Prohibitin, STAT3 and SH2D4A Physically and Functionally Interact
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genetic Basis of Simple and Complex Traits with Relevance to Avian Evolution
Genetic basis of simple and complex traits with relevance to avian evolution Małgorzata Anna Gazda Doctoral Program in Biodiversity, Genetics and Evolution D Faculdade de Ciências da Universidade do Porto 2019 Supervisor Miguel Jorge Pinto Carneiro, Auxiliary Researcher, CIBIO/InBIO, Laboratório Associado, Universidade do Porto Co-supervisor Ricardo Lopes, CIBIO/InBIO Leif Andersson, Uppsala University FCUP Genetic basis of avian traits Nota Previa Na elaboração desta tese, e nos termos do número 2 do Artigo 4º do Regulamento Geral dos Terceiros Ciclos de Estudos da Universidade do Porto e do Artigo 31º do D.L.74/2006, de 24 de Março, com a nova redação introduzida pelo D.L. 230/2009, de 14 de Setembro, foi efetuado o aproveitamento total de um conjunto coerente de trabalhos de investigação já publicados ou submetidos para publicação em revistas internacionais indexadas e com arbitragem científica, os quais integram alguns dos capítulos da presente tese. Tendo em conta que os referidos trabalhos foram realizados com a colaboração de outros autores, o candidato esclarece que, em todos eles, participou ativamente na sua conceção, na obtenção, análise e discussão de resultados, bem como na elaboração da sua forma publicada. Este trabalho foi apoiado pela Fundação para a Ciência e Tecnologia (FCT) através da atribuição de uma bolsa de doutoramento (PD/BD/114042/2015) no âmbito do programa doutoral em Biodiversidade, Genética e Evolução (BIODIV). 2 FCUP Genetic basis of avian traits Acknowledgements Firstly, I would like to thank to my all supervisors Miguel Carneiro, Ricardo Lopes and Leif Andersson, for the demanding task of supervising myself last four years. -

Noelia Díaz Blanco
Effects of environmental factors on the gonadal transcriptome of European sea bass (Dicentrarchus labrax), juvenile growth and sex ratios Noelia Díaz Blanco Ph.D. thesis 2014 Submitted in partial fulfillment of the requirements for the Ph.D. degree from the Universitat Pompeu Fabra (UPF). This work has been carried out at the Group of Biology of Reproduction (GBR), at the Department of Renewable Marine Resources of the Institute of Marine Sciences (ICM-CSIC). Thesis supervisor: Dr. Francesc Piferrer Professor d’Investigació Institut de Ciències del Mar (ICM-CSIC) i ii A mis padres A Xavi iii iv Acknowledgements This thesis has been made possible by the support of many people who in one way or another, many times unknowingly, gave me the strength to overcome this "long and winding road". First of all, I would like to thank my supervisor, Dr. Francesc Piferrer, for his patience, guidance and wise advice throughout all this Ph.D. experience. But above all, for the trust he placed on me almost seven years ago when he offered me the opportunity to be part of his team. Thanks also for teaching me how to question always everything, for sharing with me your enthusiasm for science and for giving me the opportunity of learning from you by participating in many projects, collaborations and scientific meetings. I am also thankful to my colleagues (former and present Group of Biology of Reproduction members) for your support and encouragement throughout this journey. To the “exGBRs”, thanks for helping me with my first steps into this world. Working as an undergrad with you Dr. -

Early Growth Response 1 Regulates Hematopoietic Support and Proliferation in Human Primary Bone Marrow Stromal Cells
Hematopoiesis SUPPLEMENTARY APPENDIX Early growth response 1 regulates hematopoietic support and proliferation in human primary bone marrow stromal cells Hongzhe Li, 1,2 Hooi-Ching Lim, 1,2 Dimitra Zacharaki, 1,2 Xiaojie Xian, 2,3 Keane J.G. Kenswil, 4 Sandro Bräunig, 1,2 Marc H.G.P. Raaijmakers, 4 Niels-Bjarne Woods, 2,3 Jenny Hansson, 1,2 and Stefan Scheding 1,2,5 1Division of Molecular Hematology, Department of Laboratory Medicine, Lund University, Lund, Sweden; 2Lund Stem Cell Center, Depart - ment of Laboratory Medicine, Lund University, Lund, Sweden; 3Division of Molecular Medicine and Gene Therapy, Department of Labora - tory Medicine, Lund University, Lund, Sweden; 4Department of Hematology, Erasmus MC Cancer Institute, Rotterdam, the Netherlands and 5Department of Hematology, Skåne University Hospital Lund, Skåne, Sweden ©2020 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2019.216648 Received: January 14, 2019. Accepted: July 19, 2019. Pre-published: August 1, 2019. Correspondence: STEFAN SCHEDING - [email protected] Li et al.: Supplemental data 1. Supplemental Materials and Methods BM-MNC isolation Bone marrow mononuclear cells (BM-MNC) from BM aspiration samples were isolated by density gradient centrifugation (LSM 1077 Lymphocyte, PAA, Pasching, Austria) either with or without prior incubation with RosetteSep Human Mesenchymal Stem Cell Enrichment Cocktail (STEMCELL Technologies, Vancouver, Canada) for lineage depletion (CD3, CD14, CD19, CD38, CD66b, glycophorin A). BM-MNCs from fetal long bones and adult hip bones were isolated as reported previously 1 by gently crushing bones (femora, tibiae, fibulae, humeri, radii and ulna) in PBS+0.5% FCS subsequent passing of the cell suspension through a 40-µm filter. -

Role and Regulation of the P53-Homolog P73 in the Transformation of Normal Human Fibroblasts
Role and regulation of the p53-homolog p73 in the transformation of normal human fibroblasts Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Bayerischen Julius-Maximilians-Universität Würzburg vorgelegt von Lars Hofmann aus Aschaffenburg Würzburg 2007 Eingereicht am Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Dr. Martin J. Müller Gutachter: Prof. Dr. Michael P. Schön Gutachter : Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am Erklärung Hiermit erkläre ich, dass ich die vorliegende Arbeit selbständig angefertigt und keine anderen als die angegebenen Hilfsmittel und Quellen verwendet habe. Diese Arbeit wurde weder in gleicher noch in ähnlicher Form in einem anderen Prüfungsverfahren vorgelegt. Ich habe früher, außer den mit dem Zulassungsgesuch urkundlichen Graden, keine weiteren akademischen Grade erworben und zu erwerben gesucht. Würzburg, Lars Hofmann Content SUMMARY ................................................................................................................ IV ZUSAMMENFASSUNG ............................................................................................. V 1. INTRODUCTION ................................................................................................. 1 1.1. Molecular basics of cancer .......................................................................................... 1 1.2. Early research on tumorigenesis ................................................................................. 3 1.3. Developing -

Based on Network Pharmacology and RNA Sequencing Techniques to Explore the Molecular Mechanism of Huatan Jiangzhuo Decoction for Treating Hyperlipidemia
Hindawi Evidence-Based Complementary and Alternative Medicine Volume 2021, Article ID 9863714, 16 pages https://doi.org/10.1155/2021/9863714 Research Article Based on Network Pharmacology and RNA Sequencing Techniques to Explore the Molecular Mechanism of Huatan Jiangzhuo Decoction for Treating Hyperlipidemia XiaowenZhou ,1 ZhenqianYan ,1 YaxinWang ,1 QiRen ,1 XiaoqiLiu ,2 GeFang ,3 Bin Wang ,4 and Xiantao Li 1 1Laboratory of TCM Syndrome Essence and Objectification, School of Basic Medical Sciences, Guangzhou University of Chinese Medicine, Guangzhou 510006, China 2Guangzhou Sagene Tech Co., Ltd., Guangzhou 510006, China 3College of Traditional Chinese Medicine, Hunan University of Chinese Medicine, Changsha 410208, China 4Shenzhen Traditional Chinese Medicine Hospital, Shenzhen 518000, China Correspondence should be addressed to Xiantao Li; [email protected] Received 22 May 2020; Revised 12 March 2021; Accepted 18 March 2021; Published 12 April 2021 Academic Editor: Jianbo Wan Copyright © 2021 Xiaowen Zhou et al. +is is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Background. Hyperlipidemia, due to the practice of unhealthy lifestyles of modern people, has been a disturbance to a large portion of population worldwide. Recently, several scholars have turned their attention to Chinese medicine (CM) to seek out a lipid-lowering approach with high efficiency and low toxicity. +is study aimed to explore the mechanism of Huatan Jiangzhuo decoction (HTJZD, a prescription of CM) in the treatment of hyperlipidemia and to determine the major regulation pathways and potential key targets involved in the treatment process. -

Supplementary Tables S1-S3
Supplementary Table S1: Real time RT-PCR primers COX-2 Forward 5’- CCACTTCAAGGGAGTCTGGA -3’ Reverse 5’- AAGGGCCCTGGTGTAGTAGG -3’ Wnt5a Forward 5’- TGAATAACCCTGTTCAGATGTCA -3’ Reverse 5’- TGTACTGCATGTGGTCCTGA -3’ Spp1 Forward 5'- GACCCATCTCAGAAGCAGAA -3' Reverse 5'- TTCGTCAGATTCATCCGAGT -3' CUGBP2 Forward 5’- ATGCAACAGCTCAACACTGC -3’ Reverse 5’- CAGCGTTGCCAGATTCTGTA -3’ Supplementary Table S2: Genes synergistically regulated by oncogenic Ras and TGF-β AU-rich probe_id Gene Name Gene Symbol element Fold change RasV12 + TGF-β RasV12 TGF-β 1368519_at serine (or cysteine) peptidase inhibitor, clade E, member 1 Serpine1 ARE 42.22 5.53 75.28 1373000_at sushi-repeat-containing protein, X-linked 2 (predicted) Srpx2 19.24 25.59 73.63 1383486_at Transcribed locus --- ARE 5.93 27.94 52.85 1367581_a_at secreted phosphoprotein 1 Spp1 2.46 19.28 49.76 1368359_a_at VGF nerve growth factor inducible Vgf 3.11 4.61 48.10 1392618_at Transcribed locus --- ARE 3.48 24.30 45.76 1398302_at prolactin-like protein F Prlpf ARE 1.39 3.29 45.23 1392264_s_at serine (or cysteine) peptidase inhibitor, clade E, member 1 Serpine1 ARE 24.92 3.67 40.09 1391022_at laminin, beta 3 Lamb3 2.13 3.31 38.15 1384605_at Transcribed locus --- 2.94 14.57 37.91 1367973_at chemokine (C-C motif) ligand 2 Ccl2 ARE 5.47 17.28 37.90 1369249_at progressive ankylosis homolog (mouse) Ank ARE 3.12 8.33 33.58 1398479_at ryanodine receptor 3 Ryr3 ARE 1.42 9.28 29.65 1371194_at tumor necrosis factor alpha induced protein 6 Tnfaip6 ARE 2.95 7.90 29.24 1386344_at Progressive ankylosis homolog (mouse) -

The Potential Genetic Network of Human Brain SARS-Cov-2 Infection
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.06.027318; this version posted April 6, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. The potential genetic network of human brain SARS-CoV-2 infection. Colline Lapina 1,2,3, Mathieu Rodic 1, Denis Peschanski 4,5, and Salma Mesmoudi 1, 3, 4, 5 1 Prematuration Program: linkAllBrains. CNRS. Paris. France 2 Graduate School in Cognitive Engineering (ENSC). Talence. France 3 Complex Systems Institute Paris île-de-France. Paris. France 4 CNRS, Paris-1-Panthéon-Sorbonne University. CESSP-UMR8209. Paris. France 5 MATRICE Equipex. Paris. France Abstract The literature reports several symptoms of SARS-CoV-2 in humans such as fever, cough, fatigue, pneumonia, and headache. Furthermore, patients infected with similar strains (SARS-CoV and MERS-CoV) suffered testis, liver, or thyroid damage. Angiotensin-converting enzyme 2 (ACE2) serves as an entry point into cells for some strains of coronavirus (SARS-CoV, MERS-CoV, SARS-CoV-2). Our hypothesis was that as ACE2 is essential to the SARS-CoV-2 virus invasion, then brain regions where ACE2 is the most expressed are more likely to be disturbed by the infection. Thus, the expression of other genes which are also over-expressed in those damaged areas could be affected. We used mRNA expression levels data of genes provided by the Allen Human Brain Atlas (ABA), and computed spatial correlations with the LinkRbrain platform. -

Research 1..8
Article pubs.acs.org/jpr Proteome Atlas of Human Chromosome 8 and Its Multiple 8p Deficiencies in Tumorigenesis of the Stomach, Colon, and Liver † ○ † ○ ‡ § ○ ∥ ○ † † † Yang Zhang, , Guoquan Yan, , Linhui Zhai, , , Shaohang Xu, , Huali Shen, Jun Yao, Feifei Wu, † # † † † ‡ § ‡ § Liqi Xie, Hailin Tang, Hongxiu Yu, Mingqi Liu, Pengyuan Yang, Ping Xu, , Chengpu Zhang, , ‡ § ‡ § ‡ § ‡ § ‡ § ∥ ⊥ ∥ Liwei Li, , Cheng Chang, , Ning Li, , Songfeng Wu, , Yunping Zhu, , Quanhui Wang, , Bo Wen, ∥ ∥ ∥ □ † ∥ ⊥ ‡ § Liang Lin, Yinzhu Wang, Guiyan Zheng, Lanping Zhou, Haojie Lu,*, Siqi Liu,*, , Fuchu He,*, , † and Fan Zhong*, † Institutes of Biomedical Sciences and Department of Chemistry, Fudan University, Shanghai 200032, China ‡ State Key Laboratory of Proteomics, Beijing Proteome Research Center, Beijing Institute of Radiation Medicine, Beijing 102206, China § National Engineering Research Center for Protein Drugs, Beijing 102206, China ∥ BGI-Shenzhen, Shenzhen 518083, China ⊥ Beijing Institute of Genomics, Chinese Academy of Sciences, Beijing 100029, China # College of Mechanical & Electronic Engineering and Automatization, National University of Defense Technology, Changsha 410073, China □ State Key Laboratory of Molecular Oncology, Cancer Institute & Hospital, Chinese Academy of Medical Sciences & Peking Union Medical College, Beijing 100021, China *S Supporting Information ABSTRACT: Chromosome 8, a medium-length euchromatic unit in humans that has an extraordinarily high mutation rate, can be detected not only in evolution but also in multiple -

Autocrine IFN Signaling Inducing Profibrotic Fibroblast Responses By
Downloaded from http://www.jimmunol.org/ by guest on September 23, 2021 Inducing is online at: average * The Journal of Immunology , 11 of which you can access for free at: 2013; 191:2956-2966; Prepublished online 16 from submission to initial decision 4 weeks from acceptance to publication August 2013; doi: 10.4049/jimmunol.1300376 http://www.jimmunol.org/content/191/6/2956 A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Autocrine IFN Signaling Feng Fang, Kohtaro Ooka, Xiaoyong Sun, Ruchi Shah, Swati Bhattacharyya, Jun Wei and John Varga J Immunol cites 49 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130037 6.DC1 This article http://www.jimmunol.org/content/191/6/2956.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 23, 2021. The Journal of Immunology A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Inducing Autocrine IFN Signaling Feng Fang,* Kohtaro Ooka,* Xiaoyong Sun,† Ruchi Shah,* Swati Bhattacharyya,* Jun Wei,* and John Varga* Activation of TLR3 by exogenous microbial ligands or endogenous injury-associated ligands leads to production of type I IFN. -

A Network Inference Approach to Understanding Musculoskeletal
A NETWORK INFERENCE APPROACH TO UNDERSTANDING MUSCULOSKELETAL DISORDERS by NIL TURAN A thesis submitted to The University of Birmingham for the degree of Doctor of Philosophy College of Life and Environmental Sciences School of Biosciences The University of Birmingham June 2013 University of Birmingham Research Archive e-theses repository This unpublished thesis/dissertation is copyright of the author and/or third parties. The intellectual property rights of the author or third parties in respect of this work are as defined by The Copyright Designs and Patents Act 1988 or as modified by any successor legislation. Any use made of information contained in this thesis/dissertation must be in accordance with that legislation and must be properly acknowledged. Further distribution or reproduction in any format is prohibited without the permission of the copyright holder. ABSTRACT Musculoskeletal disorders are among the most important health problem affecting the quality of life and contributing to a high burden on healthcare systems worldwide. Understanding the molecular mechanisms underlying these disorders is crucial for the development of efficient treatments. In this thesis, musculoskeletal disorders including muscle wasting, bone loss and cartilage deformation have been studied using systems biology approaches. Muscle wasting occurring as a systemic effect in COPD patients has been investigated with an integrative network inference approach. This work has lead to a model describing the relationship between muscle molecular and physiological response to training and systemic inflammatory mediators. This model has shown for the first time that oxygen dependent changes in the expression of epigenetic modifiers and not chronic inflammation may be causally linked to muscle dysfunction. -

SH2D4A Regulates Cell Proliferation Via the Erα/PLC-Γ/PKC Pathway
BMB reports SH2D4A regulates cell proliferation via the ERα/PLC-γ/PKC pathway Tingting Li, Wei Li, Jingyu Lu, Hong Liu, Yinghui Li & Yanyan Zhao* Department of Medical Genetics, China Medical University, Shenyang 110001, China SH2D4A, comprising a single SH2 domain, is a novel protein ing, Rual et al. identified four proteins that interacted with of the SH2 signaling protein family. We have previously dem- SH2D4A, including DBNL, HCLS1, MAGEA11, and SH3GL1 onstrated SH2D4A is expressed ubiquitously in various tissues (4). Through mass spectrometry, Ewing et al. found an inter- and is located in the cytoplasm. In this study we investigated action between PPP1CB and SH2D4A (5). Neither of them de- the function of SH2D4A in human embryonic kidney (HEK) tected the function of these interactions. Additionally, Lapinski 293 cells using interaction analysis, cell proliferation assays, et al. observed that SH2D4A was dispensable for T cell re- and kinase activity detection. SH2D4A was found to directly ceptor signal transduction (6). bind to estrogen receptor α (ERα), and prevent the recruitment Herein, we demonstrated an inhibitory effect of SH2D4A on of phospholipase C-γ (PLC-γ) to ERα. Moreover, we observed cell proliferation. We showed that this effect was due to sup- its inhibitory effects on estrogen-induced cell proliferation, in- pression of a non-genomic action of ERα via an ERα/PLC-γ/ volving the protein kinase C (PKC) signaling pathway. Toge- PKC signaling pathway. In addition, we confirmed a new rela- ther, these findings suggested that SH2D4A inhibited cell pro- tionship among SH2D4A, ERα, and PLC-γ. Our findings pro- liferation by suppression of the ERα/PLC-γ/PKC signaling path- vide insight into the biological function of SH2D4A. -
Downloaded from the Mouse Lysosome Gene Database, Mlgdb
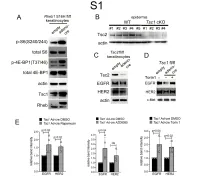
1 Supplemental Figure Legends 2 3 Supplemental Figure S1: Epidermal-specific mTORC1 gain-of-function models show 4 increased mTORC1 activation and down-regulate EGFR and HER2 protein expression in a 5 mTORC1-sensitive manner. (A) Immunoblotting of Rheb1 S16H flox/flox keratinocyte cultures 6 infected with empty or adenoviral cre recombinase for markers of mTORC1 (p-S6, p-4E-BP1) 7 activity. (B) Tsc1 cKO epidermal lysates also show decreased expression of TSC2 by 8 immunoblotting of the same experiment as in Figure 2A. (C) Immunoblotting of Tsc2 flox/flox 9 keratinocyte cultures infected with empty or adenoviral cre recombinase showing decreased EGFR 10 and HER2 protein expression. (D) Expression of EGFR and HER2 was decreased in Tsc1 cre 11 keratinocytes compared to empty controls, and up-regulated in response to Torin1 (1µM, 24 hrs), 12 by immunoblot analyses. Immunoblots are contemporaneous and parallel from the same biological 13 replicate and represent the same experiment as depicted in Figure 7B. (E) Densitometry 14 quantification of representative immunoblot experiments shown in Figures 2E and S1D (r≥3; error 15 bars represent STDEV; p-values by Student’s T-test). 16 17 18 19 20 21 22 23 Supplemental Figure S2: EGFR and HER2 transcription are unchanged with epidermal/ 24 keratinocyte Tsc1 or Rptor loss. Egfr and Her2 mRNA levels in (A) Tsc1 cKO epidermal lysates, 25 (B) Tsc1 cKO keratinocyte lysates and(C) Tsc1 cre keratinocyte lysates are minimally altered 26 compared to their respective controls. (r≥3; error bars represent STDEV; p-values by Student’s T- 27 test).