 5,516,635 A 5/1996 Ekins et al. (75) Inventors: Fang X. Zhou, New Haven, CT (US); 5,532,128 A 7/1996 Eggers Barry Schweitzer, Cheshire, CT (US) 5,538,897 A 7/1996 Yates, III et al. s s 5,541,070 A 7/1996 Kauvar (73) Assignee: Life Technologies Corporation, .. S.E. al Carlsbad, CA (US) 5,585,069 A 12/1996 Zanzucchi et al. 5,585,639 A 12/1996 Dorsel et al. (*) Notice: Subject to any disclaimer, the term of this 5,593,838 A 1/1997 Zanzucchi et al. patent is extended or adjusted under 35 5,605,662 A 2f1997 Heller et al. U.S.C. 154(b) by 0 days. 5,620,850 A 4/1997 Bamdad et al. 5,624,711 A 4/1997 Sundberg et al. (21) Appl. No.: 10/865,431 5,627,369 A 5/1997 Vestal et al. 5,629,213 A 5/1997 Kornguth et al. (22) Filed: Jun. 9, 2004 (Continued) (65) Prior Publication Data FOREIGN PATENT DOCUMENTS US 2005/O118665 A1 Jun. 2, 2005 EP 596421 10, 1993 EP 0619321 12/1994 (51) Int. Cl. EP O664452 7, 1995 CI2O 1/50 (2006.01) EP O818467 1, 1998 (52) U.S. -

Genomic, Transcriptomic and Enzymatic Insight Into Lignocellulolytic System of a Plant Pathogen Dickeya Sp
biomolecules Article Genomic, Transcriptomic and Enzymatic Insight into Lignocellulolytic System of a Plant Pathogen Dickeya sp. WS52 to Digest Sweet Pepper and Tomato Stalk 1, 2, 3, 2 2 Ying-Jie Yang y, Wei Lin y, Raghvendra Pratap Singh * , Qian Xu , Zhihou Chen , Yuan Yuan 1 , Ping Zou 1, Yiqiang Li 1 and Chengsheng Zhang 1,* 1 Marine Agriculture Research Center, Tobacco Research Institute of Chinese Academy of Agricultural Sciences, Qingdao 266101, China; [email protected] (Y.-J.Y.); [email protected] (Y.Y.); [email protected] (P.Z.); [email protected] (Y.L.) 2 Tobacco Research Institute of Nanping, Nanping, Fujian 353000, China; [email protected] (W.L.); [email protected] (Q.X.); [email protected] (Z.C.) 3 Department of Research & Development, Biotechnology, Uttaranchal University, Dehradun 248007, India * Correspondence: [email protected] (R.P.S.); [email protected] (C.Z.) These authors contributed equally to this work. y Received: 31 October 2019; Accepted: 18 November 2019; Published: 20 November 2019 Abstract: Dickeya sp., a plant pathogen, causing soft rot with strong pectin degradation capacity was taken for the comprehensive analysis of its corresponding biomass degradative system, which has not been analyzed yet. Whole genome sequence analysis of the isolated soft-rotten plant pathogen Dickeya sp. WS52, revealed various coding genes which involved in vegetable stalk degradation-related properties. A total of 122 genes were found to be encoded for putative carbohydrate-active enzymes (CAZy) in Dickeya sp. WS52. The number of pectin degradation-related genes, was higher than that of cellulolytic bacteria as well as other Dickeya spp. -

The Chemistry and Evolution of Enzyme Function
The Chemistry and Evolution of Enzyme Function: Isomerases as a Case Study Sergio Mart´ınez Cuesta EMBL - European Bioinformatics Institute Gonville and Caius College University of Cambridge A thesis submitted for the degree of Doctor of Philosophy 31st July 2014 This dissertation is the result of my own work and contains nothing which is the outcome of work done in collaboration except where specifically indicated in the text. No part of this dissertation has been submitted or is currently being submitted for any other degree or diploma or other qualification. This thesis does not exceed the specified length limit of 60.000 words as defined by the Biology Degree Committee. This thesis has been typeset in 12pt font using LATEX according to the specifications de- fined by the Board of Graduate Studies and the Biology Degree Committee. Cambridge, 31st July 2014 Sergio Mart´ınezCuesta To my parents and my sister Contents Abstract ix Acknowledgements xi List of Figures xiii List of Tables xv List of Publications xvi 1 Introduction 1 1.1 Chemistry of enzymes . .2 1.1.1 Catalytic sites, mechanisms and cofactors . .3 1.1.2 Enzyme classification . .5 1.2 Evolution of enzyme function . .6 1.3 Similarity between enzymes . .8 1.3.1 Comparing sequences and structures . .8 1.3.2 Comparing genomic context . .9 1.3.3 Comparing biochemical reactions and mechanisms . 10 1.4 Isomerases . 12 1.4.1 Metabolism . 13 1.4.2 Genome . 14 1.4.3 EC classification . 15 1.4.4 Applications . 18 1.5 Structure of the thesis . 20 2 Data Resources and Methods 21 2.1 Introduction . -

All Enzymes in BRENDA™ the Comprehensive Enzyme Information System
All enzymes in BRENDA™ The Comprehensive Enzyme Information System http://www.brenda-enzymes.org/index.php4?page=information/all_enzymes.php4 1.1.1.1 alcohol dehydrogenase 1.1.1.B1 D-arabitol-phosphate dehydrogenase 1.1.1.2 alcohol dehydrogenase (NADP+) 1.1.1.B3 (S)-specific secondary alcohol dehydrogenase 1.1.1.3 homoserine dehydrogenase 1.1.1.B4 (R)-specific secondary alcohol dehydrogenase 1.1.1.4 (R,R)-butanediol dehydrogenase 1.1.1.5 acetoin dehydrogenase 1.1.1.B5 NADP-retinol dehydrogenase 1.1.1.6 glycerol dehydrogenase 1.1.1.7 propanediol-phosphate dehydrogenase 1.1.1.8 glycerol-3-phosphate dehydrogenase (NAD+) 1.1.1.9 D-xylulose reductase 1.1.1.10 L-xylulose reductase 1.1.1.11 D-arabinitol 4-dehydrogenase 1.1.1.12 L-arabinitol 4-dehydrogenase 1.1.1.13 L-arabinitol 2-dehydrogenase 1.1.1.14 L-iditol 2-dehydrogenase 1.1.1.15 D-iditol 2-dehydrogenase 1.1.1.16 galactitol 2-dehydrogenase 1.1.1.17 mannitol-1-phosphate 5-dehydrogenase 1.1.1.18 inositol 2-dehydrogenase 1.1.1.19 glucuronate reductase 1.1.1.20 glucuronolactone reductase 1.1.1.21 aldehyde reductase 1.1.1.22 UDP-glucose 6-dehydrogenase 1.1.1.23 histidinol dehydrogenase 1.1.1.24 quinate dehydrogenase 1.1.1.25 shikimate dehydrogenase 1.1.1.26 glyoxylate reductase 1.1.1.27 L-lactate dehydrogenase 1.1.1.28 D-lactate dehydrogenase 1.1.1.29 glycerate dehydrogenase 1.1.1.30 3-hydroxybutyrate dehydrogenase 1.1.1.31 3-hydroxyisobutyrate dehydrogenase 1.1.1.32 mevaldate reductase 1.1.1.33 mevaldate reductase (NADPH) 1.1.1.34 hydroxymethylglutaryl-CoA reductase (NADPH) 1.1.1.35 3-hydroxyacyl-CoA