Catalytic Promiscuity and the Evolution of New Enzymatic Activities Patrick J O’Brien and Daniel Herschlag
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Sphingolipid Metabolism Diseases ⁎ Thomas Kolter, Konrad Sandhoff
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Biochimica et Biophysica Acta 1758 (2006) 2057–2079 www.elsevier.com/locate/bbamem Review Sphingolipid metabolism diseases ⁎ Thomas Kolter, Konrad Sandhoff Kekulé-Institut für Organische Chemie und Biochemie der Universität, Gerhard-Domagk-Str. 1, D-53121 Bonn, Germany Received 23 December 2005; received in revised form 26 April 2006; accepted 23 May 2006 Available online 14 June 2006 Abstract Human diseases caused by alterations in the metabolism of sphingolipids or glycosphingolipids are mainly disorders of the degradation of these compounds. The sphingolipidoses are a group of monogenic inherited diseases caused by defects in the system of lysosomal sphingolipid degradation, with subsequent accumulation of non-degradable storage material in one or more organs. Most sphingolipidoses are associated with high mortality. Both, the ratio of substrate influx into the lysosomes and the reduced degradative capacity can be addressed by therapeutic approaches. In addition to symptomatic treatments, the current strategies for restoration of the reduced substrate degradation within the lysosome are enzyme replacement therapy (ERT), cell-mediated therapy (CMT) including bone marrow transplantation (BMT) and cell-mediated “cross correction”, gene therapy, and enzyme-enhancement therapy with chemical chaperones. The reduction of substrate influx into the lysosomes can be achieved by substrate reduction therapy. Patients suffering from the attenuated form (type 1) of Gaucher disease and from Fabry disease have been successfully treated with ERT. © 2006 Elsevier B.V. All rights reserved. Keywords: Ceramide; Lysosomal storage disease; Saposin; Sphingolipidose Contents 1. Sphingolipid structure, function and biosynthesis ..........................................2058 1.1. -

Genetic Basis of Simple and Complex Traits with Relevance to Avian Evolution
Genetic basis of simple and complex traits with relevance to avian evolution Małgorzata Anna Gazda Doctoral Program in Biodiversity, Genetics and Evolution D Faculdade de Ciências da Universidade do Porto 2019 Supervisor Miguel Jorge Pinto Carneiro, Auxiliary Researcher, CIBIO/InBIO, Laboratório Associado, Universidade do Porto Co-supervisor Ricardo Lopes, CIBIO/InBIO Leif Andersson, Uppsala University FCUP Genetic basis of avian traits Nota Previa Na elaboração desta tese, e nos termos do número 2 do Artigo 4º do Regulamento Geral dos Terceiros Ciclos de Estudos da Universidade do Porto e do Artigo 31º do D.L.74/2006, de 24 de Março, com a nova redação introduzida pelo D.L. 230/2009, de 14 de Setembro, foi efetuado o aproveitamento total de um conjunto coerente de trabalhos de investigação já publicados ou submetidos para publicação em revistas internacionais indexadas e com arbitragem científica, os quais integram alguns dos capítulos da presente tese. Tendo em conta que os referidos trabalhos foram realizados com a colaboração de outros autores, o candidato esclarece que, em todos eles, participou ativamente na sua conceção, na obtenção, análise e discussão de resultados, bem como na elaboração da sua forma publicada. Este trabalho foi apoiado pela Fundação para a Ciência e Tecnologia (FCT) através da atribuição de uma bolsa de doutoramento (PD/BD/114042/2015) no âmbito do programa doutoral em Biodiversidade, Genética e Evolução (BIODIV). 2 FCUP Genetic basis of avian traits Acknowledgements Firstly, I would like to thank to my all supervisors Miguel Carneiro, Ricardo Lopes and Leif Andersson, for the demanding task of supervising myself last four years. -

Characterization of a Mutation in a Family with Saposin B Deficiency
Proc. Nadl. Acad. Sci. USA Vol. 87, pp. 2541-2544, April 1990 Genetics Characterization of a mutation in a family with saposin B deficiency: A glycosylation site defect (sphingolipid activator protein/SAP-1/metachromatic leukodystrophy/arylsulfatase A) KEITH A. KRETZ*, GEOFFREY S. CARSON*, SATOSHI MORIMOTO*t, YASUO KISHIMOTO*, ARVAN L. FLUHARTYt, AND JOHN S. O'BPUEN*§ *Department of Neurosciences and Center for Molecular Genetics, University of California, San Diego, School of Medicine, M-034J, La Jolla, CA 92093; and tUniversity of California, Los Angeles, Mental Retardation Research Center Group at Lanterman Developmental Center, Pomona, CA 91766 Communicated by Dan L. Lindsley, January 19, 1990 ABSTRACT Saposins are small, heat-stable glycoproteins these four saposin proteins has now been isolated and their required for the hydrolysis of sphingolipids by specific lyso- activating properties have been determined (3-14). somal hydrolases. Saposins A, B, C, and D are derived by Saposins A and C specifically activate hydrolysis of glu- proteolytic processing from a single precursor protein named cocerebroside byB-glucosylceramidase (D-glucosyl-N-acyl- prosaposin. Saposin B, previously known as SAP-1 and sul- sphingosine glucohydrolase; EC 3.2.1.45) and ofgalactocere- fatide activator, stimulates the hydrolysis of a wide variety of broside by galactosylceramidase (D-galactosyl-N-acyl- substrates including cerebroside sulfate, GM1 ganglioside, and sphingosine galactohydrolase; EC 3.2.1.46) (3, 4). Saposin D globotriaosylceramide by arylsulfatase A, acid 8-galacto- specifically activates the hydrolysis of sphingomyelin by sidase, and a-galactosidase, respectively. Human saposin B sphingomyelin phosphodiesterase (sphingomyelin choline- deficiency, transmitted as an autosomal recessive trait, results phosphohydrolase; EC 3.1.4.12) (5). -

Functional Assignments in the Enolase Superfamily: Investigations of Two Divergent Groups of D-Galacturonate Dehydratases and Galactarate Dehydratase-Iii
FUNCTIONAL ASSIGNMENTS IN THE ENOLASE SUPERFAMILY: INVESTIGATIONS OF TWO DIVERGENT GROUPS OF D-GALACTURONATE DEHYDRATASES AND GALACTARATE DEHYDRATASE-III BY FIONA PATRICIA GRONINGER-POE i DISSERTATION Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biochemistry in the Graduate College of the University of Illinois at Urbana-Champaign, 2014 Urbana, Illinois Doctoral Committee: Professor John A. Gerlt, Chair Professor John E. Cronan, Jr. Associate Professor Rutilo Fratti Associate Professor Raven H. Huang ABSTRACT More than a decade after the genomic age, full genome sequencing is cost-effective and fast, allowing for the deposit of an ever increasing number of DNA sequences. New fields have arisen from this availability of genomic information, and the way we think about biochemistry and enzymology has been transformed. Unfortunately, there is no robust method for accurately determining the functions of enzymes encoded by these sequences that matches the speed in which genomes are deposited into public databases. Functional assignment of enzymes remains of utmost importance in understanding microbial metabolism and has applications in agriculture by examining bacterial plant pathogen metabolism and additionally in human health by providing metabolic context to the human gut microbiome. To aid in the functional identification of proteins, enzymes can be grouped into superfamilies which share common structural motifs as well as mechanistic features. To this end, the enolase superfamily is an excellent model system for functional assignment because more than half of the members still lack functional identification. Structurally, these enzymes contain substrate specificity residues in the N-terminal capping domain and catalytic residues in the C-terminal barrel domain. -

A General Binding Mechanism for All Human Sulfatases by the Formylglycine-Generating Enzyme
A general binding mechanism for all human sulfatases by the formylglycine-generating enzyme Dirk Roeser*, Andrea Preusser-Kunze†, Bernhard Schmidt†, Kathrin Gasow*, Julia G. Wittmann*, Thomas Dierks‡, Kurt von Figura†, and Markus Georg Rudolph*§ *Department of Molecular Structural Biology, University of Go¨ttingen, Justus-von-Liebig-Weg 11, D-37077 Go¨ttingen, Germany; †Department of Biochemistry II, Heinrich-Du¨ker-Weg 12, University of Go¨ttingen, D-37073 Go¨ttingen, Germany; and ‡Department of Biochemistry I, Universita¨tsstrasse 25, University of Bielefeld, D-33615 Bielefeld, Germany Edited by Carolyn R. Bertozzi, University of California, Berkeley, CA, and approved November 8, 2005 (received for review September 1, 2005) The formylglycine (FGly)-generating enzyme (FGE) uses molecular tases, suggesting a general binding mechanism of substrate sulfa- oxygen to oxidize a conserved cysteine residue in all eukaryotic tases by FGE. sulfatases to the catalytically active FGly. Sulfatases degrade and The details of how O2-dependent cysteine oxidation is mediated remodel sulfate esters, and inactivity of FGE results in multiple by FGE are unknown. As a first step toward the elucidation of the sulfatase deficiency, a fatal disease. The previously determined FGE molecular mechanism of FGly formation, we have previously crystal structure revealed two crucial cysteine residues in the active determined crystal structures of FGE in various oxidation states site, one of which was thought to be implicated in substrate (8). FGE adopts a novel fold with surprisingly little regular sec- 2ϩ binding. The other cysteine residue partakes in a novel oxygenase ondary structure and contains two structural Ca ions and two mechanism that does not rely on any cofactors. -

Supplementary File 2A Revised
Supplementary file 2A. Differentially expressed genes in aldosteronomas compared to all other samples, ranked according to statistical significance. Missing values were not allowed in aldosteronomas, but to a maximum of five in the other samples. Acc UGCluster Name Symbol log Fold Change P - Value Adj. P-Value B R99527 Hs.8162 Hypothetical protein MGC39372 MGC39372 2,17 6,3E-09 5,1E-05 10,2 AA398335 Hs.10414 Kelch domain containing 8A KLHDC8A 2,26 1,2E-08 5,1E-05 9,56 AA441933 Hs.519075 Leiomodin 1 (smooth muscle) LMOD1 2,33 1,3E-08 5,1E-05 9,54 AA630120 Hs.78781 Vascular endothelial growth factor B VEGFB 1,24 1,1E-07 2,9E-04 7,59 R07846 Data not found 3,71 1,2E-07 2,9E-04 7,49 W92795 Hs.434386 Hypothetical protein LOC201229 LOC201229 1,55 2,0E-07 4,0E-04 7,03 AA454564 Hs.323396 Family with sequence similarity 54, member B FAM54B 1,25 3,0E-07 5,2E-04 6,65 AA775249 Hs.513633 G protein-coupled receptor 56 GPR56 -1,63 4,3E-07 6,4E-04 6,33 AA012822 Hs.713814 Oxysterol bining protein OSBP 1,35 5,3E-07 7,1E-04 6,14 R45592 Hs.655271 Regulating synaptic membrane exocytosis 2 RIMS2 2,51 5,9E-07 7,1E-04 6,04 AA282936 Hs.240 M-phase phosphoprotein 1 MPHOSPH -1,40 8,1E-07 8,9E-04 5,74 N34945 Hs.234898 Acetyl-Coenzyme A carboxylase beta ACACB 0,87 9,7E-07 9,8E-04 5,58 R07322 Hs.464137 Acyl-Coenzyme A oxidase 1, palmitoyl ACOX1 0,82 1,3E-06 1,2E-03 5,35 R77144 Hs.488835 Transmembrane protein 120A TMEM120A 1,55 1,7E-06 1,4E-03 5,07 H68542 Hs.420009 Transcribed locus 1,07 1,7E-06 1,4E-03 5,06 AA410184 Hs.696454 PBX/knotted 1 homeobox 2 PKNOX2 1,78 2,0E-06 -

Lysosomal Hydrolases: Conversion of Acidic to Basic Forms by Neuraminidase
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Volume 13, number 1 FEBS LETTERS February 1971 LYSOSOMAL HYDROLASES: CONVERSION OF ACIDIC TO BASIC FORMS BY NEURAMINIDASE Alfred GOLDSTONE, Philip KONECNY and Harold KOENIG Neurology Service, VA Research Hospital; and Departments of Neurology and Biochemistry, Northwestern University Medical School, Chicago, Illinois, USA Received 21 December 1970 1. Introduction intervals to avoid overheating). The lysosomal lysate was ultracentrifuged at 100,000 g for 30 min to give A number of hydrolytic enzymes of lysosomal a soluble enzyme fraction. This fraction also contains origin occur in multiple forms which differ in such an acidic lipoprotein [4]. The insoluble residue was physicochemical properties as solubility, heat and pH suspended in 0.05 M acetate buffer pH 5.0 with 0.2% stability, pH optima and electrophoretic mobility [I]. Triton X-100 and resonicated to release the bound ly- Recent studies in this laboratory indicate that essen- sosomal enzymes together with additional acidic lipo- tially all the enzyme proteins of rat kidney and liver protein ([4] and A. Goldstone and H. Koenig, unpu- lysosomes are glycoproteins, and that at least some of blished observations). All operations were carried out these glycoproteins contain sialic acid [2-41. The at 4’. acidic form of lysosomal N-acetylhexosaminidase from Aliquots of the soluble fraction containing about human spleen [ 51 and kidney [6] is converted into 2 mg of protein were incubated with neuraminidase the basic form by treatment with neuraminidase. We from Cl. perfringens (Type Vl enzyme supplied by now confirm this finding for rat kidney and show Sigma), 0.1 mg/ml, or V. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Increased CHST15 Follows Decline in Arylsulfatase B (ARSB) and Disinhibition of Non-Canonical WNT Signaling: Potential Impact on Epithelial and Mesenchymal Identity
www.oncotarget.com Oncotarget, 2020, Vol. 11, (No. 24), pp: 2327-2344 Research Paper Increased CHST15 follows decline in arylsulfatase B (ARSB) and disinhibition of non-canonical WNT signaling: potential impact on epithelial and mesenchymal identity Sumit Bhattacharyya1,2, Leo Feferman1,2, Xiaorui Han3, Ke Xia3, Fuming Zhang3, Robert J. Linhardt3, Joanne K. Tobacman1,2 1Department of Medicine, University of Illinois at Chicago, Chicago, IL, USA 2Jesse Brown VAMC, Chicago, IL, USA 3Department of Chemistry and Chemical Biology Rensselaer Polytechnic Insitute, Troy, NY, USA Correspondence to: Joanne K. Tobacman, email: [email protected] Keywords: sulfotransferase; sulfatase; chondroitin sulfate; Wnt; EMT Received: February 18, 2020 Accepted: May 20, 2020 Published: June 16, 2020 Copyright: Bhattacharyya et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Expression of CHST15 (carbohydrate sulfotransferase 15; chondroitin 4-sulfate- 6-sulfotransferase; BRAG), the sulfotransferase enzyme that adds 6-sulfate to chondroitin 4-sulfate (C4S) to make chondroitin 4,6-disulfate (chondroitin sulfate E, CSE), was increased in malignant prostate epithelium obtained by laser capture microdissection and following arylsulfatase B (ARSB; N-acetylgalactosamine- 4-sulfatase) silencing in human prostate epithelial cells. Experiments in normal and malignant human prostate epithelial and stromal cells and tissues, in HepG2 cells, and in the ARSB-null mouse were performed to determine the pathway by which CHST15 expression is up-regulated when ARSB expression is reduced. Effects of Wnt- containing prostate stromal cell spent media and selective inhibitors of WNT, JNK, p38, SHP2, β-catenin, Rho, and Rac-1 signaling pathways were determined. -

Early Growth Response 1 Regulates Hematopoietic Support and Proliferation in Human Primary Bone Marrow Stromal Cells
Hematopoiesis SUPPLEMENTARY APPENDIX Early growth response 1 regulates hematopoietic support and proliferation in human primary bone marrow stromal cells Hongzhe Li, 1,2 Hooi-Ching Lim, 1,2 Dimitra Zacharaki, 1,2 Xiaojie Xian, 2,3 Keane J.G. Kenswil, 4 Sandro Bräunig, 1,2 Marc H.G.P. Raaijmakers, 4 Niels-Bjarne Woods, 2,3 Jenny Hansson, 1,2 and Stefan Scheding 1,2,5 1Division of Molecular Hematology, Department of Laboratory Medicine, Lund University, Lund, Sweden; 2Lund Stem Cell Center, Depart - ment of Laboratory Medicine, Lund University, Lund, Sweden; 3Division of Molecular Medicine and Gene Therapy, Department of Labora - tory Medicine, Lund University, Lund, Sweden; 4Department of Hematology, Erasmus MC Cancer Institute, Rotterdam, the Netherlands and 5Department of Hematology, Skåne University Hospital Lund, Skåne, Sweden ©2020 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2019.216648 Received: January 14, 2019. Accepted: July 19, 2019. Pre-published: August 1, 2019. Correspondence: STEFAN SCHEDING - [email protected] Li et al.: Supplemental data 1. Supplemental Materials and Methods BM-MNC isolation Bone marrow mononuclear cells (BM-MNC) from BM aspiration samples were isolated by density gradient centrifugation (LSM 1077 Lymphocyte, PAA, Pasching, Austria) either with or without prior incubation with RosetteSep Human Mesenchymal Stem Cell Enrichment Cocktail (STEMCELL Technologies, Vancouver, Canada) for lineage depletion (CD3, CD14, CD19, CD38, CD66b, glycophorin A). BM-MNCs from fetal long bones and adult hip bones were isolated as reported previously 1 by gently crushing bones (femora, tibiae, fibulae, humeri, radii and ulna) in PBS+0.5% FCS subsequent passing of the cell suspension through a 40-µm filter. -

2016 Mock Exam General Pathology Answer Sheet
Name___________________________ 2016 Mock Exam General Pathology 1. You have 1 HOUR to complete this 50-question multiple choice exam. 2. Write your name on all pages of the exam packet. 3. Use capital letters on the answer sheet. 4. For each question, select the ONE best answer and mark it on the answer sheet. 5. Credit will be given only for correct answers recorded on the answer sheet. 6. All questions for which more than one answer is marked will be recorded as incorrect. 7. No credit will be awarded or deducted for incorrect answers. 8. Turn in the entire exam packet when you are done. 2016 Mock Exam General Pathology Answer sheet 1. ______ 26. ______ 2. ______ 27. ______ 3. ______ 28. ______ 4. ______ 29. ______ 5. ______ 30. ______ 6. ______ 31. ______ 7. ______ 32. ______ 8. ______ 33. ______ 9. ______ 34. ______ 10. ______ 35. ______ 11. ______ 36. ______ 12. ______ 37. ______ 13. ______ 38. ______ 14. ______ 39. ______ 15. ______ 40. ______ 16. ______ 41. ______ 17. ______ 42. ______ 18. ______ 43. ______ 19. ______ 44. ______ 20. ______ 45. ______ 21. ______ 46. ______ 22. ______ 47. ______ 23. ______ 48. ______ 24. ______ 49. ______ 25. ______ 50. ______ ii 2016 Mock Exam Name___________________________ General Pathology 1. Which of the following toxins reaches its target cell using retrograde axonal transport? a. tetanospasmin b. botulinum toxin c. syntaxin d. SNAP-25 2. All of the following EXCEPT which are true regarding high mobility group box protein 1 (HMGB-1)? a. -
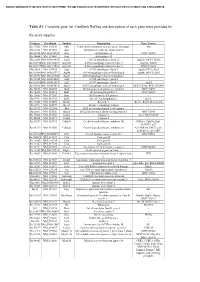
Table S1. Complete Gene List. Genbank Refseq and Description of Each Gene Were Provided By
Document downloaded from http://www.elsevier.es, day 24/09/2021. This copy is for personal use. Any transmission of this document by any media or format is strictly prohibited. Table S1. Complete gene list. GenBank RefSeq and description of each gene were provided by the array supplier. Unigene GeneBank Symbol Description Gene Name/s Rn.11422 NM_033230 Akt1 V-akt murine thymoma viral oncogene homolog 1 Akt Rn.2104 NM_019288 App Amyloid beta (A4) precursor protein - Rn.23323 NM_001034933 Arsa Arylsulfatase A MGC125207 Rn.94004 NM_033443 Arsb Arylsulfatase B - Rn.6224 NM_001038495 Atg12 ATG12 autophagy related 12 Apg12l, MGC125080 Rn.101734NM_001108809 Atg16l1 ATG16 autophagy related 16-like 1 Apg16l, Wdr30 Rn.104199NM_001191560 Atg16l2 ATG16 autophagy related 16-like 2 RGD1311400 Rn.3084 NM_134394 Atg3 ATG3 autophagy related 3 Apg3l, PIG-1, Pig1 Rn.163086NM_001025711 Atg4b ATG4 autophagy related 4 homolog B Apg4b, MGC112887 Rn.23378 NM_001107948 Atg4c ATG4 autophagy related 4 homolog C - Rn.98385 NM_001014250 Atg5 ATG5 autophagy related 5 - Rn.162765NM_001012097 Atg7 ATG7 autophagy related 7 Apg7l Rn.35248 NM_001014218 Atg9a ATG9 autophagy related 9 homolog A MGC105908, RGD1310450 Rn.36696 NM_022698 Bad BCL2-associated agonist of cell death MGC72439 Rn.14598 NM_053812 Bak1 BCL2-antagonist/killer 1 MGC108627 Rn.10668 NM_017059 Bax Bcl2-associated X protein - Rn.9996 NM_016993 Bcl2 B-cell CLL/lymphoma 2 Bcl-2 Rn.10323 NM_031535 Bcl2l1 Bcl2-like 1 Bcl-xl, Bcl2l, Bclx, bcl-X Rn.2776 NM_053739 Becn1 Beclin 1, autophagy related - Rn.31142 NM_022684