Mouse Rabepk Conditional Knockout Project (CRISPR/Cas9)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Genetic and Genomic Analysis of Hyperlipidemia, Obesity and Diabetes Using (C57BL/6J × TALLYHO/Jngj) F2 Mice
University of Tennessee, Knoxville TRACE: Tennessee Research and Creative Exchange Nutrition Publications and Other Works Nutrition 12-19-2010 Genetic and genomic analysis of hyperlipidemia, obesity and diabetes using (C57BL/6J × TALLYHO/JngJ) F2 mice Taryn P. Stewart Marshall University Hyoung Y. Kim University of Tennessee - Knoxville, [email protected] Arnold M. Saxton University of Tennessee - Knoxville, [email protected] Jung H. Kim Marshall University Follow this and additional works at: https://trace.tennessee.edu/utk_nutrpubs Part of the Animal Sciences Commons, and the Nutrition Commons Recommended Citation BMC Genomics 2010, 11:713 doi:10.1186/1471-2164-11-713 This Article is brought to you for free and open access by the Nutrition at TRACE: Tennessee Research and Creative Exchange. It has been accepted for inclusion in Nutrition Publications and Other Works by an authorized administrator of TRACE: Tennessee Research and Creative Exchange. For more information, please contact [email protected]. Stewart et al. BMC Genomics 2010, 11:713 http://www.biomedcentral.com/1471-2164/11/713 RESEARCH ARTICLE Open Access Genetic and genomic analysis of hyperlipidemia, obesity and diabetes using (C57BL/6J × TALLYHO/JngJ) F2 mice Taryn P Stewart1, Hyoung Yon Kim2, Arnold M Saxton3, Jung Han Kim1* Abstract Background: Type 2 diabetes (T2D) is the most common form of diabetes in humans and is closely associated with dyslipidemia and obesity that magnifies the mortality and morbidity related to T2D. The genetic contribution to human T2D and related metabolic disorders is evident, and mostly follows polygenic inheritance. The TALLYHO/ JngJ (TH) mice are a polygenic model for T2D characterized by obesity, hyperinsulinemia, impaired glucose uptake and tolerance, hyperlipidemia, and hyperglycemia. -

1 Title 1 Loss of PABPC1 Is Compensated by Elevated PABPC4
bioRxiv preprint doi: https://doi.org/10.1101/2021.02.07.430165; this version posted February 15, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 1 Title 2 Loss of PABPC1 is compensated by elevated PABPC4 and correlates with transcriptome 3 changes 4 5 Jingwei Xie1, 2, Xiaoyu Wei1, Yu Chen1 6 7 1 Department of Biochemistry and Groupe de recherche axé sur la structure des 8 protéines, McGill University, Montreal, Quebec H3G 0B1, Canada 9 10 2 To whom correspondence should be addressed: Dept. of Biochemistry, McGill 11 University, Montreal, QC H3G 0B1, Canada. E-mail: [email protected]. 12 13 14 15 Abstract 16 Cytoplasmic poly(A) binding protein (PABP) is an essential translation factor that binds to 17 the 3' tail of mRNAs to promote translation and regulate mRNA stability. PABPC1 is the 18 most abundant of several PABP isoforms that exist in mammals. Here, we used the 19 CRISPR/Cas genome editing system to shift the isoform composition in HEK293 cells. 20 Disruption of PABPC1 elevated PABPC4 levels. Transcriptome analysis revealed that the 21 shift in the dominant PABP isoform was correlated with changes in key transcriptional 22 regulators. This study provides insight into understanding the role of PABP isoforms in 23 development and differentiation. 24 Keywords 25 PABPC1, PABPC4, c-Myc 26 bioRxiv preprint doi: https://doi.org/10.1101/2021.02.07.430165; this version posted February 15, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

RAB9 Antibody (R32125)
RAB9 Antibody (R32125) Catalog No. Formulation Size R32125 0.5mg/ml if reconstituted with 0.2ml sterile DI water 100 ug Bulk quote request Availability 1-3 business days Species Reactivity Human, Mouse, Rat Format Antigen affinity purified Clonality Polyclonal (rabbit origin) Isotype Rabbit IgG Purity Antigen affinity Buffer Lyophilized from 1X PBS with 2.5% BSA and 0.025% sodium azide UniProt P51151 Applications Western blot : 0.1-0.5ug/ml Limitations This RAB9 antibody is available for research use only. Western blot testing of 1) rat brain, 2) mouse brain, 3) human HeLa and 4) human MCF7 lysate with RAB9 antibody. Expected/observed molecular weight: ~23 kDa. Description Ras-related protein Rab-9A, also called RAB9, is a protein that in humans is encoded by the RAB9A gene. This gene is mapped to Xp22.2. RAB9 has been localized to components of the endocytic/exocytic pathway and has been implicated in recycling of membrane receptors. It has been found that downregulation of RAB9A gene expression in HeLa cells induced severe cell vacuolation. RAB9A GTPase is directly bound by TIP47 in its active, GTP-bound conformation. Moreover, RAB9A increases the affinity of TIP47 for its cargo. What’s more, this gene may involved in the transport of proteins between the endosomes and the trans Golgi network. RAB9A has been shown to interact with RABEPK and TIP47. Application Notes Optimal dilution of the RAB9 antibody should be determined by the researcher. Immunogen Amino acids KDATNVAAAFEEAVRRVLATEDRSDHLIQTDTVNLHRK of human RAB9A were used as the immunogen for the RAB9 antibody. Storage After reconstitution, the RAB9 antibody can be stored for up to one month at 4oC. -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Mapping of Craniofacial Traits in Outbred Mice Identifies Major Developmental Genes Involved in Shape Determination
Mapping of craniofacial traits in outbred mice identifies major developmental genes involved in shape determination Luisa F Pallares1, Peter Carbonetto2,3, Shyam Gopalakrishnan2,4, Clarissa C Parker2,5, Cheryl L Ackert-Bicknell6, Abraham A Palmer2,7, Diethard Tautz1 # 1Max Planck Institute for Evolutionary Biology, Plön, Germany 2University of Chicago, Chicago, Illinois, USA 3AncestryDNA, San Francisco, California, USA 4Museum of Natural History, Copenhagen University, Copenhagen, Denmark 5Middlebury College, Department of Psychology and Program in Neuroscience, Middlebury VT, USA 6Center for Musculoskeletal Research, University of Rochester, Rochester, NY USA 7University of California San Diego, La Jolla, CA, USA # corresponding author: [email protected] short title: craniofacial shape mapping Abstract The vertebrate cranium is a prime example of the high evolvability of complex traits. While evidence of genes and developmental pathways underlying craniofacial shape determination 1 is accumulating, we are still far from understanding how such variation at the genetic level is translated into craniofacial shape variation. Here we used 3D geometric morphometrics to map genes involved in shape determination in a population of outbred mice (Carworth Farms White, or CFW). We defined shape traits via principal component analysis of 3D skull and mandible measurements. We mapped genetic loci associated with shape traits at ~80,000 candidate single nucleotide polymorphisms in ~700 male mice. We found that craniofacial shape and size are highly heritable, polygenic traits. Despite the polygenic nature of the traits, we identified 17 loci that explain variation in skull shape, and 8 loci associated with variation in mandible shape. Together, the associated variants account for 11.4% of skull and 4.4% of mandible shape variation, however, the total additive genetic variance associated with phenotypic variation was estimated in ~45%. -

Understanding and Improving the Identification of Somatic Variants
Understanding and Improving the Identification of Somatic Variants Vinaya Vijayan Dissertation submitted to the faculty of the Virginia Polytechnic Institute and State University in partial fulfillment of the requirements for the degree of Doctor of Philosophy In Genetics, Bioinformatics and Computational Biology Chair: Liqing Zhang Christopher Franck Lenwood S. Heath Xiaowei Wu August 12th, 2016 Blacksburg, VA, USA Keywords: Somatic variants, Somatic variant callers, Somatic point mutations, Short tandem repeat variation, Lung squamous cell carcinoma Understanding and Improving the Identification of Somatic Variants Vinaya Vijayan ABSTRACT It is important to understand the entire spectrum of somatic variants to gain more insight into mutations that occur in different cancers for development of better diagnostic, prognostic and therapeutic tools. This thesis outlines our work in understanding somatic variant calling, improving the identification of somatic variants from whole genome and whole exome platforms and identification of biomarkers for lung cancer. Integrating somatic variants from whole genome and whole exome platforms poses a challenge as variants identified in the exonic regions of the whole genome platform may not be identified on the whole exome platform and vice-versa. Taking a simple union or intersection of the somatic variants from both platforms would lead to inclusion of many false positives (through union) and exclusion of many true variants (through intersection). We develop the first framework to improve the identification of somatic variants on whole genome and exome platforms using a machine learning approach by combining the results from two popular somatic variant callers. Testing on simulated and real data sets shows that our framework identifies variants more accurately than using only one somatic variant caller or using variants from only one platform. -

Proteomicsdb: a Multi-Omics and Multi-Organism Resource for Life Science
Published online 30 October 2019 Nucleic Acids Research, 2020, Vol. 48, Database issue D1153–D1163 doi: 10.1093/nar/gkz974 ProteomicsDB: a multi-omics and multi-organism resource for life science research Patroklos Samaras 1, Tobias Schmidt 1,MartinFrejno1, Siegfried Gessulat1,2, Maria Reinecke1,3,4, Anna Jarzab1, Jana Zecha1, Julia Mergner1, Piero Giansanti1, Hans-Christian Ehrlich2, Stephan Aiche2, Johannes Rank5,6, Harald Kienegger5,6, Helmut Krcmar5,6, Bernhard Kuster1,7,* and Mathias Wilhelm1,* 1Chair of Proteomics and Bioanalytics, Technical University of Munich (TUM), Freising, Bavaria, Germany, 2Innovation Center Network, SAP SE, Potsdam, Germany, 3German Cancer Consortium (DKTK), Partner Site Munich, Munich, Germany, 4German Cancer Research Center (DKFZ), Heidelberg, Germany, 5Chair for Information Systems, Technical University of Munich (TUM), Garching, Germany, 6SAP University Competence Center, Technical University of Munich (TUM), Garching, Germany and 7Bavarian Biomolecular Mass Spectrometry Center (BayBioMS), Technical University of Munich (TUM), Freising, Bavaria, Germany Received September 14, 2019; Revised October 11, 2019; Editorial Decision October 11, 2019; Accepted October 15, 2019 ABSTRACT INTRODUCTION ProteomicsDB (https://www.ProteomicsDB.org) ProteomicsDB (https://www.ProteomicsDB.org) is an in- started as a protein-centric in-memory database for memory database initially developed for the exploration of the exploration of large collections of quantitative large quantities of quantitative human mass spectrometry- mass spectrometry-based proteomics data. The based proteomics data including the first draft of the hu- data types and contents grew over time to include man proteome (1). Among many features, it allows the real- time exploration and retrieval of protein abundance values RNA-Seq expression data, drug-target interactions across different tissues, cell lines, and body fluids via inter- and cell line viability data. -

Predict AID Targeting in Non-Ig Genes Multiple Transcription Factor
Downloaded from http://www.jimmunol.org/ by guest on September 26, 2021 is online at: average * The Journal of Immunology published online 20 March 2013 from submission to initial decision 4 weeks from acceptance to publication Multiple Transcription Factor Binding Sites Predict AID Targeting in Non-Ig Genes Jamie L. Duke, Man Liu, Gur Yaari, Ashraf M. Khalil, Mary M. Tomayko, Mark J. Shlomchik, David G. Schatz and Steven H. Kleinstein J Immunol http://www.jimmunol.org/content/early/2013/03/20/jimmun ol.1202547 Submit online. Every submission reviewed by practicing scientists ? is published twice each month by http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://www.jimmunol.org/content/suppl/2013/03/20/jimmunol.120254 7.DC1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 26, 2021. Published March 20, 2013, doi:10.4049/jimmunol.1202547 The Journal of Immunology Multiple Transcription Factor Binding Sites Predict AID Targeting in Non-Ig Genes Jamie L. Duke,* Man Liu,†,1 Gur Yaari,‡ Ashraf M. Khalil,x Mary M. Tomayko,{ Mark J. Shlomchik,†,x David G. -
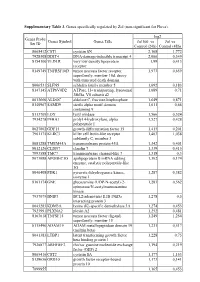
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis -
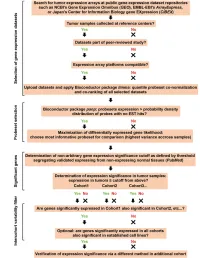
Supplementary Data
SUPPLEMENTAL INFORMATION A study restricted to chemokine receptors as well as a genome-wide transcript analysis uncovered CXCR4 as preferentially expressed in Ewing's sarcoma (Ewing's sarcoma) cells of metastatic origin (Figure 4). Transcriptome analyses showed that in addition to CXCR4, genes known to support cell motility and invasion topped the list of genes preferentially expressed in metastasis-derived cells (Figure 4D). These included kynurenine 3-monooxygenase (KMO), galectin-1 (LGALS1), gastrin-releasing peptide (GRP), procollagen C-endopeptidase enhancer (PCOLCE), and ephrin receptor B (EPHB3). KMO, a key enzyme of tryptophan catabolism, has not been linked to metastasis. Tryptophan and its catabolites, however, are involved in immune evasion by tumors, a process that can assist in tumor progression and metastasis (1). LGALS1, GRP, PCOLCE and EPHB3 have been linked to tumor progression and metastasis of several cancers (2-4). Top genes preferentially expressed in L-EDCL included genes that suppress cell motility and/or potentiate cell adhesion such as plakophilin 1 (PKP1), neuropeptide Y (NPY), or the metastasis suppressor TXNIP (5-7) (Figure 4D). Overall, L-EDCL were enriched in gene sets geared at optimizing nutrient transport and usage (Figure 4D; Supplementary Table 3), a state that may support the early stages of tumor growth. Once tumor growth outpaces nutrient and oxygen supplies, gene expression programs are usually switched to hypoxic response and neoangiogenesis, which ultimately lead to tumor egress and metastasis. Accordingly, gene sets involved in extracellular matrix remodeling, MAPK signaling, and response to hypoxia were up-regulated in M-EDCL (Figure 4D; Supplementary Table 4), consistent with their association to metastasis in other cancers (8, 9). -

Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations Fall 2010 Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress Renuka Nayak University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Computational Biology Commons, and the Genomics Commons Recommended Citation Nayak, Renuka, "Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress" (2010). Publicly Accessible Penn Dissertations. 1559. https://repository.upenn.edu/edissertations/1559 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/1559 For more information, please contact [email protected]. Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress Abstract Genes interact in networks to orchestrate cellular processes. Here, we used coexpression networks based on natural variation in gene expression to study the functions and interactions of human genes. We asked how these networks change in response to stress. First, we studied human coexpression networks at baseline. We constructed networks by identifying correlations in expression levels of 8.9 million gene pairs in immortalized B cells from 295 individuals comprising three independent samples. The resulting networks allowed us to infer interactions between biological processes. We used the network to predict the functions of poorly-characterized human genes, and provided some experimental support. Examining genes implicated in disease, we found that IFIH1, a diabetes susceptibility gene, interacts with YES1, which affects glucose transport. Genes predisposing to the same diseases are clustered non-randomly in the network, suggesting that the network may be used to identify candidate genes that influence disease susceptibility.