A Detailed RFLP Map of Cotton, Gossypium Hirsutum X Gossypium
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Natural Materials for the Textile Industry Alain Stout
English by Alain Stout For the Textile Industry Natural Materials for the Textile Industry Alain Stout Compiled and created by: Alain Stout in 2015 Official E-Book: 10-3-3016 Website: www.TakodaBrand.com Social Media: @TakodaBrand Location: Rotterdam, Holland Sources: www.wikipedia.com www.sensiseeds.nl Translated by: Microsoft Translator via http://www.bing.com/translator Natural Materials for the Textile Industry Alain Stout Table of Contents For Word .............................................................................................................................. 5 Textile in General ................................................................................................................. 7 Manufacture ....................................................................................................................... 8 History ................................................................................................................................ 9 Raw materials .................................................................................................................... 9 Techniques ......................................................................................................................... 9 Applications ...................................................................................................................... 10 Textile trade in Netherlands and Belgium .................................................................... 11 Textile industry ................................................................................................................... -

Gossypium Barbadense: an Approach for in Situ Conservation in Cerrado, Brazil
Journal of Agricultural Science; Vol. 8, No. 8; 2016 ISSN 1916-9752 E-ISSN 1916-9760 Published by Canadian Center of Science and Education Gossypium barbadense: An Approach for in Situ Conservation in Cerrado, Brazil Andrezza Arantes Castro1, Lúcia Vieira Hoffmann2, Thiago Henrique Lima1, Aryanny Irene Domingos Oliveira1, Rafaela Ribeiro Brito1, Letícia de Maria Oliveira Mendes1, Caio César Oliveira Pereira1, Guilherme Malafaia1 & Ivandilson Pessoa Pinto de Menezes1 1 Genetic Molecular Laboratory, Instituto Federal Goiano, Urutaí, Goiás, Brazil 2 Embrapa Algodão, Campina Grande, Paraíba, Brazil Correspondence: Ivandilson Pessoa Pinto de Menezes, School Genetic Molecular Laboratory, Instituto Federal Goiano, Urutaí, Brazil. Tel: 55-64-9279-9708. E-mail: [email protected] Received: May 27, 2016 Accepted: June 16, 2016 Online Published: July 15, 2016 doi:10.5539/jas.v8n8p59 URL:http://dx.doi.org/10.5539/jas.v8n8p59 Abstract Abandonment of planting of Gossypium barbadense has endangered its existence. The objective was to determine the characteristicof the maintenance of Gossypium barbadense in the Central-West Region of Brazil, with the aim to foster the conservation of the species. Expeditions were conducted in 2014-2015 in Southeast Goiás, where cotton collection has not been reported before. Data from previous collections in Goiás, Mato Grosso, Mato Grosso do Sul and Distrito Federal available in Albrana database were considered this study. In the Central-West Region of Brazil, 466 accesses of G. barbadense were recorded, found most frequently in backyards (91.4%), but also spontaneous plants (7.5%), farm boundary (0.8%) and commercial farming (0.2%) have also been found. The main use indicated by VDU was as medicinal plant (0.66), therefore this is the main reason for in situ preservation. -

Complete Sequence of Kenaf (Hibiscus Cannabinus)
www.nature.com/scientificreports OPEN Complete sequence of kenaf (Hibiscus cannabinus) mitochondrial genome and comparative analysis Received: 2 November 2017 Accepted: 27 July 2018 with the mitochondrial genomes of Published: xx xx xxxx other plants Xiaofang Liao1,2,3, Yanhong Zhao3, Xiangjun Kong2, Aziz Khan2, Bujin Zhou 2, Dongmei Liu4, Muhammad Haneef Kashif2, Peng Chen2, Hong Wang5 & Ruiyang Zhou2 Plant mitochondrial (mt) genomes are species specifc due to the vast of foreign DNA migration and frequent recombination of repeated sequences. Sequencing of the mt genome of kenaf (Hibiscus cannabinus) is essential for elucidating its evolutionary characteristics. In the present study, single- molecule real-time sequencing technology (SMRT) was used to sequence the complete mt genome of kenaf. Results showed that the complete kenaf mt genome was 569,915 bp long and consisted of 62 genes, including 36 protein-coding, 3 rRNA and 23 tRNA genes. Twenty-fve introns were found among nine of the 36 protein-coding genes, and fve introns were trans-spliced. A comparative analysis with other plant mt genomes showed that four syntenic gene clusters were conserved in all plant mtDNAs. Fifteen chloroplast-derived fragments were strongly associated with mt genes, including the intact sequences of the chloroplast genes psaA, ndhB and rps7. According to the plant mt genome evolution analysis, some ribosomal protein genes and succinate dehydrogenase genes were frequently lost during the evolution of angiosperms. Our data suggest that the kenaf mt genome retained evolutionarily conserved characteristics. Overall, the complete sequencing of the kenaf mt genome provides additional information and enhances our better understanding of mt genomic evolution across angiosperms. -
Polyploidy) / Ancient Genome Duplications (Paleopolyploidy
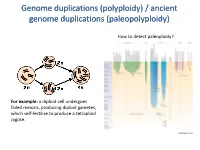
Genome duplications (polyploidy) / ancient genome duplications (paleopolyploidy) How to detect paleoploidy? For example: a diploid cell undergoes failed meiosis, producing diploid gametes, which self-fertilize to produce a tetraploid zygote. Timing of duplication by trees (phylogenetic timing) Phylogenetic timing of duplicates b Paramecium genome duplications Comparison of two scaffolds originating from a common ancestor at the recent WGD Saccharomyces cerevisiae Just before genome duplication Just after genome duplication More time after genome duplication Unaligned view (removing gaps just like in cerev has occurred) Saccharomyces cerevisiae Problem reciprocal gene loss (extreme case); how to solve? Problem reciprocal gene loss (extreme case); how to solve? Just before genome duplication Outgroup! Just after genome duplication Outgroup Just after genome duplication Outgroup More time after genome duplication Outgroup Problem (extreme case); how to solve? Outgroup Outgroup Outgroup Outgroup Outgroup Using other genomes Wong et al. 2002 PNAS Centromeres Vertebrate genome duplication Nature. 2011 Apr 10. [Epub ahead of print] Ancestral polyploidy in seed plants and angiosperms. Jiao Y, Wickett NJ, Ayyampalayam S, Chanderbali AS, Landherr L, Ralph PE, Tomsho LP, Hu Y, Liang H, Soltis PS, Soltis DE, Clifton SW, Schlarbaum SE, Schuster SC, Ma H, Leebens-Mack J, Depamphilis CW. Flowering plants Flowering MOSS Vertebrates Teleosts S. serevisiae and close relatives Paramecium Reconstructed map of genome duplications allows unprecedented mapping -

Polyploidy and the Evolutionary History of Cotton
POLYPLOIDY AND THE EVOLUTIONARY HISTORY OF COTTON Jonathan F. Wendel1 and Richard C. Cronn2 1Department of Botany, Iowa State University, Ames, Iowa 50011, USA 2Pacific Northwest Research Station, USDA Forest Service, 3200 SW Jefferson Way, Corvallis, Oregon 97331, USA I. Introduction II. Taxonomic, Cytogenetic, and Phylogenetic Framework A. Origin and Diversification of the Gossypieae, the Cotton Tribe B. Emergence and Diversification of the Genus Gossypium C. Chromosomal Evolution and the Origin of the Polyploids D. Phylogenetic Relationships and the Temporal Scale of Divergence III. Speciation Mechanisms A. A Fondness for Trans-oceanic Voyages B. A Propensity for Interspecific Gene Exchange IV. Origin of the Allopolyploids A. Time of Formation B. Parentage of the Allopolyploids V. Polyploid Evolution A. Repeated Cycles of Genome Duplication B. Chromosomal Stabilization C. Increased Recombination in Polyploid Gossypium D. A Diverse Array of Genic and Genomic Interactions E. Differential Evolution of Cohabiting Genomes VI. Ecological Consequences of Polyploidization VII. Polyploidy and Fiber VIII. Concluding Remarks References The cotton genus (Gossypium ) includes approximately 50 species distributed in arid to semi-arid regions of the tropic and subtropics. Included are four species that have independently been domesticated for their fiber, two each in Africa–Asia and the Americas. Gossypium species exhibit extraordinary morphological variation, ranging from herbaceous perennials to small trees with a diverse array of reproductive and vegetative -

A 'David and Goliath'
Current Genomics, 2002, 3, 563-576 563 Nucleolar Dominance: A ‘David and Goliath’ Chromatin Imprinting Process W. Viegas1,*, N. Neves1,2, M. Silva1, A. Caperta1,2, and L. Morais-Cecílio1 1Centro de Botânica Aplicada à Agricultura, Secção de Genética/DBEB, Instituto Superior de Agronomia, 1349-017 Lisboa, 2Departamento de Ciências Biológicas e Naturais, Universidade Lusófona de Humanidades e Tecnologias, Campo Grande 376, 1749-042 Lisboa, Portugal Abstract: Nucleolar dominance is an enigma. The puzzle of differential amphiplasty has remained unresolved since it was first recognised and described in Crepis hybrids by Navashin in 1934. Here we review the body of knowledge that has grown out of the many models that have tried to find the genetic basis for differential rRNA gene expression in hybrids, and present a new interpretation. We propose and discuss a chromatin imprinting model which re-interprets differential amphiplasty in terms of two genomes of differing size occupying a common space within the nucleus, and with heterochromatin as a key player in the scenario. Difference in size between two parental genomes induces an inherited epigenetic mark in the hybrid that allows patterns of chromatin organization to have positional effects on the neighbouring domains. This chromatin imprinting model can be also used to explain complex genomic interactions which transcend nucleolar dominance and which can account for the overall characteristics of hybrids. Gene expression in hybrids, relative to parentage, is seen as being based on the nuclear location of the sequences concerned within their genomic environment, and where the presence of particular repetitive DNA sequences are ‘sensed’, and render silent the adjacent information. -

Gene Loss and Adaptation in Saccharomyces Genomes
Genetics: Published Articles Ahead of Print, published on December 1, 2005 as 10.1534/genetics.105.048900 After the duplication: gene loss and adaptation in Saccharomyces genomes Paul F. Cliften*,1, Robert S. Fulton§, Richard K. Wilson*, §, and Mark Johnston* *Department of Genetics and §Genome Sequencing Center, Washington University School of Medicine, 660 South Euclid Ave., St. Louis, MO, 63110; 1Current address: Department of Biology, Utah State University, 5305 Old Main Hill, Logan, UT, 84322 Running head: Saccharomyces genomic duplications Key words: genomic duplication, comparative sequence analysis, Saccharomyces phylogeny, yeast Corresponding author: Mark Johnston Department of Genetics Campus Box 8232 Washington University Medical School 4566 Scott Ave. St. Louis, MO 63110 TEL: 314-362-2735 FAX: 314-362-2157 [email protected] ABSTRACT The ancient duplication of the Saccharomyces cerevisiae genome and subsequent massive loss of duplicated genes is apparent when it is compared to the genomes of related species that diverged before the duplication event. To learn more about the evolutionary effects of the duplication event, we compared the S. cerevisiae genome to other Saccharomyces genomes. We demonstrate that the whole genome duplication occurred before S. castellii diverged from S. cerevisiae. In addition to more accurately dating the duplication event, this finding allowed us to study the effects of the duplication on two separate lineages. Analyses of the duplication regions of the genomes indicate that most of the duplicated genes (approximately 85%) were lost before the speciation. Only a small amount of paralogous gene loss (4-6%) occurred after speciation. On the other hand, S. castellii appears to have lost several hundred genes that were not retained as duplicated paralogs. -

Iv Gene Duplication and the Evolution of Silenced
Gene Duplication and the Evolution of Silenced Chromatin in Yeasts by Meleah A. Hickman University Program in Genetics and Genomics Duke University Date:_______________________ Approved: ___________________________ Laura Rusche, Advisor ___________________________ Paul Magwene, Chair ___________________________ Fred Dietrich ___________________________ Joe Heitman ___________________________ Beth Sullivan Dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the University Program in Genetics and Genomics in the Graduate School of Duke University 2010 i v ABSTRACT Gene Duplication and the Evolution of Silenced Chromatin in Yeasts by Meleah A. Hickman University Program in Genetics and Genomics Duke University Date:_______________________ Approved: ___________________________ Laura Rusche, Advisor ___________________________ Paul Magwene, Chair ___________________________ Fred Dietrich ___________________________ Joe Heitman ___________________________ Beth Sullivan An abstract of a dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the University Program in Genetics and Genomics in the Graduate School of Duke University 2010 Copyright by Meleah A. Hickman 2010 Abstract In Saccharomyces cerevisiae, proper maintenance of haploid cell identity requires the SIR complex to mediate the silenced chromatin found at the cryptic mating-type loci, HML and HMR. This complex consists of Sir2, a histone deacetylase, and the histone binding proteins Sir3 and Sir4. Interestingly, both Sir2 and Sir3 have paralogs from a genome duplication that occurred after the divergence of Saccharomyces and Kluyveromyces species, approximately 100 million years ago. The histone deacetylase HST1 is the paralog of SIR2 and works with the promoter-specific SUM1 complex to repress sporulation and alpha-specific genes. ORC1 is the paralog of SIR3 and is an essential subunit of the Origin Recognition Complex and also recruits SIR proteins to the HM loci. -

Genetic Variability Studies in Gossypium Barbadense L
Electronic Journal of Plant Breeding, 1(4): 961-965 (July 2010) Research Article Genetic variability studies in Gossypium barbadense L. genotypes for seed cotton yield and its yield components K. P. M. Dhamayanathi , S. Manickam and K. Rathinavel Abstract A study was carried out during kharif 2006-07 with twenty five Gossypium barbadense L genotypes to obtain information on genetic variability, heritability and genetic advance for seed cotton yield and its yield attributes. Significant differences were observed for characters among genotypes. High genetic differences were recorded for nodes/plant, sympodia, bolls as well as fruiting points per plant, seed cotton yield, lint index indicating ample scope for genetic improvement of these characters through selection. Results also revealed high heritability coupled with high genetic advance for yield and most of the yield components as well as fibre quality traits. Sympodia/plant, fruiting point /plant, number of nodes/plant, number of bolls per plant, and lint index were positively correlated with seed cotton yield per plant and appeared to be interrelated with each other. It is suggested that these characters could be considered as selection criteria in improving the seed cotton yield of G. barbadense , L genotypes. Key words : Gossypium barbadense , genetic variability, heritability, genetic advance, lint index, selection criteria Introduction Seed cotton yield is a complex trait governed by Cotton is the most widely used vegetable fibre and several yield contributing characters such as plant also the most important raw material for the textile height, number of monopodia, number of industry, grown in tropical and subtropical regions sympodia, number of bolls, number of fruiting in more than 80 countries all over the world. -

Whole-Genome Duplications and Paleopolyploidy Whole-Genome Duplications
Whole-genome duplications and paleopolyploidy Whole-genome duplications AUTOPOLYPLOIDY ALLOPOLYPLOIDY Hegarty and HiscockHegarty 2008, 18 Current Biology Examples of allopolyploid speciation Hegarty and Hiscock 2008, Current Biology 18 Whole-genome duplications of different age time neopolyploidy mesopolyploidy paleopolyploidy time Whole-genome duplications in protozoa •Aury et al. (2006) analyzed the unicellular eukaryote Paramecium tetraurelia • most of 40,000 genes arose through at least 3 successive whole-genome duplications (WGDs) • most recent duplication most likely caused an explosion of speciation events that gave rise to the P. aurelia complex (15 sibling species) • some genes have been lost, some retained • many retained (duplicated) genes do not generate functional innovations but are important because of the gene dosage effect Whole-genome duplications in yeast • genome comparison between two yeast species, Saccharomyces cerevisiae (n=16) and Kluyveromyces waltii (n=8) • each region of K. waltii corresponding to two regions of S. cerevisiae •the S. cerevisiae genome underwent a WGD after the two yeast species diverged • in nearly every case (95%), accelerated evolution was confined to only one of the two paralogues (= one of the paralogues retained an ancestral function, the other was free to evolve more rapidly and acquired a derived function) Kellis et al. 2004, Nature 428 Whole-genome duplications in yeast Kellis et al. 2004, Nature 428 a) after divergence from K. waltii, the Saccharomyces lineage underwent a genome duplication event (2 copies of every gene and chromosome) b) duplicated genes were mutated and some lost c) two copies kept for only a small minority of duplicated genes d) the conserved order of duplicated genes (nos. -

Transposable Elements in Polyploid Evolution Final.Pdf
Transposable elements and polyploid evolution in animals Fernando Rodriguez and Irina R. Arkhipova Address Josephine Bay Paul Center for Comparative Molecular Biology and Evolution, Marine Biological Laboratory, Woods Hole, MA 02543, USA Corresponding author: Arkhipova, Irina R. ([email protected]) Polyploidy in animals is much less common than in plants, where it is thought to be pervasive in all higher plant lineages. Recent studies have highlighted the impact of polyploidization and the associated process of diploidy restoration on the evolution and speciation of selected taxonomic groups in the animal kingdom: from vertebrates represented by salmonid fishes and African clawed frogs to invertebrates represented by parasitic root-knot nematodes and bdelloid rotifers. In this review, we focus on the unique and diverse roles that transposable elements may play in these processes, from marking and diversifying subgenome-specific chromosome sets prior to hybridization, to influencing genome restructuring during rediploidization, to affecting subgenome-specific regulatory evolution, and occasionally providing opportunities for domestication and gene amplification to restore and improve functionality. There is still much to be learned from the future comparative genomic studies of chromosome-sized and haplotype-aware assemblies, and from post-genomic studies elucidating genetic and epigenetic regulatory phenomena across short and long evolutionary distances in the metazoan tree of life. Glossary Polyploidy: having more than the usual diploid set of homologous chromosomes Allopolyploidy: a form of polyploidy that results from interspecific hybridization Autopolyploidy: a form of polyploidy that results from chromosome redoubling in the same species Paleopolyploidy: ancient polyploidy Neopolyploidy: recent polyploidy Homeologs: pairs of genes originated by speciation and brought back together by allopolyploidization Ohnologs: same as homeologs, named after S. -

The Chromosome-Based Lavender Genome Provides New Insights Into
Li et al. Horticulture Research (2021) 8:53 Horticulture Research https://doi.org/10.1038/s41438-021-00490-6 www.nature.com/hortres ARTICLE Open Access The chromosome-based lavender genome provides new insights into Lamiaceae evolution and terpenoid biosynthesis Jingrui Li1,2, Yiming Wang 3,YanmeiDong1,2, Wenying Zhang1,2,DiWang1, Hongtong Bai1,KuiLi3,HuiLi1 and Lei Shi1 Abstract The aromatic shrub Lavandula angustifolia produces various volatile terpenoids that serve as resources for essential oils and function in plant-insect communication. To better understand the genetic basis of the terpenoid diversity in lavender, we present a high-quality reference genome for the Chinese lavender cultivar ‘Jingxun 2’ using PacBio and Hi-C technologies to anchor the 894.50 Mb genome assembly into 27 pseudochromosomes. In addition to the γ triplication event, lavender underwent two rounds of whole-genome duplication (WGD) during the Eocene–Oligocene (29.6 MYA) and Miocene–Pliocene (6.9 MYA) transitions. As a result of tandem duplications and lineage-specific WGDs, gene families related to terpenoid biosynthesis in lavender are substantially expanded compared to those of five other species in Lamiaceae. Many terpenoid biosynthesis transcripts are abundant in glandular trichomes. We further integrated the contents of ecologically functional terpenoids and coexpressed terpenoid biosynthetic genes to construct terpenoid-gene networks. Typical gene clusters, including TPS-TPS, TPS- CYP450, and TPS-BAHD, linked with compounds that primarily function as attractants or repellents, were identified by fl 1234567890():,; 1234567890():,; 1234567890():,; 1234567890():,; their similar patterns of change during ower development or in response to methyl jasmonate. Comprehensive analysis of the genetic basis of the production of volatiles in lavender could serve as a foundation for future research into lavender evolution, phytochemistry, and ecology.