Non-Homologous Isofunctional Enzymes: a Systematic Analysis Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Aryl Hydrocarbon Receptor: Structural Analysis and Activation Mechanisms
The Aryl Hydrocarbon Receptor: Structural Analysis and Activation Mechanisms This thesis is submitted in fulfilment of the requirements for the degree of Doctor of Philosophy in the School of Molecular and Biomedical Sciences (Biochemistry), The University of Adelaide, Australia Fiona Whelan, B.Sc. (Hons) 2009 2 Table of Contents THESIS SUMMARY................................................................................. 6 DECLARATION....................................................................................... 7 PUBLICATIONS ARISING FROM THIS THESIS.................................... 8 ACKNOWLEDGEMENTS...................................................................... 10 ABBREVIATIONS ................................................................................. 12 CHAPTER 1: INTRODUCTION ............................................................. 17 1.1 BHLH.PAS PROTEINS ............................................................................................17 1.1.1 General background..................................................................................17 1.1.2 bHLH.PAS Class I Proteins.........................................................................18 1.2 THE ARYL HYDROCARBON RECEPTOR......................................................................19 1.2.1 Domain Structure and Ligand Activation ..............................................19 1.2.2 AhR Expression and Developmental Activity .......................................21 1.2.3 Mouse AhR Knockout Phenotype ...........................................................23 -

An Atpase Domain Common to Prokaryotic Cell Cycle Proteins
Proc. Natl. Acad. Sci. USA Vol. 89, pp. 7290-7294, August 1992 Biochemistry An ATPase domain common to prokaryotic cell cycle proteins, sugar kinases, actin, and hsp7O heat shock proteins (structural comparison/property pattern/remote homology) PEER BORK, CHRIS SANDER, AND ALFONSO VALENCIA European Molecular Biology Laboratory, D-6900 Heidelberg, Federal Republic of Germany Communicated by Russell F. Doolittle, March 6, 1992 ABSTRACT The functionally diverse actin, hexokinase, and hsp7O protein families have in common an ATPase domain of known three-dimensional structure. Optimal superposition ofthe three structures and alignment ofmany sequences in each of the three families has revealed a set of common conserved residues, distributed in five sequence motifs, which are in- volved in ATP binding and in a putative interdomain hinge. From the multiple sequence aliment in these motifs a pattern of amino acid properties required at each position is defined. The discriminatory power of the pattern is in part due to the use of several known three-dimensional structures and many sequences and in part to the "property" method ofgeneralizing from observed amino acid frequencies to amino acid fitness at each sequence position. A sequence data base search with the pattern significantly matches sugar kinases, such as fuco-, glucono-, xylulo-, ribulo-, and glycerokinase, as well as the prokaryotic cell cycle proteins MreB, FtsA, and StbA. These are predicted to have subdomains with the same tertiary structure as the ATPase subdomains Ia and Ha of hexokinase, actin, and Hsc7O, a very similar ATP binding pocket, and the capacity for interdomain hinge motion accompanying func- tional state changes. -

Biosynthesis in Vitro of Bacillamide Intermediate-Heterocyclic Alacysthiazole by Heterologous Expression of Nonribosomal Peptide Synthetase (NRPS) T
Journal of Biotechnology 292 (2019) 5–11 Contents lists available at ScienceDirect Journal of Biotechnology journal homepage: www.elsevier.com/locate/jbiotec Biosynthesis in vitro of bacillamide intermediate-heterocyclic AlaCysthiazole by heterologous expression of nonribosomal peptide synthetase (NRPS) T Fengli Zhang, Nayila Mulati, Yukun Wang, Yingxin Li, Sanqiang Gong, Loganathan Karthik, ⁎ Wei Sun, Zhiyong Li Marine Biotechnology Laboratory, State Key Laboratory of Microbial Metabolism and School of Life Sciences & Biotechnology, Shanghai Jiao Tong University, Shanghai, China ARTICLE INFO ABSTRACT Keywords: Bacillamide C, a potential natural antialgae active compound, is produced by Bacillus atrophaeus C89 derived Bacillus atrophaeus from marine sponge Dysidea avara. A nonribosomal peptide synthetase (NRPS) cluster is hypothesized to be Bacillamides involved in the biosynthesis of bacillamide C. The NRPS with a domain string of A1-PCP1-Cy-A2-PCP2-C can be Heterologous expression divided into three functional modules. After heterologous expression and purification of module A1-PCP1 and Nonribosomal peptide synthetase (NRPS) module Cy-A2-PCP2, their catalytic activities were biochemically proven in vitro by the reaction with the apo- Thiazole PCP domain transformed to the holo-PCP domain through a phosphopantetheinyl transferase, ATP, and substrate amino acids. Five– membered heterocyclic AlaCysthiazole with molecular weight of 172.0389 was detected. This proved the formation of the heterocyclic dipeptide AlaCysthiazole, which is considered to be a building block for the biosynthesis of bacillamide. This study provides a basis for further biosynthesis of bacillamides. 1. Introduction et al., 2017). Even though the biosynthesis of bacillamide C was opti- mized, the yield was very low (Jin et al., 2011; Yu et al., 2015). -

Guanylate Kinase (Ec 2.7.4.8)
Enzymatic Assay of GUANYLATE KINASE (EC 2.7.4.8) PRINCIPLE: Guanylate Kinase GMP + ATP > GDP + ADP Pyruvate Kinase ADP + PEP > ATP + Pyruvate Pyruvate Kinase GDP + PEP > GTP + Pyruvate Lactic Dehydrogenase 2 Pyruvate + 2 ß-NADH > 2 Lactate + 2 ß-NAD Abbreviations used: GMP = Guanosine 5'-Monophosphate ATP = Adenosine 5'-Triphosphate GDP = Guanosine 5'-Diphosphate ADP = Adenosine 5'-Diphosphate PEP = Phospho(enol)phosphate ß-NADH = ß-Nicotinamide Adenine Dinucleotide, Reduced Form ß-NAD = ß-Nicotinamide Adenine Dinucleotide, Oxidized Form CONDITIONS: T = 30°C, pH = 7.5, A340nm, Light path = 1 cm METHOD: Continuous Spectrophotometric Rate Determination REAGENTS: A. 200 mM Tris HCl Buffer, pH 7.5 at 30°C (Prepare 50 ml in deionized water using Trizma Base, Sigma Prod. No. T-1503. Adjust to pH 7.5 at 30°C with 1 M HCl.) B. 1 M Potassium Chloride Solution (KCl) (Prepare 10 ml in deionized water using Potassium Chloride, Sigma Prod. No. P-4504.) C. 60 mM Magnesium Sulfate Solution (MgSO4) (Prepare 20 ml in deionized water using Magnesium Sulfate, Heptahydrate, Sigma Prod. No. M-1880.) D. 40 mM Phospho(enol)pyruvate Solution (PEP) (Prepare 50 ml in deionized water using Phospho(enol)Pyruvate, Trisodium Salt, Hydrate, Sigma Prod. No. P-7002. PREPARE FRESH.) Revised: 03/10/94 Page 1 of 4 Enzymatic Assay of GUANYLATE KINASE (EC 2.7.4.8) REAGENTS: (continued) E. 100 mM Ethylenediaminetetraacetic Acid Solution (EDTA) (Prepare 10 ml in deionized water using Ethylenediaminetetraacetic Acid, Tetrasodium Salt, Hydrate, Sigma Stock No. ED4SS.) F. 3.8 mM ß-Nicotinamide Adenine Dinucleotide, Reduced Form (ß-NADH) (Prepare 2 ml in deionized water using ß-Nicotinamide Adenine Dinucleotide, Reduced Form, Dipotassium Salt, Sigma Prod. -

WO 2013/180584 Al 5 December 2013 (05.12.2013) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2013/180584 Al 5 December 2013 (05.12.2013) P O P C T (51) International Patent Classification: AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, C12N 1/21 (2006.01) C12N 15/74 (2006.01) BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, C12N 15/52 (2006.01) C12P 5/02 (2006.01) DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, C12N 15/63 (2006.01) HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KN, KP, KR, KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, ME, (21) International Application Number: MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, PCT/NZ20 13/000095 OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SC, (22) International Filing Date: SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, 4 June 2013 (04.06.2013) TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (25) Filing Language: English (84) Designated States (unless otherwise indicated, for every kind of regional protection available): ARIPO (BW, GH, (26) Publication Language: English GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, SZ, TZ, (30) Priority Data: UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, TJ, 61/654,412 1 June 2012 (01 .06.2012) US TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, DK, EE, ES, FI, FR, GB, GR, HR, HU, IE, IS, IT, LT, LU, LV, (71) Applicant: LANZATECH NEW ZEALAND LIMITED MC, MK, MT, NL, NO, PL, PT, RO, RS, SE, SI, SK, SM, [NZ/NZ]; 24 Balfour Road, Parnell, Auckland, 1052 (NZ). -

Increased Whitening Efficacy and Reduced Cytotoxicity Are Achieved by the Chemical Activation of a Highly Concentrated Hydrogen Peroxide Bleaching Gel
Original Article http://dx.doi.org/10.1590/1678-7757-2018-0453 Increased whitening efficacy and reduced cytotoxicity are achieved by the chemical activation of a highly concentrated hydrogen peroxide bleaching gel Abstract Diana Gabriela SOARES1 Objective: This study was designed for the chemical activation of a 35% hydrogen peroxide (H O ) bleaching gel to increase its whitening effectiveness Natália MARCOMINI² 2 2 and reduce its toxicity. Methodology: First, the bleaching gel - associated or Carla Caroline de Oliveira DUQUE³ not with ferrous sulfate (FS), manganese chloride (MC), peroxidase (PR), Ester Alves Ferreira BORDINI³ or catalase (CT) - was applied (3x 15 min) to enamel/dentin discs adapted Uxua Ortecho ZUTA³ to artificial pulp chambers. Then, odontoblast-like MDPC-23 cells were Fernanda Gonçalves BASSO⁴ exposed for 1 h to the extracts (culture medium + components released Josimeri HEBLING⁵ from the product), for the assessment of viability (MTT assay) and oxidative D Carlos Alberto de Souza COSTA⁴ stress (H2DCFDA). Residual H2O2 and bleaching effectiveness ( E) were also evaluated. Data were analyzed with one-way ANOVA complemented with Tukey’s test (n=8. p<0.05). Results: All chemically activated groups minimized MDPC-23 oxidative stress generation; however, significantly higher cell viability was detected for MC, PR, and CT than for plain 35% H2O2 gel. Nevertheless, FS, MC, PR, and CT reduced the amount of residual H2O2 and increased bleaching effectiveness. Conclusion: Chemical activation of 35% H2O2 gel with MC, PR, and CT minimized residual H2O2 and pulp cell toxicity; but PR duplicated the whitening potential of the bleaching gel after a single 45-minute session. -

(12) Patent Application Publication (10) Pub. No.: US 2014/0155567 A1 Burk Et Al
US 2014O155567A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2014/0155567 A1 Burk et al. (43) Pub. Date: Jun. 5, 2014 (54) MICROORGANISMS AND METHODS FOR (60) Provisional application No. 61/331,812, filed on May THE BIOSYNTHESIS OF BUTADENE 5, 2010. (71) Applicant: Genomatica, Inc., San Diego, CA (US) Publication Classification (72) Inventors: Mark J. Burk, San Diego, CA (US); (51) Int. Cl. Anthony P. Burgard, Bellefonte, PA CI2P 5/02 (2006.01) (US); Jun Sun, San Diego, CA (US); CSF 36/06 (2006.01) Robin E. Osterhout, San Diego, CA CD7C II/6 (2006.01) (US); Priti Pharkya, San Diego, CA (52) U.S. Cl. (US) CPC ................. CI2P5/026 (2013.01); C07C II/I6 (2013.01); C08F 136/06 (2013.01) (73) Assignee: Genomatica, Inc., San Diego, CA (US) USPC ... 526/335; 435/252.3:435/167; 435/254.2: (21) Appl. No.: 14/059,131 435/254.11: 435/252.33: 435/254.21:585/16 (22) Filed: Oct. 21, 2013 (57) ABSTRACT O O The invention provides non-naturally occurring microbial Related U.S. Application Data organisms having a butadiene pathway. The invention addi (63) Continuation of application No. 13/101,046, filed on tionally provides methods of using Such organisms to produce May 4, 2011, now Pat. No. 8,580,543. butadiene. Patent Application Publication Jun. 5, 2014 Sheet 1 of 4 US 2014/O155567 A1 ?ueudos!SMS |?un61– Patent Application Publication Jun. 5, 2014 Sheet 2 of 4 US 2014/O155567 A1 VOJ OO O Z?un61– Patent Application Publication US 2014/O155567 A1 {}}} Hººso Patent Application Publication Jun. -

Chloride Peroxidase Cat
chloride peroxidase Cat. No. EXWM-0491 Lot. No. (See product label) Introduction Description Brings about the chlorination of a range of organic molecules, forming stable C-Cl bonds. Also oxidizes bromide and iodide. Enzymes of this type are either heme-thiolate proteins, or contain vanadate. A secreted enzyme produced by the ascomycetous fungus Caldariomyces fumago (Leptoxyphium fumago) is an example of the heme-thiolate type. It catalyses the production of hypochlorous acid by transferring one oxygen atom from H2O2 to chloride. At a separate site it catalyses the chlorination of activated aliphatic and aromatic substrates, via HClO and derived chlorine species. In the absence of halides, it shows peroxidase (e.g. phenol oxidation) and peroxygenase activities. The latter inserts oxygen from H2O2 into, for example, styrene (side chain epoxidation) and toluene (benzylic hydroxylation), however, these activities are less pronounced than its activity with halides. Has little activity with non-activated substrates such as aromatic rings, ethers or saturated alkanes. The chlorinating peroxidase produced by ascomycetous fungi (e.g. Curvularia inaequalis) is an example of a vanadium chloroperoxidase, and is related to bromide peroxidase (EC 1.11.1.18). It contains vanadate and oxidizes chloride, bromide and iodide into hypohalous acids. In the absence of halides, it peroxygenates organic sulfides and oxidizes ABTS [2,2'-azinobis(3-ethylbenzthiazoline-6-sulfonic acid)] but no phenols. Synonyms chloroperoxidase; CPO; vanadium haloperoxidase Product Information Form Liquid or lyophilized powder EC Number EC 1.11.1.10 CAS No. 9055-20-3 Reaction RH + chloride + H2O2 = RCl + 2 H2O Notes This item requires custom production and lead time is between 5-9 weeks. -

The Mechanism of GHMP Kinases by QM/MM Studies
The Role of Arg228 in the phosphorylation of galactokinase: The mechanism of GHMP kinases by QM/MM studies Huang, M., Li, X., Zou, J-W., & Timson, D. J. (2013). The Role of Arg228 in the phosphorylation of galactokinase: The mechanism of GHMP kinases by QM/MM studies. Biochemistry, 52(28), 4858. https://doi.org/10.1021/bi400228e Published in: Biochemistry Document Version: Early version, also known as pre-print Queen's University Belfast - Research Portal: Link to publication record in Queen's University Belfast Research Portal General rights Copyright for the publications made accessible via the Queen's University Belfast Research Portal is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The Research Portal is Queen's institutional repository that provides access to Queen's research output. Every effort has been made to ensure that content in the Research Portal does not infringe any person's rights, or applicable UK laws. If you discover content in the Research Portal that you believe breaches copyright or violates any law, please contact [email protected]. Download date:01. Oct. 2021 Article pubs.acs.org/biochemistry Role of Arg228 in the Phosphorylation of Galactokinase: The Mechanism of GHMP Kinases by Quantum Mechanics/Molecular Mechanics Studies † † ‡ § Meilan Huang,*, Xiaozhou Li, Jian-Wei Zou, and David J. Timson † School of Chemistry and Chemical Engineering, Queen’s University Belfast, David Keir Building, Stranmillis Road, Belfast BT9 5AG, U.K. -

The Role of the Salvage Pathway in Nucleotide Sugar Biosynthesis
THE ROLE OF THE SALVAGE PATHWAY IN NUCLEOTIDE SUGAR BIOSYNTHESIS: IDENTIFICATION OF SUGAR KINASES AND NDP-SUGAR PYROPHOSPHORYLASES by TING YANG (Under the Direction of Maor Bar-Peled) ABSTRACT The synthesis of polysaccharides, glycoproteins, glycolipids, glycosylated secondary metabolites and hormones requires a large number of glycosyltransferases and a constant supply of nucleotide sugars. In plants, photosynthesis and the NDP-sugar inter-conversion pathway are the major entry points to form NDP-sugars. In addition to these pathways is the salvage pathway, a less understood metabolism that provides the flux of NDP-sugars. This latter pathway involves the hydrolysis of glycans to free sugars, sugar transport, sugar phosphorylation and nucleotidylation. The balance between glycan synthesis and recycling as well as its regulation at various plant developmental stages remains elusive as many of the molecular components are unknown. To understand how the salvage pathway contributes to the sugar flux and cell wall biosynthesis, my research focused on the functional identification of salvage pathway sugar kinases and NDP-sugar pyrophosphorylases. This research led to the first identification and enzymatic characterization of galacturonic acid kinase (GalA kinase), galactokinase (GalK), a broad UDP-sugar pyrophosphorylase (sloppy), two promiscuous UDP-GlcNAc pyrophosphorylases (GlcNAc-1-P uridylyltransferases), as well as UDP-sugar pyrophosphorylase paralogs from Trypanosoma cruzi and Leishmania major. To evaluate the salvage pathway in plant biology, we further investigated a sugar kinase mutant: galacturonic acid kinase mutant (galak) and determined if and how galak KO mutant affects the synthesis of glycans in Arabidopsis. Feeding galacturonic acid to the seedlings exhibited a 40-fold accumulation of free GalA in galak mutant, while the wild type (WT) plant readily metabolizes the fed-sugar. -

Recent Advances in Trypanosomatid Research: Genome Organization, Expression, Metabolism, Taxonomy and Evolution
Parasitology Recent advances in trypanosomatid research: genome organization, expression, metabolism, cambridge.org/par taxonomy and evolution 1 2 3,4 5,6 Review Dmitri A. Maslov , Fred R. Opperdoes , Alexei Y. Kostygov , Hassan Hashimi , Julius Lukeš5,6 and Vyacheslav Yurchenko3,5,7 Cite this article: Maslov DA, Opperdoes FR, Kostygov AY, Hashimi H, Lukeš J, Yurchenko V 1Department of Molecular, Cell and Systems Biology, University of California – Riverside, Riverside, California, USA; (2018). Recent advances in trypanosomatid 2de Duve Institute, Université Catholique de Louvain, Brussels, Belgium; 3Life Science Research Centre, Faculty of research: genome organization, expression, 4 metabolism, taxonomy and evolution. Science, University of Ostrava, Ostrava, Czech Republic; Zoological Institute of the Russian Academy of Sciences, 5 Parasitology 1–27. https://doi.org/10.1017/ St. Petersburg, Russia; Biology Centre, Institute of Parasitology, Czech Academy of Sciences, České Budejovice 6 S0031182018000951 (Budweis), Czech Republic; University of South Bohemia, Faculty of Sciences, České Budejovice (Budweis), Czech Republic and 7Martsinovsky Institute of Medical Parasitology, Tropical and Vector Borne Diseases, Sechenov Received: 30 January 2018 University, Moscow, Russia Revised: 23 April 2018 Accepted: 23 April 2018 Abstract Key words: Unicellular flagellates of the family Trypanosomatidae are obligatory parasites of inverte- Gene exchange; kinetoplast; metabolism; molecular and cell biology; taxonomy; brates, vertebrates and plants. Dixenous species are aetiological agents of a number of diseases trypanosomatidae in humans, domestic animals and plants. Their monoxenous relatives are restricted to insects. Because of the high biological diversity, adaptability to dramatically different environmental Author for correspondence: conditions, and omnipresence, these protists have major impact on all biotic communities Vyacheslav Yurchenko, E-mail: vyacheslav. -
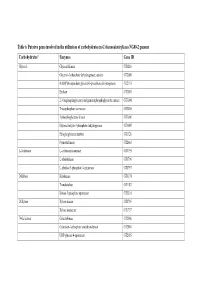
Table 6. Putative Genes Involved in the Utilization of Carbohydrates in G
Table 6. Putative genes involved in the utilization of carbohydrates in G. thermodenitrificans NG80-2 genome Carbohydrates* Enzymes Gene ID Glycerol Glycerol Kinase GT1216 Glycerol-3-phosphate dehydrogenase, aerobic GT2089 NAD(P)H-dependent glycerol-3-phosphate dehydrogenase GT2153 Enolase GT3003 2,3-bisphosphoglycerate-independentphosphoglycerate mutase GT3004 Triosephosphate isomerase GT3005 3-phosphoglycerate kinase GT3006 Glyceraldehyde-3-phosphate dehydrogenase GT3007 Phosphoglycerate mutase GT1326 Pyruvate kinase GT2663 L-Arabinose L-arabinose isomerase GT1795 L-ribulokinase GT1796 L-ribulose 5-phosphate 4-epimerase GT1797 D-Ribose Ribokinase GT3174 Transketolase GT1187 Ribose 5-phosphate epimerase GT3316 D-Xylose Xylose kinase GT1756 Xylose isomerase GT1757 D-Galactose Galactokinase GT2086 Galactose-1-phosphate uridyltransferase GT2084 UDP-glucose 4-epimerase GT2085 Carbohydrates* Enzymes Gene ID D-Fructose 1-phosphofructokinase GT1727 Fructose-1,6-bisphosphate aldolase GT1805 Fructose-1,6-bisphosphate aldolase type II GT3331 Triosephosphate isomerase GT3005 D-Mannose Mannnose-6 phospate isomelase GT3398 6-phospho-1-fructokinase GT2664 D-Mannitol Mannitol-1-phosphate dehydrogenase GT1844 N-Acetylglucosamine N-acetylglucosamine-6-phosphate deacetylase GT2205 N-acetylglucosamine-6-phosphate isomerase GT2204 D-Maltose Alpha-1,4-glucosidase GT0528, GT1643 Sucrose Sucrose phosphorylase GT3215 D-Trehalose Alpha-glucosidase GT1643 Glucose kinase GT2381 Inositol Myo-inositol catabolism protein iolC;5-dehydro-2- GT1807 deoxygluconokinase