Molecular Mechanisms in Muscular Dystrophy : a Gene Expression Profiling Study Turk, R
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Whole Exome Sequencing in ADHD Trios from Single and Multi-Incident Families Implicates New Candidate Genes and Highlights Polygenic Transmission
European Journal of Human Genetics (2020) 28:1098–1110 https://doi.org/10.1038/s41431-020-0619-7 ARTICLE Whole exome sequencing in ADHD trios from single and multi-incident families implicates new candidate genes and highlights polygenic transmission 1,2 1 1,2 1,3 1 Bashayer R. Al-Mubarak ● Aisha Omar ● Batoul Baz ● Basma Al-Abdulaziz ● Amna I. Magrashi ● 1,2 2 2,4 2,4,5 6 Eman Al-Yemni ● Amjad Jabaan ● Dorota Monies ● Mohamed Abouelhoda ● Dejene Abebe ● 7 1,2 Mohammad Ghaziuddin ● Nada A. Al-Tassan Received: 26 September 2019 / Revised: 26 February 2020 / Accepted: 10 March 2020 / Published online: 1 April 2020 © The Author(s) 2020. This article is published with open access Abstract Several types of genetic alterations occurring at numerous loci have been described in attention deficit hyperactivity disorder (ADHD). However, the role of rare single nucleotide variants (SNVs) remains under investigated. Here, we sought to identify rare SNVs with predicted deleterious effect that may contribute to ADHD risk. We chose to study ADHD families (including multi-incident) from a population with a high rate of consanguinity in which genetic risk factors tend to 1234567890();,: 1234567890();,: accumulate and therefore increasing the chance of detecting risk alleles. We employed whole exome sequencing (WES) to interrogate the entire coding region of 16 trios with ADHD. We also performed enrichment analysis on our final list of genes to identify the overrepresented biological processes. A total of 32 rare variants with predicted damaging effect were identified in 31 genes. At least two variants were detected per proband, most of which were not exclusive to the affected individuals. -
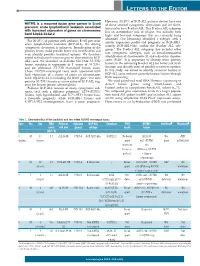
NUTM1 Is a Recurrent Fusion Gene Partner in B-Cell Precursor Acute
LETTERS TO THE EDITOR However, 20-25% of BCP-ALL patients do not have one NUTM1 is a recurrent fusion gene partner in B-cell of these sentinel cytogenetic aberrations and are there- precursor acute lymphoblastic leukemia associated fore said to have B-other ALL. This B-other ALL subgroup with increased expression of genes on chromosome has an intermediate risk of relapse, but includes both band 10p12.31-12.2 high- and low-risk subgroups that are currently being identified. Our laboratory identified a subtype with a For 20-25% of patients with pediatric B-cell precursor similar expression profile and prognosis as BCR-ABL1, acute lymphoblastic leukemia (BCP-ALL), the driving namely BCR-ABL1-like, within the B-other ALL sub- cytogenetic aberration is unknown. Identification of the group.2 The B-other ALL subgroup also includes other primary lesion could provide better risk stratification and rare cytogenetic subtypes, such as intrachromosomal even identify possible treatment options. We therefore amplification of chromosome 21 and a dicentric chromo- aimed to find novel recurrent genetic aberrations in BCP- 1 ALL cases. We identified an in-frame SLC12A6-NUTM1 some (9;20). It is important to identify more primary fusion, resulting in expression of 3’ exons of NUTM1, lesions in the remaining B-other ALL for better risk strat- and six additional NUTM1-rearranged fusion cases. ification and identification of possible treatment options. These NUTM1-rearranged cases were associated with In this study, we aimed to identify recurrent fusions in high expression of a cluster of genes on chromosome BCP-ALL cases without currently known lesions through band 10p12.31-12.2, including the BMI1 gene. -

Viewed and Published Immediately Upon Acceptance Cited in Pubmed and Archived on Pubmed Central Yours — You Keep the Copyright
BMC Genomics BioMed Central Research article Open Access Differential gene expression in ADAM10 and mutant ADAM10 transgenic mice Claudia Prinzen1, Dietrich Trümbach2, Wolfgang Wurst2, Kristina Endres1, Rolf Postina1 and Falk Fahrenholz*1 Address: 1Johannes Gutenberg-University, Institute of Biochemistry, Mainz, Johann-Joachim-Becherweg 30, 55128 Mainz, Germany and 2Helmholtz Zentrum München – German Research Center for Environmental Health, Institute for Developmental Genetics, Ingolstädter Landstraße 1, 85764 Neuherberg, Germany Email: Claudia Prinzen - [email protected]; Dietrich Trümbach - [email protected]; Wolfgang Wurst - [email protected]; Kristina Endres - [email protected]; Rolf Postina - [email protected]; Falk Fahrenholz* - [email protected] * Corresponding author Published: 5 February 2009 Received: 19 June 2008 Accepted: 5 February 2009 BMC Genomics 2009, 10:66 doi:10.1186/1471-2164-10-66 This article is available from: http://www.biomedcentral.com/1471-2164/10/66 © 2009 Prinzen et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: In a transgenic mouse model of Alzheimer disease (AD), cleavage of the amyloid precursor protein (APP) by the α-secretase ADAM10 prevented amyloid plaque formation, and alleviated cognitive deficits. Furthermore, ADAM10 overexpression increased the cortical synaptogenesis. These results suggest that upregulation of ADAM10 in the brain has beneficial effects on AD pathology. Results: To assess the influence of ADAM10 on the gene expression profile in the brain, we performed a microarray analysis using RNA isolated from brains of five months old mice overexpressing either the α-secretase ADAM10, or a dominant-negative mutant (dn) of this enzyme. -

Improved Detection of Gene Fusions by Applying Statistical Methods Reveals New Oncogenic RNA Cancer Drivers
bioRxiv preprint doi: https://doi.org/10.1101/659078; this version posted June 3, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Improved detection of gene fusions by applying statistical methods reveals new oncogenic RNA cancer drivers Roozbeh Dehghannasiri1, Donald Eric Freeman1,2, Milos Jordanski3, Gillian L. Hsieh1, Ana Damljanovic4, Erik Lehnert4, Julia Salzman1,2,5* Author affiliation 1Department of Biochemistry, Stanford University, Stanford, CA 94305 2Department of Biomedical Data Science, Stanford University, Stanford, CA 94305 3Department of Computer Science, University of Belgrade, Belgrade, Serbia 4Seven Bridges Genomics, Cambridge, MA 02142 5Stanford Cancer Institute, Stanford, CA 94305 *Corresponding author [email protected] Short Abstract: The extent to which gene fusions function as drivers of cancer remains a critical open question. Current algorithms do not sufficiently identify false-positive fusions arising during library preparation, sequencing, and alignment. Here, we introduce a new algorithm, DEEPEST, that uses statistical modeling to minimize false-positives while increasing the sensitivity of fusion detection. In 9,946 tumor RNA-sequencing datasets from The Cancer Genome Atlas (TCGA) across 33 tumor types, DEEPEST identifies 31,007 fusions, 30% more than identified by other methods, while calling ten-fold fewer false-positive fusions in non-transformed human tissues. We leverage the increased precision of DEEPEST to discover new cancer biology. For example, 888 new candidate oncogenes are identified based on over-representation in DEEPEST-Fusion calls, and 1,078 previously unreported fusions involving long intergenic noncoding RNAs partners, demonstrating a previously unappreciated prevalence and potential for function. -

Cytogenetic and Molecular Characterization of the Macro- And
University of Ulm Department of Human Genetics Prof. Dr. med. Walther Vogel Cytogenetic and Molecular Characterization of the Macro- and Micro-inversions, which Distinguish the Human and the Chimpanzee Karyotypes - from Speciation to Polymorphism Thesis Applying for the Degree of Doctor of Human Biology (Dr. hum. biol.) Faculty of Medicine University of Ulm Presented by Justyna Monika Szamalek from Wrze śnia in Poland 2006 Amtierender Dekan: Prof. Dr. Klaus-Michael Debatin 1. Berichterstatter: Prof. Dr. med. Horst Hameister 2. Berichterstatter: Prof. Dr. med. Konstanze Döhner Tag der Promotion: 28.07.2006 Content Content 1. Introduction ...................................................................................................................7 1.1. Primate phylogeny........................................................................................................7 1.2. Africa as the place of human origin and the living area of the present-day chimpanzee populations .................................................................9 1.3. Cytogenetic and molecular differences between human and chimpanzee genomes.............................................................................................10 1.4. Cytogenetic and molecular differences between common chimpanzee and bonobo genomes................................................................................17 1.5. Theory of speciation .....................................................................................................18 1.6. Theory of selection -

Genome-Wide Expression Profiling Establishes Novel Modulatory Roles
Batra et al. BMC Genomics (2017) 18:252 DOI 10.1186/s12864-017-3635-4 RESEARCHARTICLE Open Access Genome-wide expression profiling establishes novel modulatory roles of vitamin C in THP-1 human monocytic cell line Sakshi Dhingra Batra, Malobi Nandi, Kriti Sikri and Jaya Sivaswami Tyagi* Abstract Background: Vitamin C (vit C) is an essential dietary nutrient, which is a potent antioxidant, a free radical scavenger and functions as a cofactor in many enzymatic reactions. Vit C is also considered to enhance the immune effector function of macrophages, which are regarded to be the first line of defence in response to any pathogen. The THP- 1 cell line is widely used for studying macrophage functions and for analyzing host cell-pathogen interactions. Results: We performed a genome-wide temporal gene expression and functional enrichment analysis of THP-1 cells treated with 100 μM of vit C, a physiologically relevant concentration of the vitamin. Modulatory effects of vitamin C on THP-1 cells were revealed by differential expression of genes starting from 8 h onwards. The number of differentially expressed genes peaked at the earliest time-point i.e. 8 h followed by temporal decline till 96 h. Further, functional enrichment analysis based on statistically stringent criteria revealed a gamut of functional responses, namely, ‘Regulation of gene expression’, ‘Signal transduction’, ‘Cell cycle’, ‘Immune system process’, ‘cAMP metabolic process’, ‘Cholesterol transport’ and ‘Ion homeostasis’. A comparative analysis of vit C-mediated modulation of gene expression data in THP-1cells and human skin fibroblasts disclosed an overlap in certain functional processes such as ‘Regulation of transcription’, ‘Cell cycle’ and ‘Extracellular matrix organization’, and THP-1 specific responses, namely, ‘Regulation of gene expression’ and ‘Ion homeostasis’. -

Evolution and Diversity of Copy Number Variation in the Great Ape Lineage
Downloaded from genome.cshlp.org on September 24, 2021 - Published by Cold Spring Harbor Laboratory Press Evolution and diversity of copy number variation in the great ape lineage Peter H. Sudmant1, John Huddleston1,7, Claudia R. Catacchio2, Maika Malig1, LaDeana W. Hillier3, Carl Baker1, Kiana Mohajeri1, Ivanela Kondova4, Ronald E. Bontrop4, Stephan Persengiev4, Francesca Antonacci2, Mario Ventura2, Javier Prado-Martinez5, Tomas 5,6 1,7 Marques-Bonet , and Evan E. Eichler 1. Department of Genome Sciences, University of Washington, Seattle, WA, USA 2. University of Bari, Bari, Italy 3. The Genome Institute, Washington University School of Medicine, St. Louis, MO, USA 4. Department of Comparative Genetics, Biomedical Primate Research Centre, Rijswijk, The Netherlands 5. Institut de Biologia Evolutiva, (UPF-CSIC) Barcelona, Spain 6. Institució Catalana de Recerca i Estudis Avançats (ICREA), Barcelona, Spain 7. Howard Hughes Medical Institute, University of Washington, Seattle, WA, USA Correspondence to: Evan Eichler Department of Genome Sciences University of Washington School of Medicine Foege S-413A, Box 355065 3720 15th Ave NE Seattle, WA 98195 E-mail: [email protected] 1 Downloaded from genome.cshlp.org on September 24, 2021 - Published by Cold Spring Harbor Laboratory Press ABSTRACT Copy number variation (CNV) contributes to the genetic basis of disease and has significantly restructured the genomes of humans and great apes. The diversity and rate of this process, however, has not been extensively explored among the great ape lineages. We analyzed 97 deeply sequenced great ape and human genomes and estimate that 16% (469 Mbp) of the hominid genome has been affected by recent copy number changes. -

Improved Detection of Gene Fusions by Applying Statistical Methods Reveals Oncogenic RNA Cancer Drivers Roozbeh Dehghannasiria, Donald E
Improved detection of gene fusions by applying statistical methods reveals oncogenic RNA cancer drivers Roozbeh Dehghannasiria, Donald E. Freemana,b, Milos Jordanskic, Gillian L. Hsieha, Ana Damljanovicd, Erik Lehnertd, and Julia Salzmana,b,e,1 aDepartment of Biochemistry, Stanford University, Stanford, CA 94305; bDepartment of Biomedical Data Science, Stanford University, Stanford, CA 94305; cDepartment of Computer Science, University of Belgrade, 11000 Belgrade, Serbia; dSeven Bridges Genomics Inc., Cambridge, MA 02142; and eStanford Cancer Institute, Stanford University, Stanford, CA 94305 Edited by Ali Torkamani, The Scripps Research Institute, La Jolla, CA, and accepted by Editorial Board Member Peter K. Vogt June 14, 2019 (received for review January 10, 2019) The extent to which gene fusions function as drivers of can- of multiple algorithms and filtering lists of fusions using manual cer remains a critical open question. Current algorithms do not approaches (13–15). These approaches lead to what third-party sufficiently identify false-positive fusions arising during library reviews agree is imprecise fusion discovery and bias against dis- preparation, sequencing, and alignment. Here, we introduce Data- covering novel oncogenes (15–17). This suboptimal performance Enriched Efficient PrEcise STatistical fusion detection (DEEPEST), becomes more problematic when fusion detection is deployed on an algorithm that uses statistical modeling to minimize false- large cancer sequencing datasets that contain thousands or tens positives while increasing the sensitivity of fusion detection. In of thousands of samples. In such scenarios, precise fusion detec- 9,946 tumor RNA-sequencing datasets from The Cancer Genome tion must overcome the problem of multiple hypothesis testing: Atlas (TCGA) across 33 tumor types, DEEPEST identifies 31,007 each algorithm is testing for fusions thousands of times, a regime fusions, 30% more than identified by other methods, while known to introduce FPs. -

High-Resolution Gene Maps of Horse Chromosomes 14 and 21: Additional Insights Into Evolution and Rearrangements of HSA5 Homologs in Mammals
Genomics 89 (2007) 89–112 www.elsevier.com/locate/ygeno High-resolution gene maps of horse chromosomes 14 and 21: Additional insights into evolution and rearrangements of HSA5 homologs in mammals Glenda Goh a,1, Terje Raudsepp a,1, Keith Durkin a, Michelle L. Wagner b, Alejandro A. Schäffer c, Richa Agarwala c, Teruaki Tozaki d, ⁎ James R. Mickelson b, Bhanu P. Chowdhary a,e, a Department of Veterinary Integrative Biosciences, College of Veterinary Medicine and Biomedical Sciences, Texas A&M University, College Station, TX 77843, USA b Department of Veterinary Biosciences, University of Minnesota, 295f AS/VM, St. Paul, MN 55108, USA c National Center for Biotechnology Information, National Institutes of Health, Department of Health and Human Services, Bethesda, MD 20894, USA d Molecular Genetics Section, Laboratory of Racing Chemistry, Tochigi 320-0851, Japan e Department of Animal Science, College of Agriculture and Life Science, Texas A&M University, College Station, TX 77843, USA Received 12 May 2006; accepted 19 June 2006 Available online 17 August 2006 Abstract High-resolution physically ordered gene maps for equine homologs of human chromosome 5 (HSA5), viz., horse chromosomes 14 and 21 (ECA14 and ECA21), were generated by adding 179 new loci (131 gene-specific and 48 microsatellites) to the existing maps of the two chromosomes. The loci were mapped primarily by genotyping on a 5000-rad horse×hamster radiation hybrid panel, of which 28 were mapped by fluorescence in situ hybridization. The approximately fivefold increase in the number of mapped markers on the two chromosomes improves the average resolution of the map to 1 marker/0.9 Mb. -

Downloaded Per Proteome Cohort Via the Web- Site Links of Table 1, Also Providing Information on the Deposited Spectral Datasets
www.nature.com/scientificreports OPEN Assessment of a complete and classifed platelet proteome from genome‑wide transcripts of human platelets and megakaryocytes covering platelet functions Jingnan Huang1,2*, Frauke Swieringa1,2,9, Fiorella A. Solari2,9, Isabella Provenzale1, Luigi Grassi3, Ilaria De Simone1, Constance C. F. M. J. Baaten1,4, Rachel Cavill5, Albert Sickmann2,6,7,9, Mattia Frontini3,8,9 & Johan W. M. Heemskerk1,9* Novel platelet and megakaryocyte transcriptome analysis allows prediction of the full or theoretical proteome of a representative human platelet. Here, we integrated the established platelet proteomes from six cohorts of healthy subjects, encompassing 5.2 k proteins, with two novel genome‑wide transcriptomes (57.8 k mRNAs). For 14.8 k protein‑coding transcripts, we assigned the proteins to 21 UniProt‑based classes, based on their preferential intracellular localization and presumed function. This classifed transcriptome‑proteome profle of platelets revealed: (i) Absence of 37.2 k genome‑ wide transcripts. (ii) High quantitative similarity of platelet and megakaryocyte transcriptomes (R = 0.75) for 14.8 k protein‑coding genes, but not for 3.8 k RNA genes or 1.9 k pseudogenes (R = 0.43–0.54), suggesting redistribution of mRNAs upon platelet shedding from megakaryocytes. (iii) Copy numbers of 3.5 k proteins that were restricted in size by the corresponding transcript levels (iv) Near complete coverage of identifed proteins in the relevant transcriptome (log2fpkm > 0.20) except for plasma‑derived secretory proteins, pointing to adhesion and uptake of such proteins. (v) Underrepresentation in the identifed proteome of nuclear‑related, membrane and signaling proteins, as well proteins with low‑level transcripts. -

Effect of Prematurity on Genome Wide Methylation in the Placenta Jessica Schuster1, Alper Uzun2, Joan Stablia1, Christoph Schorl3, Mari Mori4 and James F
Schuster et al. BMC Medical Genetics (2019) 20:116 https://doi.org/10.1186/s12881-019-0835-6 RESEARCH ARTICLE Open Access Effect of prematurity on genome wide methylation in the placenta Jessica Schuster1, Alper Uzun2, Joan Stablia1, Christoph Schorl3, Mari Mori4 and James F. Padbury2,5* Abstract Background: Preterm birth is a significant clinical problem and an enormous burden on society, affecting one in eight pregnant women and their newborns. Despite decades of research, the molecular mechanism underlying its pathogenesis remains unclear. Many studies have shown that preterm birth is associated with health risks across the later life course. The “fetal origins” hypothesis postulates that adverse intrauterine exposures are associated with later disease susceptibility. Our recent studies have focused on the placental epigenome at term. We extended these studies to genome-wide placental DNA methylation across a wide range of gestational ages. We applied methylation dependent immunoprecipitation/DNA sequencing (MeDIP-seq) to 9 placentas with gestational age from 25 weeks to term to identify differentially methylated regions (DMRs). Results: Enrichment analysis revealed 427 DMRs with nominally significant differences in methylation between preterm and term placentas (p < 0.01) and 21 statistically significant DMRs after multiple comparison correction (FDR p < 0.05), of which 62% were hypo-methylated in preterm placentas vs term placentas. The majority of DMRs were in distal intergenic regions and introns. Significantly enriched pathways identified by Ingenuity Pathway Analysis (IPA) included Citrulline-Nitric Oxide Cycle and Fcy Receptor Mediated Phagocytosis in macrophages. The DMR gene set overlapped placental gene expression data, genes and pathways associated evolutionarily with preterm birth. -

Variation in Protein Coding Genes Identifies Information Flow
bioRxiv preprint doi: https://doi.org/10.1101/679456; this version posted June 21, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Animal complexity and information flow 1 1 2 3 4 5 Variation in protein coding genes identifies information flow as a contributor to 6 animal complexity 7 8 Jack Dean, Daniela Lopes Cardoso and Colin Sharpe* 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Institute of Biological and Biomedical Sciences 25 School of Biological Science 26 University of Portsmouth, 27 Portsmouth, UK 28 PO16 7YH 29 30 * Author for correspondence 31 [email protected] 32 33 Orcid numbers: 34 DLC: 0000-0003-2683-1745 35 CS: 0000-0002-5022-0840 36 37 38 39 40 41 42 43 44 45 46 47 48 49 Abstract bioRxiv preprint doi: https://doi.org/10.1101/679456; this version posted June 21, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Animal complexity and information flow 2 1 Across the metazoans there is a trend towards greater organismal complexity. How 2 complexity is generated, however, is uncertain. Since C.elegans and humans have 3 approximately the same number of genes, the explanation will depend on how genes are 4 used, rather than their absolute number.