JANG-DISSERTATION-2017.Pdf (6.114Mb)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Functional Compensation Among HMGN Variants Modulates the Dnase I Hypersensitive Sites at Enhancers
Downloaded from genome.cshlp.org on October 9, 2021 - Published by Cold Spring Harbor Laboratory Press Research Functional compensation among HMGN variants modulates the DNase I hypersensitive sites at enhancers Tao Deng,1,12 Z. Iris Zhu,2,12 Shaofei Zhang,1 Yuri Postnikov,1 Di Huang,2 Marion Horsch,3 Takashi Furusawa,1 Johannes Beckers,3,4,5 Jan Rozman,3,5 Martin Klingenspor,6,7 Oana Amarie,3,8 Jochen Graw,3,8 Birgit Rathkolb,3,5,9 Eckhard Wolf,9 Thure Adler,3 Dirk H. Busch,10 Valérie Gailus-Durner,3 Helmut Fuchs,3 Martin Hrabeˇ de Angelis,3,4,5 Arjan van der Velde,2,13 Lino Tessarollo,11 Ivan Ovcherenko,2 David Landsman,2 and Michael Bustin1 1Protein Section, Laboratory of Metabolism, Center for Cancer Research, National Cancer Institute, National Institutes of Health, Bethesda, Maryland 20892, USA; 2Computational Biology Branch, National Center for Biotechnology Information, National Library of Medicine, Bethesda, Maryland 20892, USA; 3German Mouse Clinic, Institute of Experimental Genetics, Helmholtz Zentrum München, German Research Center for Environmental Health, 85764 Neuherberg, Germany; 4Experimental Genetics, Center of Life and Food Sciences Weihenstephan, Technische Universität München, 85354 Freising-Weihenstephan, Germany; 5German Center for Diabetes Research (DZD), 85764 Neuherberg, Germany; 6Molecular Nutritional Medicine, Technische Universität München, 85350 Freising, Germany; 7Center for Nutrition and Food Sciences, Technische Universität München, 85350 Freising, Germany; 8Institute of Developmental Genetics (IDG), 85764 -

HMGB1 in Health and Disease R
Donald and Barbara Zucker School of Medicine Journal Articles Academic Works 2014 HMGB1 in health and disease R. Kang R. C. Chen Q. H. Zhang W. Hou S. Wu See next page for additional authors Follow this and additional works at: https://academicworks.medicine.hofstra.edu/articles Part of the Emergency Medicine Commons Recommended Citation Kang R, Chen R, Zhang Q, Hou W, Wu S, Fan X, Yan Z, Sun X, Wang H, Tang D, . HMGB1 in health and disease. 2014 Jan 01; 40():Article 533 [ p.]. Available from: https://academicworks.medicine.hofstra.edu/articles/533. Free full text article. This Article is brought to you for free and open access by Donald and Barbara Zucker School of Medicine Academic Works. It has been accepted for inclusion in Journal Articles by an authorized administrator of Donald and Barbara Zucker School of Medicine Academic Works. Authors R. Kang, R. C. Chen, Q. H. Zhang, W. Hou, S. Wu, X. G. Fan, Z. W. Yan, X. F. Sun, H. C. Wang, D. L. Tang, and +8 additional authors This article is available at Donald and Barbara Zucker School of Medicine Academic Works: https://academicworks.medicine.hofstra.edu/articles/533 NIH Public Access Author Manuscript Mol Aspects Med. Author manuscript; available in PMC 2015 December 01. NIH-PA Author ManuscriptPublished NIH-PA Author Manuscript in final edited NIH-PA Author Manuscript form as: Mol Aspects Med. 2014 December ; 0: 1–116. doi:10.1016/j.mam.2014.05.001. HMGB1 in Health and Disease Rui Kang1,*, Ruochan Chen1, Qiuhong Zhang1, Wen Hou1, Sha Wu1, Lizhi Cao2, Jin Huang3, Yan Yu2, Xue-gong Fan4, Zhengwen Yan1,5, Xiaofang Sun6, Haichao Wang7, Qingde Wang1, Allan Tsung1, Timothy R. -
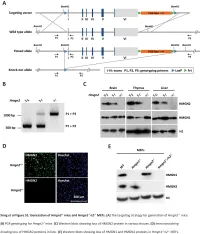
Supplemental Data.Pdf
Table S1. Summary of sequencing results A. DNase‐seq data % Align % Mismatch % >=Q30 bases Sample ID # Reads % unique reads (PF) Rate (PF) (PF) WT_1 84,301,522 86.76 0.52 93.34 96.30 WT_2 98,744,222 84.97 0.51 93.75 89.94 Hmgn1‐/‐_1 79,620,656 83.94 0.86 90.58 88.65 Hmgn1‐/‐_2 62,673,782 84.13 0.87 91.30 89.18 Hmgn2‐/‐_1 87,734,440 83.49 0.71 91.81 90.00 Hmgn2‐/‐_2 82,498,808 83.25 0.69 92.73 90.66 Hmgn1‐/‐n2‐/‐_1 71,739,638 68.51 2.31 81.11 89.22 Hmgn1‐/‐n2‐/‐_2 74,113,682 68.19 2.37 81.16 86.57 B. ChIP‐seq data Histone % Align % Mismatch % >=Q30 % unique Genotypes # Reads marks (PF) Rate (PF) bases (PF) reads H3K4me1 100670054 92.99 0.28 91.21 87.29 H3K4me3 67064272 91.97 0.35 89.11 27.15 WT H3K27ac 90,340,242 93.57 0.28 95.02 89.80 input 111,292,572 78.24 0.55 96.07 86.99 H3K4me1 84598176 92.34 0.33 91.2 81.69 H3K4me3 90032064 92.19 0.44 88.76 15.81 Hmgn1‐/‐n2‐/‐ H3K27ac 86,260,526 93.40 0.29 94.94 87.49 input 78,142,334 78.47 0.56 95.82 81.11 C. MNase‐seq data % Mismatch % >=Q30 bases % unique Sample ID # Reads % Align (PF) Rate (PF) (PF) reads WT_1_Extensive 45,232,694 55.23 1.49 90.22 81.73 WT_1_Limited 105,460,950 58.03 1.39 90.81 79.62 WT_2_Extensive 40,785,338 67.34 1.06 89.76 89.60 WT_2_Limited 105,738,078 68.34 1.05 90.29 85.96 Hmgn1‐/‐n2‐/‐_1_Extensive 117,927,050 55.74 1.49 89.50 78.01 Hmgn1‐/‐n2‐/‐_1_Limited 61,846,742 63.76 1.22 90.57 84.55 Hmgn1‐/‐n2‐/‐_2_Extensive 137,673,830 60.04 1.30 89.28 78.99 Hmgn1‐/‐n2‐/‐_2_Limited 45,696,614 62.70 1.21 90.71 85.52 D. -

Binding RXFP1 in Brain Tumors
Functional role of C1Q-TNF related peptide 8 (CTRP8)- binding RXFP1 in brain tumors By THATCHAWAN THANASUPAWAT A Thesis submitted to the Faculty of Graduate Studies of The University of Manitoba in partial fulfillment of the requirements of the degree of DOCTOR OF PHILOSOPHY Department of Human Anatomy and Cell Science University of Manitoba Winnipeg, Manitoba Canada Copyright © 2017 by Thatchawan Thanasupawat TABLE OF CONTENTS ABSTRACT ....................................................................................................................... v ACKNOWLEDGEMENTS ........................................................................................... vii LIST OF TABLES ........................................................................................................... ix LIST OF FIGURES .......................................................................................................... x LIST OF ABBREVIATIONS ........................................................................................ xii LIST OF COPYRIGHTED MATERIALS ................................................................. xix CHAPTER 1: INTRODUCTION .................................................................................... 1 1.1 Relaxin family peptide receptors (RXFPs) ............................................................... 1 1.2 Relaxin family peptide receptor 1 (RXFP1) ............................................................. 1 1.2.1 Structural features of RXFP1 ............................................................................ -

Identification of Proteins Involved in the Maintenance of Genome Stability
Identification of Proteins Involved in the Maintenance of Genome Stability by Edith Hang Yu Cheng A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Department of Biochemistry University of Toronto ©Copyright by Edith Cheng2015 Identification of Proteins Involved in the Maintenance of Genome Stability Edith Cheng Doctor of Philosophy Department of Biochemistry University of Toronto 2015 Abstract Aberrant changes to the genome structure underlie numerous human diseases such as cancers. The functional characterization ofgenesand proteins that maintain chromosome stability will be important in understanding disease etiology and developing therapeutics. I took a multi-faceted approach to identify and characterize genes involved in the maintenance of genome stability. As biological pathways involved in genome maintenance are highly conserved in evolution, results from model organisms can greatly facilitate functional discovery in humans. In S. cerevisiae, I identified 47 essential gene depletions with elevated levels of spontaneous DNA damage foci and 92 depletions that caused elevated levels of chromosome rearrangements. Of these, a core subset of 15 DNA replication genes demonstrated both phenotypes when depleted. Analysis of rearrangement breakpoints revealed enrichment at yeast fragile sites, Ty retrotransposons, early origins of replication and replication termination sites. Together, thishighlighted the integral role of DNA replicationin genome maintenance. In light of my findings in S. cerevisiae, I identified a list of 153 human proteins that interact with the nascentDNA at replication forks, using a DNA pull down strategy (iPOND) in human cell lines. As a complementary approach for identifying human proteins involved in genome ii maintenance, I usedthe BioID techniqueto discernin vivo proteins proximal to the human BLM- TOP3A-RMI1-RMI2 genome stability complex, which has an emerging role in DNA replication progression. -

The Hmgn Family of Chromatin Binding Proteins Regulate Stem Cell Pluripotency and Neuronal Differentiation
The Hmgn family of chromatin binding proteins regulate stem cell pluripotency and neuronal differentiation Gokula Mohan, Ohoud Rehbini, Tomoko Iwata, Katherine West Institute of Cancer Sciences, University of Glasgow Hmgn proteins Hmgn proteins are enriched across Hmgn2 is expressed in neural progenitor cells the Oct4 and Nanog gene loci during embryogenesis E11.5 E13.5 E15.5 Bind as dimers to nucleosomes svz Hmgn1 Hmgn2 svz 3 Regulate the deposition of histone 3 3 2.5 modifications 2.5 2.5 2 2 2 1.5 Compete with linker histones 1.5 1.5 1 enrichment 1 1 Interact with several transcription 0.5 0.5 0.5 factors. 0 0 0 svz: ventricular/ -1781 -399 410 2070 3342 -1081 -219 929 1740 2636 Actin Msat c subventricular zone Kato H et al. PNAS 2011;108:12283-12288 Oct4 Nanog H3 dg dg: dentate gyrus (in hippocampus) control cells NLS Nucleosome-binding domain NLS Regulatory domain 2 2 2 very highly conserved Competes with H1 c: cerebellum 1.5 1.5 1.5 100 aa 1 1 1 P14 Inhibits phosphorylation Increases acetylation enrichment 0.5 0.5 0.5 of H3S10 of H3K14 0 0 0 -1781 -399 410 2070 3342 -1081 -219 929 1740 2636 Actin Msat Retinoic acid-induced differentiation of EC P Oct4 Nanog cells down the neuronal lineage. M A A P Reprogramming Differentiation Hmgn1 and Hmgn2 binding profiles in control cells. Enrichment at each PCR set was normalised to Histone H3 N-terminal tail ARTKQTARKSTGGKAPRKQLATKAARKSA... stage stage input and then normalised to average H3 enrichment across all primer sets to control for differences M between chromatin preps. -

HMGA2-Mediated Epigenetic Regulation of Gata6 Controls Epithelial Canonical WNT Signaling During Lung Development and Homeostasis
HMGA2-mediated epigenetic regulation of Gata6 controls epithelial canonical WNT signaling during lung development and homeostasis INAUGURALDISSERTATION zur Erlangung des Doktorgrades der Naturwissenschaften - Doctor rerum naturalium - (Dr. rer. nat.) eingereicht am Fachbereich Biologie und Chemie der Justus-Liebig-Universität Giessen vorgelegt von Indrabahadur Singh Bad Nauheim, 2014 Die vorliegende Arbeit wurde am Max-Planck-Institut für Herz- und Lungenforschung in der Abteilung "Epigenetik des Lungenkrebs" unter der Leitung von Herrn Dr. Guillermo Barreto angefertigt. Erstgutachter: Prof. Dr. Dr. Thomas Braun Abteilung Entwicklung und Umbau des Herzens Max-Planck-Institut für Herz- und Lungenforschung Ludwigstraße 43, 61231 Bad Nauheim Zweitgutachter: Prof. Dr. Rainer Renkawitz Institut für Genetik Justus-Liebig-Universität Giessen Heinrich-Buff-Ring 58, 35392 Giessen Disputation am 24-09-2014 DEDICATED TO MY PARENTS & GRANDPARENTS Whose perpetual affection and blessing always Inspired me for higher ambition in life TABLE OF CONTENTS Table of Contents Table of Contents ....................................................................................................................... VIII List of Figures and Tables............................................................................................................. XI ZUSAMMENFASSUNG ......................................................................................................... XIV ABSTRACT .................................................................................................................................. -

The HMGA Proteins: a Myriad of Functions (Review)
289-305 9/1/08 10:59 Page 289 INTERNATIONAL JOURNAL OF ONCOLOGY 32: 289-305, 2008 289 The HMGA proteins: A myriad of functions (Review) ISABELLE CLEYNEN and WIM J.M. VAN DE VEN Laboratory of Molecular Oncology, Department of Human Genetics, University of Leuven, Herestraat 49, B-3000 Leuven, Belgium Received September 28, 2007; Accepted November 5, 2007 Abstract. The ‘high mobility group’ HMGA protein family 3. The HMGA genes and corresponding proteins consists of four members: HMGA1a, HMGA1b and HMGA1c, 4. The HMGA proteins: mechanism of action which result from translation of alternative spliced forms of 5 Cellular functions of the HMGA proteins one gene and HMGA2, which is encoded for by another 6. Regulation of HMGA expression gene. HMGA proteins are characterized by three DNA-binding 7. Conclusion domains, called AT-hooks, and an acidic carboxy-terminal tail. HMGA proteins are architectural transcription factors that both positively and negatively regulate the transcription 1. Introduction of a variety of genes. They do not display direct transcriptional activation capacity, but regulate gene expression by changing Life depends on the ability of a cell to store, retrieve, process the DNA conformation by binding to AT-rich regions in and translate the genetic instructions required to make and the DNA and/or direct interaction with several transcription maintain a living organism. This hereditary information is factors. In this way, they influence a diverse array of normal passed on from a cell to its daughter cells at cell division, and biological processes including cell growth, proliferation, from generation to generation of organisms through the differentiation and death. -

E2F1 Coregulates Cell Cycle Genes and Chromatin Components During the Transition of Oligodendrocyte Progenitors from Proliferation to Differentiation
The Journal of Neuroscience, January 22, 2014 • 34(4):1481–1493 • 1481 Development/Plasticity/Repair E2F1 Coregulates Cell Cycle Genes and Chromatin Components during the Transition of Oligodendrocyte Progenitors from Proliferation to Differentiation Laura Magri,1* Victoria A. Swiss,1* Beata Jablonska,4 Liang Lei,5 Xiomara Pedre,1 Martin Walsh,2 Weijia Zhang,3 Vittorio Gallo,4 Peter Canoll,5 and Patrizia Casaccia1,2 Departments of 1Neuroscience, 2Genetics and Genomics, and 3Medicine, Icahn School of Medicine at Mount Sinai, New York, New York 10029, 4Center for Neuroscience Research, Children’s National Medical Center, George Washington University, Washington, DC 20010-2970, and 5Department of Pathology and Cell Biology, Columbia University Medical Center, New York, New York 10032 Cell cycle exit is an obligatory step for the differentiation of oligodendrocyte progenitor cells (OPCs) into myelinating cells. A key regulator of the transition from proliferation to quiescence is the E2F/Rb pathway, whose activity is highly regulated in physiological conditions and deregulated in tumors. In this paper we report a lineage-specific decline of nuclear E2F1 during differentiation of rodent OPC into oligodendrocytes (OLs) in developing white matter tracts and in cultured cells. Using chromatin immunoprecipitation (ChIP) and deep-sequencing in mouse and rat OPCs, we identified cell cycle genes (i.e., Cdc2) and chromatin components (i.e., Hmgn1, Hmgn2), including those modulating DNA methylation (i.e., Uhrf1), as E2F1 targets. Binding of E2F1 to chromatin on the gene targets was validated and their expression assessed in developing white matter tracts and cultured OPCs. Increased expression of E2F1 gene targets was also detected in mouse gliomas (that were induced by retroviral transformation of OPCs) compared with normal brain. -

Baz1a, Which Is the Closest Mammalian Homolog of Acf1 from Drosophila
Copyright © 2012 by James A. Dowdle All rights reserved DEDICATION This work is dedicated to my loving parents and grandparents who have never stood in the way of letting me be me and for selflessly ensuring and continuing to ensure that I have every opportunity to succeed. Also, in loving memory of my Grams who made the best damn Michigan the world has ever known. A fellow scientist once reminded me that: “Science is boring, boring, boring, boring (breath), boring, boring, exciting! and that is why we keep doing what we do.” iii ABSTRACT Acf1 was isolated almost 15 years ago from Drosophila embryo extracts as a subunit of several complexes that possess chromatin assembly activity. Acf1 binds to the ATPase Iswi (SNF2H in mammals) to form the ACF and CHRAC chromatin remodeling complexes, which can slide nucleosomes and assemble arrays of regularly spaced nucleosomes in vitro. Evidence from Drosophila and from human and mouse cell culture studies implicates Acf1 in a wide range of cellular processes including transcriptional repression, heterochromatin formation and replication, and DNA damage checkpoints and repair. However, despite these studies, little is known about the in vivo function of Acf1 in mammals. This thesis is focused on elucidating the role of mouse Baz1a, which is the closest mammalian homolog of Acf1 from Drosophila. The study began by characterizing the expression of Baz1a, which revealed high expression in the testis, and continued with analysis of unique splice variants and the subcellular localization of the protein product in this organ. Next, Baz1a-deficient mice were generated to investigate the role of this factor during development. -

The Drosophila Homologue of Vertebrate Myogenic-Determination
Proc. Natl. Acad. Sci. USA Vol. 88, pp. 3782-3786, May 1991 Developmental Biology The Drosophila homologue of vertebrate myogenic-determination genes encodes a transiently expressed nuclear protein marking primary myogenic cells (insect myogenesis/helix-loop-helix/invertebrate MyoD) BRUCE M. PATERSON*t, UWE WALLDORFt, JUANITA ELDRIDGE*, ANDREAS DUBENDORFERt, MANFRED FRASCH§, AND WALTER J. GEHRINGt *Laboratory of Biochemistry, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892; tDepartment of Cell Biology, Biozentrum, University of Basel, Klingelbergstrasse 70, CH-4056 Basel, Switzerland; tZoology Institute, University of Zurich, Winterthurerstrasse 190, CH-8057 Zurich, Switzerland; and §Max Planck Institute for Developmental Biology, Abteilung III Genetik, D-7400 Tubingen, Federal Republic of Germany Contributed by Walter J. Gehring, January 22, 1991 10 30 50 ABSTRACT We have isolated a cDNA clone, called Dmyd GGAAAAATCCCGAAAGTGAAACTATAACAAATTAAACTAAATAGAAACCACAGCCTAAAA 70 90 110 that encodes a CTTGTGTGTAACCATAACTCATAAATTTGTGTTAATGACCTAGCATACCTAGAAAAGGAG for Drosophila myogenic-determination gene, 130 150 170 CTTAAACCACGTTAAAAATGGTAATAATCAACTGACTAAATTAACTGTGCCATCGTAATT protein with structural and functional characteristics similar to 190 210 230 AGTATCAGCAATTAAATTCAGCAACTTGTTGAACTACCAAAATTGTTTTGTAAAATATAA the members of the vertebrate MyoD family. Dmyd clone 250 270 290 CAAAATCGTAGTGAAGTGAAAAATGACCAAGTATAkATAGTGGCAGCAGTGAAATGCCTGC encodes a polypeptide of 332 amino acids with 82% identity to M TK Y N S G S S E M P A 310 330 350 MyoD in the 41 amino acids of the putative helix-loop-helix GGCTCAAACCATCAAGCAGGAGTACCACAATGGCTATGGTCAGCCGACACATCCTGGATA A Q T I K 0 E Y H N G YG Q P T H P G Y region and 100% identity in the 13 amino acids of the basic 37S0 390 4 10 CGGATTTAGCGCCTATAGCCAACAGAATCCGATAGCCCATCCCGGCCAGAATCCACACCA domain proposed to contain the essential recognition code for G FS A Y S QQ N P I A H P G Q N P H Q 430 450 470 muscle-specific gene activation. -

The Drosophila Homologue of Vertebrate Myogenic-Determination
Proc. Natl. Acad. Sci. USA Vol. 88, pp. 3782-3786, May 1991 Developmental Biology The Drosophila homologue of vertebrate myogenic-determination genes encodes a transiently expressed nuclear protein marking primary myogenic cells (insect myogenesis/helix-loop-helix/invertebrate MyoD) BRUCE M. PATERSON*t, UWE WALLDORFt, JUANITA ELDRIDGE*, ANDREAS DUBENDORFERt, MANFRED FRASCH§, AND WALTER J. GEHRINGt *Laboratory of Biochemistry, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892; tDepartment of Cell Biology, Biozentrum, University of Basel, Klingelbergstrasse 70, CH-4056 Basel, Switzerland; tZoology Institute, University of Zurich, Winterthurerstrasse 190, CH-8057 Zurich, Switzerland; and §Max Planck Institute for Developmental Biology, Abteilung III Genetik, D-7400 Tubingen, Federal Republic of Germany Contributed by Walter J. Gehring, January 22, 1991 10 30 50 ABSTRACT We have isolated a cDNA clone, called Dmyd GGAAAAATCCCGAAAGTGAAACTATAACAAATTAAACTAAATAGAAACCACAGCCTAAAA 70 90 110 that encodes a CTTGTGTGTAACCATAACTCATAAATTTGTGTTAATGACCTAGCATACCTAGAAAAGGAG for Drosophila myogenic-determination gene, 130 150 170 CTTAAACCACGTTAAAAATGGTAATAATCAACTGACTAAATTAACTGTGCCATCGTAATT protein with structural and functional characteristics similar to 190 210 230 AGTATCAGCAATTAAATTCAGCAACTTGTTGAACTACCAAAATTGTTTTGTAAAATATAA the members of the vertebrate MyoD family. Dmyd clone 250 270 290 CAAAATCGTAGTGAAGTGAAAAATGACCAAGTATAkATAGTGGCAGCAGTGAAATGCCTGC encodes a polypeptide of 332 amino acids with 82% identity to M TK Y N S G S S E M P A 310 330 350 MyoD in the 41 amino acids of the putative helix-loop-helix GGCTCAAACCATCAAGCAGGAGTACCACAATGGCTATGGTCAGCCGACACATCCTGGATA A Q T I K 0 E Y H N G YG Q P T H P G Y region and 100% identity in the 13 amino acids of the basic 37S0 390 4 10 CGGATTTAGCGCCTATAGCCAACAGAATCCGATAGCCCATCCCGGCCAGAATCCACACCA domain proposed to contain the essential recognition code for G FS A Y S QQ N P I A H P G Q N P H Q 430 450 470 muscle-specific gene activation.