The Photoreceptor Program in Group 3 Medulloblastoma: Role of the Tfs NRL and CRX
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Core Transcriptional Regulatory Circuitries in Cancer
Oncogene (2020) 39:6633–6646 https://doi.org/10.1038/s41388-020-01459-w REVIEW ARTICLE Core transcriptional regulatory circuitries in cancer 1 1,2,3 1 2 1,4,5 Ye Chen ● Liang Xu ● Ruby Yu-Tong Lin ● Markus Müschen ● H. Phillip Koeffler Received: 14 June 2020 / Revised: 30 August 2020 / Accepted: 4 September 2020 / Published online: 17 September 2020 © The Author(s) 2020. This article is published with open access Abstract Transcription factors (TFs) coordinate the on-and-off states of gene expression typically in a combinatorial fashion. Studies from embryonic stem cells and other cell types have revealed that a clique of self-regulated core TFs control cell identity and cell state. These core TFs form interconnected feed-forward transcriptional loops to establish and reinforce the cell-type- specific gene-expression program; the ensemble of core TFs and their regulatory loops constitutes core transcriptional regulatory circuitry (CRC). Here, we summarize recent progress in computational reconstitution and biologic exploration of CRCs across various human malignancies, and consolidate the strategy and methodology for CRC discovery. We also discuss the genetic basis and therapeutic vulnerability of CRC, and highlight new frontiers and future efforts for the study of CRC in cancer. Knowledge of CRC in cancer is fundamental to understanding cancer-specific transcriptional addiction, and should provide important insight to both pathobiology and therapeutics. 1234567890();,: 1234567890();,: Introduction genes. Till now, one critical goal in biology remains to understand the composition and hierarchy of transcriptional Transcriptional regulation is one of the fundamental mole- regulatory network in each specified cell type/lineage. -

The Title of the Dissertation
UNIVERSITY OF CALIFORNIA SAN DIEGO Novel network-based integrated analyses of multi-omics data reveal new insights into CD8+ T cell differentiation and mouse embryogenesis A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Bioinformatics and Systems Biology by Kai Zhang Committee in charge: Professor Wei Wang, Chair Professor Pavel Arkadjevich Pevzner, Co-Chair Professor Vineet Bafna Professor Cornelis Murre Professor Bing Ren 2018 Copyright Kai Zhang, 2018 All rights reserved. The dissertation of Kai Zhang is approved, and it is accept- able in quality and form for publication on microfilm and electronically: Co-Chair Chair University of California San Diego 2018 iii EPIGRAPH The only true wisdom is in knowing you know nothing. —Socrates iv TABLE OF CONTENTS Signature Page ....................................... iii Epigraph ........................................... iv Table of Contents ...................................... v List of Figures ........................................ viii List of Tables ........................................ ix Acknowledgements ..................................... x Vita ............................................. xi Abstract of the Dissertation ................................. xii Chapter 1 General introduction ............................ 1 1.1 The applications of graph theory in bioinformatics ......... 1 1.2 Leveraging graphs to conduct integrated analyses .......... 4 1.3 References .............................. 6 Chapter 2 Systematic -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplemental Materials ZNF281 Enhances Cardiac Reprogramming
Supplemental Materials ZNF281 enhances cardiac reprogramming by modulating cardiac and inflammatory gene expression Huanyu Zhou, Maria Gabriela Morales, Hisayuki Hashimoto, Matthew E. Dickson, Kunhua Song, Wenduo Ye, Min S. Kim, Hanspeter Niederstrasser, Zhaoning Wang, Beibei Chen, Bruce A. Posner, Rhonda Bassel-Duby and Eric N. Olson Supplemental Table 1; related to Figure 1. Supplemental Table 2; related to Figure 1. Supplemental Table 3; related to the “quantitative mRNA measurement” in Materials and Methods section. Supplemental Table 4; related to the “ChIP-seq, gene ontology and pathway analysis” and “RNA-seq” and gene ontology analysis” in Materials and Methods section. Supplemental Figure S1; related to Figure 1. Supplemental Figure S2; related to Figure 2. Supplemental Figure S3; related to Figure 3. Supplemental Figure S4; related to Figure 4. Supplemental Figure S5; related to Figure 6. Supplemental Table S1. Genes included in human retroviral ORF cDNA library. Gene Gene Gene Gene Gene Gene Gene Gene Symbol Symbol Symbol Symbol Symbol Symbol Symbol Symbol AATF BMP8A CEBPE CTNNB1 ESR2 GDF3 HOXA5 IL17D ADIPOQ BRPF1 CEBPG CUX1 ESRRA GDF6 HOXA6 IL17F ADNP BRPF3 CERS1 CX3CL1 ETS1 GIN1 HOXA7 IL18 AEBP1 BUD31 CERS2 CXCL10 ETS2 GLIS3 HOXB1 IL19 AFF4 C17ORF77 CERS4 CXCL11 ETV3 GMEB1 HOXB13 IL1A AHR C1QTNF4 CFL2 CXCL12 ETV7 GPBP1 HOXB5 IL1B AIMP1 C21ORF66 CHIA CXCL13 FAM3B GPER HOXB6 IL1F3 ALS2CR8 CBFA2T2 CIR1 CXCL14 FAM3D GPI HOXB7 IL1F5 ALX1 CBFA2T3 CITED1 CXCL16 FASLG GREM1 HOXB9 IL1F6 ARGFX CBFB CITED2 CXCL3 FBLN1 GREM2 HOXC4 IL1F7 -

TWIST1 and TWIST2 Regulate Glycogen Storage and Inflammatory
J M MUDRY and others TWIST regulation of metabolism 224:3 303–313 Research in muscle TWIST1 and TWIST2 regulate glycogen storage and inflammatory genes in skeletal muscle Correspondence 1 1 2 1,2 Jonathan M Mudry , Julie Massart , Ferenc L M Szekeres and Anna Krook should be addressed to A Krook 1Section for Integrative Physiology, Department of Molecular Medicine and Surgery, and 2Section for Integrative Email Physiology, Department of Physiology and Pharmacology, Karolinska Institutet, SE-171 77 Stockholm, Sweden [email protected] Abstract TWIST proteins are important for development of embryonic skeletal muscle and play a role Key Words in the metabolism of tumor and white adipose tissue. The impact of TWIST on metabolism " TWIST in skeletal muscle is incompletely studied. Our aim was to assess the impact of TWIST1 " metabolism and TWIST2 overexpression on glucose and lipid metabolism. In intact mouse muscle, " glycogen overexpression of Twist reduced total glycogen content without altering glucose uptake. " skeletal muscle Expression of TWIST1 or TWIST2 reduced Pdk4 mRNA, while increasing mRNA levels of Il6, Tnfa, and Il1b. Phosphorylation of AKT was increased and protein abundance of acetyl CoA carboxylase (ACC) was decreased in skeletal muscle overexpressing TWIST1 or TWIST2. Glycogen synthesis and fatty acid oxidation remained stable in C2C12 cells overexpressing TWIST1 or TWIST2. Finally, skeletal muscle mRNA levels remain unaltered in ob/ob mice, type 2 diabetic patients, or in healthy subjects before and after 3 months of exercise training. Journal of Endocrinology Collectively, our results indicate that TWIST1 and TWIST2 are expressed in skeletal muscle. Overexpression of these proteins impacts proteins in metabolic pathways and mRNA level of cytokines. -

Accompanies CD8 T Cell Effector Function Global DNA Methylation
Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer, Benjamin G. Barwick, Benjamin A. Youngblood, Rafi Ahmed and Jeremy M. Boss This information is current as of October 1, 2021. J Immunol 2013; 191:3419-3429; Prepublished online 16 August 2013; doi: 10.4049/jimmunol.1301395 http://www.jimmunol.org/content/191/6/3419 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130139 Material 5.DC1 References This article cites 81 articles, 25 of which you can access for free at: http://www.jimmunol.org/content/191/6/3419.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists by guest on October 1, 2021 • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer,* Benjamin G. Barwick,* Benjamin A. Youngblood,*,† Rafi Ahmed,*,† and Jeremy M. -

A Range of Clinical Phenotypes Associated with Mutations in CRX, a Photoreceptor Transcription-Factor Gene Melanie M
Am. J. Hum. Genet. 63:1307–1315, 1998 A Range of Clinical Phenotypes Associated with Mutations in CRX, a Photoreceptor Transcription-Factor Gene Melanie M. Sohocki,1 Lori S. Sullivan,1,2 Helen A. Mintz-Hittner,2 David Birch,3 John R. Heckenlively,4 Carol L. Freund,5 Roderick R. McInnes,5 and Stephen P. Daiger1,2 1Human Genetics Center, School of Public Health, and 2Department of Ophthalmology and Visual Science, The University of Texas Health Science Center, Houston; 3Retina Foundation of the Southwest, Dallas; 4Jules Stein Eye Institute, University of California, Los Angeles; and 5Program in Developmental Biology, Research Institute, Hospital for Sick Children, Toronto Summary is the diagnosis of some of these diseases difficult because of overlapping phenotypes, but mutations within a single Mutations in the retinal-expressed gene CRX (cone-rod gene may cause very different phenotypes. For example, homeobox gene) have been associated with dominant different mutations in one gene, peripherin/RDS, have cone-rod dystrophy and with de novo Leber congenital been associated with several forms of retinal degenera- amaurosis. However, CRX is a transcription factor for tion, such as cone-rod dystrophy, cone degeneration, and several retinal genes, including the opsins and the gene retinitis pigmentosa, affecting both the macula and the for interphotoreceptor retinoid binding protein. Because panretinal structures (Weleber et al. 1993; Wells et al. loss of CRX function could alter the expression of a 1993; Keen et al. 1994; Nakazawa et al. 1994, 1996). number of other retinal proteins, we screened for mu- Recently, mutations within the photoreceptor-ex- tations in the CRX gene in probands with a range of pressed gene CRX (cone-rod homeobox gene; MIM degenerative retinal diseases. -

An All-To-All Approach to the Identification of Sequence-Specific Readers for Epigenetic DNA Modifications on Cytosine
bioRxiv preprint doi: https://doi.org/10.1101/638700; this version posted May 16, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. An All-to-All Approach to the Identification of Sequence-Specific Readers for Epigenetic DNA Modifications on Cytosine Guang Song1,6, Guohua Wang2,6, Ximei Luo2,3,6, Ying Cheng4, Qifeng Song1, Jun Wan3, Cedric Moore1, Hongjun Song5, Peng Jin4, Jiang Qian3,7,*, Heng Zhu1,7,8,* 1Department of Pharmacology and Molecular Sciences, Johns Hopkins University School of Medicine, Baltimore, MD 21205, USA 2School of Computer Science and Technology, Harbin Institute of Technology, Harbin, Heilongjiang 150001, China 3Department of Ophthalmology, Johns Hopkins University School of Medicine, Baltimore, MD 21205, USA 4Department of Human Genetics, Emory University School of Medicine, Atlanta, GA 30322, USA 5Department of Neuroscience and Mahoney Institute for Neurosciences, University of Pennsylvania, Philadelphia, PA 19104, USA 6These authors contributed equally 7Senior author 8Lead Contact *Correspondence: [email protected] (H.Z.), [email protected] (J.Q.). 1 bioRxiv preprint doi: https://doi.org/10.1101/638700; this version posted May 16, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. SUMMARY Epigenetic modifications of DNA in mammals play important roles in many biological processes. Identification of readers of these epigenetic marks is a critical step towards understanding the underlying molecular mechanisms. Here, we report the invention and application of an all-to-all approach, dubbed Digital Affinity Profiling via Proximity Ligation (DAPPL), to simultaneously profile human TF-DNA interactions using mixtures of random DNA libraries carrying four different epigenetic modifications (i.e., 5-methylcytosine, 5- hydroxymethylcytosine, 5-formylcytosine, and 5-carboxylcytosine). -
The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas
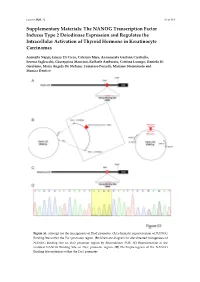
Cancers 2020, 12 S1 of S18 Supplementary Materials: The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas Annarita Nappi, Emery Di Cicco, Caterina Miro, Annunziata Gaetana Cicatiello, Serena Sagliocchi, Giuseppina Mancino, Raffaele Ambrosio, Cristina Luongo, Daniela Di Girolamo, Maria Angela De Stefano, Tommaso Porcelli, Mariano Stornaiuolo and Monica Dentice Figure S1. Strategy for the mutagenesis of Dio2 promoter. (A) Schematic representation of NANOG Binding Site within the Dio2 promoter region. (B) Schematic diagram for site‐directed mutagenesis of NANOG Binding Site on Dio2 promoter region by Recombinant PCR. (C) Representation of the mutated NANOG Binding Site on Dio2 promoter region. (D) Electropherogram of the NANOG Binding Site mutation within the Dio2 promoter. Cancers 2020, 12 S2 of S18 Figure S2. Strategy for the silencing of NANOG expression. (A) Cloning strategies for the generation of NANOG shRNA expression vectors. (B) Electropherograms of the NANOG shRNA sequences cloned into pcDNA3.1 vector. (C) Validation of effective NANOG down-modulation by two different NANOG shRNA vectors was assessed by Western Blot analysis of NANOG expression in BCC cells. (D) Quantification of NANOG protein levels versus Tubulin levels in the same experiment as in C is represented by histograms. Cancers 2020, 12 S3 of S18 Figure S3. The CD34+ cells are characterized by the expression of typical epithelial stemness genes. The mRNA levels of a panel of indicated stemness markers of epidermis were measured by Real Time PCR in the same experiment indicated in figure 3F and G. Cancers 2020, 12 S4 of S18 Figure S4. -

Gene Networks Activated by Specific Patterns of Action Potentials in Dorsal Root Ganglia Neurons Received: 10 August 2016 Philip R
www.nature.com/scientificreports OPEN Gene networks activated by specific patterns of action potentials in dorsal root ganglia neurons Received: 10 August 2016 Philip R. Lee1,*, Jonathan E. Cohen1,*, Dumitru A. Iacobas2,3, Sanda Iacobas2 & Accepted: 23 January 2017 R. Douglas Fields1 Published: 03 March 2017 Gene regulatory networks underlie the long-term changes in cell specification, growth of synaptic connections, and adaptation that occur throughout neonatal and postnatal life. Here we show that the transcriptional response in neurons is exquisitely sensitive to the temporal nature of action potential firing patterns. Neurons were electrically stimulated with the same number of action potentials, but with different inter-burst intervals. We found that these subtle alterations in the timing of action potential firing differentially regulates hundreds of genes, across many functional categories, through the activation or repression of distinct transcriptional networks. Our results demonstrate that the transcriptional response in neurons to environmental stimuli, coded in the pattern of action potential firing, can be very sensitive to the temporal nature of action potential delivery rather than the intensity of stimulation or the total number of action potentials delivered. These data identify temporal kinetics of action potential firing as critical components regulating intracellular signalling pathways and gene expression in neurons to extracellular cues during early development and throughout life. Adaptation in the nervous system in response to external stimuli requires synthesis of new gene products in order to elicit long lasting changes in processes such as development, response to injury, learning, and memory1. Information in the environment is coded in the pattern of action-potential firing, therefore gene transcription must be regulated by the pattern of neuronal firing. -

Genequery™ Human Circadian Rhythm Qpcr Array Kit (GQH-CIR) Catalog #GK060
GeneQuery™ Human Circadian Rhythm qPCR Array Kit (GQH-CIR) Catalog #GK060 Product Description ScienCell's GeneQuery™ Human Circadian Rhythm qPCR Array Kit (GQH-CIR) profiles 88 key genes involved in the regulating changes that occur on a roughly 24-hr cycle. Circadian rhythms are primarily responses to changes in light and are found in most living organisms. Not to be confused with biological clocks, circadian rhythms can influence sleep patterns, hormone release, body temperature, and other homeostatic functions. Below are brief examples of how included genes may be grouped according to their function: Clock regulation: ARNTL, BHLHE40, CLOCK, PER1, RORA Casein kinases: CSNK1A1, CSNK1D, CSNK1E, CSNK2A1 CREB pathway: CAMK2A, HTR7, MAPK1, PRKACB, AANAT Transcription factors: SMAD4, IRF1, NFIL3, RORC, SP1, GeneQuery™ qPCR array kits are qPCR ready in a 96-well plate format, with each well containing one primer set that recognizes and efficiently amplifies a specific target gene's cDNA. The carefully designed primers ensure that: (i) the optimal annealing temperature in qPCR analysis is 65°C (with 2 mM Mg2+ and no DMSO); (ii) the primer set recognizes all known transcript variants of the target gene, unless otherwise noted; and (iii) only one gene is amplified. Each primer set has been validated by qPCR with melt curve analysis and gel electrophoresis. GeneQuery™ qPCR Array Kit Controls Each GeneQuery™ plate contains eight controls (Figure 1): Five target housekeeping genes (β-actin, GAPDH, LDHA, NONO, and PPIH), which enable normalization of data. The Genomic DNA (gDNA) Control (GDC), which detects gDNA contamination in cDNA samples. This primer set targets a non-transcribed region of the genome. -

Estrogen Promotes Mandibular Condylar Fibrocartilage
www.nature.com/scientificreports OPEN Estrogen Promotes Mandibular Condylar Fibrocartilage Chondrogenesis and Inhibits Received: 27 November 2017 Accepted: 18 May 2018 Degeneration via Estrogen Published: xx xx xxxx Receptor Alpha in Female Mice Jennifer L. Robinson 1,3, Paola Soria2, Manshan Xu2, Mark Vrana1, Jefrey Luchetti1, Helen H. Lu3, Jing Chen2 & Sunil Wadhwa2 Temporomandibular joint degenerative disease (TMJ-DD) is a chronic form of TMJ disorder that specifcally aficts people over the age of 40 and targets women at a higher rate than men. Prevalence of TMJ-DD in this population suggests that estrogen loss plays a role in the disease pathogenesis. Thus, the goal of the present study was to determine the role of estrogen on chondrogenesis and homeostasis via estrogen receptor alpha (ERα) during growth and maturity of the joint. Young and mature WT and ERαKO female mice were subjected to ovariectomy procedures and then given placebo or estradiol treatment. The efect of estrogen via ERα on fbrocartilage morphology, matrix production, and protease activity was assessed. In the young mice, estrogen via ERα promoted mandibular condylar fbrocartilage chondrogenesis partly by inhibiting the canonical Wnt signaling pathway through upregulation of sclerostin (Sost). In the mature mice, protease activity was partly inhibited with estrogen treatment via the upregulation and activity of protease inhibitor 15 (Pi15) and alpha-2- macroglobulin (A2m). The results from this work provide a mechanistic understanding of estradiol on TMJ growth and homeostasis and can be utilized for development of therapeutic targets to promote regeneration and inhibit degeneration of the mandibular condylar fbrocartilage. Temporomandibular joint degenerative disease (TMJ-DD) is marked by degradation and premature calcifcation of the extracellular matrix (ECM) of the articular mandibular condylar fbrocartilage.