University of California, San Diego
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genetic Basis of Simple and Complex Traits with Relevance to Avian Evolution
Genetic basis of simple and complex traits with relevance to avian evolution Małgorzata Anna Gazda Doctoral Program in Biodiversity, Genetics and Evolution D Faculdade de Ciências da Universidade do Porto 2019 Supervisor Miguel Jorge Pinto Carneiro, Auxiliary Researcher, CIBIO/InBIO, Laboratório Associado, Universidade do Porto Co-supervisor Ricardo Lopes, CIBIO/InBIO Leif Andersson, Uppsala University FCUP Genetic basis of avian traits Nota Previa Na elaboração desta tese, e nos termos do número 2 do Artigo 4º do Regulamento Geral dos Terceiros Ciclos de Estudos da Universidade do Porto e do Artigo 31º do D.L.74/2006, de 24 de Março, com a nova redação introduzida pelo D.L. 230/2009, de 14 de Setembro, foi efetuado o aproveitamento total de um conjunto coerente de trabalhos de investigação já publicados ou submetidos para publicação em revistas internacionais indexadas e com arbitragem científica, os quais integram alguns dos capítulos da presente tese. Tendo em conta que os referidos trabalhos foram realizados com a colaboração de outros autores, o candidato esclarece que, em todos eles, participou ativamente na sua conceção, na obtenção, análise e discussão de resultados, bem como na elaboração da sua forma publicada. Este trabalho foi apoiado pela Fundação para a Ciência e Tecnologia (FCT) através da atribuição de uma bolsa de doutoramento (PD/BD/114042/2015) no âmbito do programa doutoral em Biodiversidade, Genética e Evolução (BIODIV). 2 FCUP Genetic basis of avian traits Acknowledgements Firstly, I would like to thank to my all supervisors Miguel Carneiro, Ricardo Lopes and Leif Andersson, for the demanding task of supervising myself last four years. -

Epigenetic Variation During the Adult Lifespan: Cross-Sectional and Longitudinal Data on Monozygotic Twin Pairs
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by DSpace at VU Aging Cell (2012) 11, pp694–703 Doi: 10.1111/j.1474-9726.2012.00835.x Epigenetic variation during the adult lifespan: cross-sectional and longitudinal data on monozygotic twin pairs Rudolf P. Talens,1 Kaare Christensen,2,3,4 Hein Putter,5 Introduction Gonneke Willemsen6, Lene Christiansen,2,3,4 Dennis The risk of most common diseases increases with age. A lifetime of accu- Kremer,1 H. Eka D. Suchiman,1 P. Eline Slagboom,1,7 6 1,7 mulated epigenetic changes was proposed to contribute to the develop- Dorret I. Boomsma and Bastiaan T. Heijmans ment of such diseases (Bjornsson et al., 2004). Epigenetic mechanisms 1Department of Molecular Epidemiology, Leiden University Medical Center, determine the expression potential of genes without changing the DNA Leiden, The Netherlands sequence (Jaenisch & Bird, 2003). The molecular basis includes the methyl- 2 The Danish Aging Research Center and The Danish Twin Registry, ation of cytosines in CpG dinucleotides, which, together with histone mod- University of Southern Denmark, Odense C, Denmark ifications, noncoding RNAs, and localization, influence the accessibility of 3Department of Clinical Genetics, Odense University Hospital, Odense C, Denmark a genomic locus to the transcriptional machinery (Bernstein et al., 2007; 4Department of Clinical Biochemistry and Pharmacology, Odense University Cedar & Bergman, 2009). DNA methylation can be measured on DNA sam- Hospital, Odense C, Denmark ples that are commonly available in biobanks (Talens et al., 2010). 5 Department of Medical Statistics and Bioinformatics, Leiden University Various studies have investigated whether DNA methylation can Medical Center, Leiden, The Netherlands change with increasing calendar age. -

A General Binding Mechanism for All Human Sulfatases by the Formylglycine-Generating Enzyme
A general binding mechanism for all human sulfatases by the formylglycine-generating enzyme Dirk Roeser*, Andrea Preusser-Kunze†, Bernhard Schmidt†, Kathrin Gasow*, Julia G. Wittmann*, Thomas Dierks‡, Kurt von Figura†, and Markus Georg Rudolph*§ *Department of Molecular Structural Biology, University of Go¨ttingen, Justus-von-Liebig-Weg 11, D-37077 Go¨ttingen, Germany; †Department of Biochemistry II, Heinrich-Du¨ker-Weg 12, University of Go¨ttingen, D-37073 Go¨ttingen, Germany; and ‡Department of Biochemistry I, Universita¨tsstrasse 25, University of Bielefeld, D-33615 Bielefeld, Germany Edited by Carolyn R. Bertozzi, University of California, Berkeley, CA, and approved November 8, 2005 (received for review September 1, 2005) The formylglycine (FGly)-generating enzyme (FGE) uses molecular tases, suggesting a general binding mechanism of substrate sulfa- oxygen to oxidize a conserved cysteine residue in all eukaryotic tases by FGE. sulfatases to the catalytically active FGly. Sulfatases degrade and The details of how O2-dependent cysteine oxidation is mediated remodel sulfate esters, and inactivity of FGE results in multiple by FGE are unknown. As a first step toward the elucidation of the sulfatase deficiency, a fatal disease. The previously determined FGE molecular mechanism of FGly formation, we have previously crystal structure revealed two crucial cysteine residues in the active determined crystal structures of FGE in various oxidation states site, one of which was thought to be implicated in substrate (8). FGE adopts a novel fold with surprisingly little regular sec- 2ϩ binding. The other cysteine residue partakes in a novel oxygenase ondary structure and contains two structural Ca ions and two mechanism that does not rely on any cofactors. -

Supplementary File 2A Revised
Supplementary file 2A. Differentially expressed genes in aldosteronomas compared to all other samples, ranked according to statistical significance. Missing values were not allowed in aldosteronomas, but to a maximum of five in the other samples. Acc UGCluster Name Symbol log Fold Change P - Value Adj. P-Value B R99527 Hs.8162 Hypothetical protein MGC39372 MGC39372 2,17 6,3E-09 5,1E-05 10,2 AA398335 Hs.10414 Kelch domain containing 8A KLHDC8A 2,26 1,2E-08 5,1E-05 9,56 AA441933 Hs.519075 Leiomodin 1 (smooth muscle) LMOD1 2,33 1,3E-08 5,1E-05 9,54 AA630120 Hs.78781 Vascular endothelial growth factor B VEGFB 1,24 1,1E-07 2,9E-04 7,59 R07846 Data not found 3,71 1,2E-07 2,9E-04 7,49 W92795 Hs.434386 Hypothetical protein LOC201229 LOC201229 1,55 2,0E-07 4,0E-04 7,03 AA454564 Hs.323396 Family with sequence similarity 54, member B FAM54B 1,25 3,0E-07 5,2E-04 6,65 AA775249 Hs.513633 G protein-coupled receptor 56 GPR56 -1,63 4,3E-07 6,4E-04 6,33 AA012822 Hs.713814 Oxysterol bining protein OSBP 1,35 5,3E-07 7,1E-04 6,14 R45592 Hs.655271 Regulating synaptic membrane exocytosis 2 RIMS2 2,51 5,9E-07 7,1E-04 6,04 AA282936 Hs.240 M-phase phosphoprotein 1 MPHOSPH -1,40 8,1E-07 8,9E-04 5,74 N34945 Hs.234898 Acetyl-Coenzyme A carboxylase beta ACACB 0,87 9,7E-07 9,8E-04 5,58 R07322 Hs.464137 Acyl-Coenzyme A oxidase 1, palmitoyl ACOX1 0,82 1,3E-06 1,2E-03 5,35 R77144 Hs.488835 Transmembrane protein 120A TMEM120A 1,55 1,7E-06 1,4E-03 5,07 H68542 Hs.420009 Transcribed locus 1,07 1,7E-06 1,4E-03 5,06 AA410184 Hs.696454 PBX/knotted 1 homeobox 2 PKNOX2 1,78 2,0E-06 -

Lysosomal Hydrolases: Conversion of Acidic to Basic Forms by Neuraminidase
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Volume 13, number 1 FEBS LETTERS February 1971 LYSOSOMAL HYDROLASES: CONVERSION OF ACIDIC TO BASIC FORMS BY NEURAMINIDASE Alfred GOLDSTONE, Philip KONECNY and Harold KOENIG Neurology Service, VA Research Hospital; and Departments of Neurology and Biochemistry, Northwestern University Medical School, Chicago, Illinois, USA Received 21 December 1970 1. Introduction intervals to avoid overheating). The lysosomal lysate was ultracentrifuged at 100,000 g for 30 min to give A number of hydrolytic enzymes of lysosomal a soluble enzyme fraction. This fraction also contains origin occur in multiple forms which differ in such an acidic lipoprotein [4]. The insoluble residue was physicochemical properties as solubility, heat and pH suspended in 0.05 M acetate buffer pH 5.0 with 0.2% stability, pH optima and electrophoretic mobility [I]. Triton X-100 and resonicated to release the bound ly- Recent studies in this laboratory indicate that essen- sosomal enzymes together with additional acidic lipo- tially all the enzyme proteins of rat kidney and liver protein ([4] and A. Goldstone and H. Koenig, unpu- lysosomes are glycoproteins, and that at least some of blished observations). All operations were carried out these glycoproteins contain sialic acid [2-41. The at 4’. acidic form of lysosomal N-acetylhexosaminidase from Aliquots of the soluble fraction containing about human spleen [ 51 and kidney [6] is converted into 2 mg of protein were incubated with neuraminidase the basic form by treatment with neuraminidase. We from Cl. perfringens (Type Vl enzyme supplied by now confirm this finding for rat kidney and show Sigma), 0.1 mg/ml, or V. -

Increased CHST15 Follows Decline in Arylsulfatase B (ARSB) and Disinhibition of Non-Canonical WNT Signaling: Potential Impact on Epithelial and Mesenchymal Identity
www.oncotarget.com Oncotarget, 2020, Vol. 11, (No. 24), pp: 2327-2344 Research Paper Increased CHST15 follows decline in arylsulfatase B (ARSB) and disinhibition of non-canonical WNT signaling: potential impact on epithelial and mesenchymal identity Sumit Bhattacharyya1,2, Leo Feferman1,2, Xiaorui Han3, Ke Xia3, Fuming Zhang3, Robert J. Linhardt3, Joanne K. Tobacman1,2 1Department of Medicine, University of Illinois at Chicago, Chicago, IL, USA 2Jesse Brown VAMC, Chicago, IL, USA 3Department of Chemistry and Chemical Biology Rensselaer Polytechnic Insitute, Troy, NY, USA Correspondence to: Joanne K. Tobacman, email: [email protected] Keywords: sulfotransferase; sulfatase; chondroitin sulfate; Wnt; EMT Received: February 18, 2020 Accepted: May 20, 2020 Published: June 16, 2020 Copyright: Bhattacharyya et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Expression of CHST15 (carbohydrate sulfotransferase 15; chondroitin 4-sulfate- 6-sulfotransferase; BRAG), the sulfotransferase enzyme that adds 6-sulfate to chondroitin 4-sulfate (C4S) to make chondroitin 4,6-disulfate (chondroitin sulfate E, CSE), was increased in malignant prostate epithelium obtained by laser capture microdissection and following arylsulfatase B (ARSB; N-acetylgalactosamine- 4-sulfatase) silencing in human prostate epithelial cells. Experiments in normal and malignant human prostate epithelial and stromal cells and tissues, in HepG2 cells, and in the ARSB-null mouse were performed to determine the pathway by which CHST15 expression is up-regulated when ARSB expression is reduced. Effects of Wnt- containing prostate stromal cell spent media and selective inhibitors of WNT, JNK, p38, SHP2, β-catenin, Rho, and Rac-1 signaling pathways were determined. -

Early Growth Response 1 Regulates Hematopoietic Support and Proliferation in Human Primary Bone Marrow Stromal Cells
Hematopoiesis SUPPLEMENTARY APPENDIX Early growth response 1 regulates hematopoietic support and proliferation in human primary bone marrow stromal cells Hongzhe Li, 1,2 Hooi-Ching Lim, 1,2 Dimitra Zacharaki, 1,2 Xiaojie Xian, 2,3 Keane J.G. Kenswil, 4 Sandro Bräunig, 1,2 Marc H.G.P. Raaijmakers, 4 Niels-Bjarne Woods, 2,3 Jenny Hansson, 1,2 and Stefan Scheding 1,2,5 1Division of Molecular Hematology, Department of Laboratory Medicine, Lund University, Lund, Sweden; 2Lund Stem Cell Center, Depart - ment of Laboratory Medicine, Lund University, Lund, Sweden; 3Division of Molecular Medicine and Gene Therapy, Department of Labora - tory Medicine, Lund University, Lund, Sweden; 4Department of Hematology, Erasmus MC Cancer Institute, Rotterdam, the Netherlands and 5Department of Hematology, Skåne University Hospital Lund, Skåne, Sweden ©2020 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2019.216648 Received: January 14, 2019. Accepted: July 19, 2019. Pre-published: August 1, 2019. Correspondence: STEFAN SCHEDING - [email protected] Li et al.: Supplemental data 1. Supplemental Materials and Methods BM-MNC isolation Bone marrow mononuclear cells (BM-MNC) from BM aspiration samples were isolated by density gradient centrifugation (LSM 1077 Lymphocyte, PAA, Pasching, Austria) either with or without prior incubation with RosetteSep Human Mesenchymal Stem Cell Enrichment Cocktail (STEMCELL Technologies, Vancouver, Canada) for lineage depletion (CD3, CD14, CD19, CD38, CD66b, glycophorin A). BM-MNCs from fetal long bones and adult hip bones were isolated as reported previously 1 by gently crushing bones (femora, tibiae, fibulae, humeri, radii and ulna) in PBS+0.5% FCS subsequent passing of the cell suspension through a 40-µm filter. -
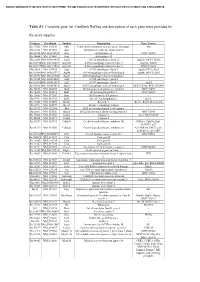
Table S1. Complete Gene List. Genbank Refseq and Description of Each Gene Were Provided By
Document downloaded from http://www.elsevier.es, day 24/09/2021. This copy is for personal use. Any transmission of this document by any media or format is strictly prohibited. Table S1. Complete gene list. GenBank RefSeq and description of each gene were provided by the array supplier. Unigene GeneBank Symbol Description Gene Name/s Rn.11422 NM_033230 Akt1 V-akt murine thymoma viral oncogene homolog 1 Akt Rn.2104 NM_019288 App Amyloid beta (A4) precursor protein - Rn.23323 NM_001034933 Arsa Arylsulfatase A MGC125207 Rn.94004 NM_033443 Arsb Arylsulfatase B - Rn.6224 NM_001038495 Atg12 ATG12 autophagy related 12 Apg12l, MGC125080 Rn.101734NM_001108809 Atg16l1 ATG16 autophagy related 16-like 1 Apg16l, Wdr30 Rn.104199NM_001191560 Atg16l2 ATG16 autophagy related 16-like 2 RGD1311400 Rn.3084 NM_134394 Atg3 ATG3 autophagy related 3 Apg3l, PIG-1, Pig1 Rn.163086NM_001025711 Atg4b ATG4 autophagy related 4 homolog B Apg4b, MGC112887 Rn.23378 NM_001107948 Atg4c ATG4 autophagy related 4 homolog C - Rn.98385 NM_001014250 Atg5 ATG5 autophagy related 5 - Rn.162765NM_001012097 Atg7 ATG7 autophagy related 7 Apg7l Rn.35248 NM_001014218 Atg9a ATG9 autophagy related 9 homolog A MGC105908, RGD1310450 Rn.36696 NM_022698 Bad BCL2-associated agonist of cell death MGC72439 Rn.14598 NM_053812 Bak1 BCL2-antagonist/killer 1 MGC108627 Rn.10668 NM_017059 Bax Bcl2-associated X protein - Rn.9996 NM_016993 Bcl2 B-cell CLL/lymphoma 2 Bcl-2 Rn.10323 NM_031535 Bcl2l1 Bcl2-like 1 Bcl-xl, Bcl2l, Bclx, bcl-X Rn.2776 NM_053739 Becn1 Beclin 1, autophagy related - Rn.31142 NM_022684 -

A Possible Role for Arylsulfatase G in Dermatan Sulfate Metabolism
International Journal of Molecular Sciences Article A Possible Role for Arylsulfatase G in Dermatan Sulfate Metabolism Aleksandra Poterala-Hejmo 1,*, Adam Golda 2 , Marcin Pacholczyk 1 , Sebastian Student 1, Anna Tylki-Szyma´nska 3 and Anna Lalik 1,* 1 Department of Systems Biology and Engineering, Silesian University of Technology, 44-100 Gliwice, Poland; [email protected] (M.P.); [email protected] (S.S.) 2 Department of Cardiology, 4th Municipal Hospital, 44-100 Gliwice, Poland; [email protected] 3 Department of Pediatrics, Nutrition and Metabolic Diseases, The Children’s Memorial Health Institute, 04-730 Warsaw, Poland; [email protected] * Correspondence: [email protected] (A.P.-H.); [email protected] (A.L.); Tel.: +48-32-2371168 (A.P.-H.); +48-32-2372769 (A.L.) Received: 2 June 2020; Accepted: 6 July 2020; Published: 12 July 2020 Abstract: Perturbations of glycosaminoglycan metabolism lead to mucopolysaccharidoses (MPS)—lysosomal storage diseases. One type of MPS (type VI) is associated with a deficiency of arylsulfatase B (ARSB), for which we previously established a cellular model using pulmonary artery endothelial cells with a silenced ARSB gene. Here, we explored the effects of silencing the ARSB gene on the growth of human pulmonary artery smooth muscle cells in the presence of different concentrations of dermatan sulfate (DS). The viability of pulmonary artery smooth muscle cells with a silenced ARSB gene was stimulated by the dermatan sulfate. In contrast, the growth of pulmonary artery endothelial cells was not affected. As shown by microarray analysis, the expression of the arylsulfatase G (ARSG) in pulmonary artery smooth muscle cells increased after silencing the arylsulfatase B gene, but the expression of genes encoding other enzymes involved in the degradation of dermatan sulfate did not. -

Recombinant Human Arylsulfatase B/ARSB
Recombinant Human Arylsulfatase B/ARSB Catalog Number: 4415-SU DESCRIPTION Source Chinese Hamster Ovary cell line, CHOderived Ser37Met533, with a Cterminal 10His tag Accession # P15848 Nterminal Sequence Ser37 Analysis Predicted Molecular 57 kDa Mass SPECIFICATIONS SDSPAGE 80 kDa, reducing conditions Activity Measured by its ability to hydrolyze the substrate 4Nitrocatechol Sulfate (PNCS). The specific activity is >3,000 pmol/min/µg, as measured under the described conditions. Endotoxin Level <1.0 EU per 1 μg of the protein by the LAL method. Purity >95%, by SDSPAGE under reducing conditions and visualized by silver stain. Formulation Supplied as a 0.2 μm filtered solution in Tris and NaCl. See Certificate of Analysis for details. Activity Assay Protocol Materials l Assay Buffer: 50 mM MES, pH 6.5 l Recombinant Human Arylsulfatase B/ARSB (rhARSB) (Catalog # 4415SU) l Substrate: 4Nitrocatechol Sulfate (4PNCS) (Sigma, Catalog # N7251) l NaOH (Sigma, Catalog # S0899) l 96well Clear Plate (Costar, Catalog # 92592) l Plate Reader (Model: SpectraMax Plus by Molecular Devices) or equivalent Assay 1. Dilute rhARSB to 1 µg/mL in Assay Buffer. 2. Dilute Substrate to 2 mM in Assay Buffer. 3. Combine 75 µL of 1 µg/mL rhARSB and 75 µL of 2 mM Substrate. Include a Substrate Blank containing 75 µL Assay Buffer and 75 µL Substrate. 4. Incubate at 37 °C for 1 hour. 5. Stop reaction by adding 150 µL of 0.2 M NaOH. 6. Load 200 µL of reaction into a plate. 7. Read at 510 nm (absorbance) in endpoint mode. -

A Genome-Wide Analysis of DNA Methylation in Buccal Cells
Genes 2014, 5, 347-365; doi:10.3390/genes5020347 OPEN ACCESS genes ISSN 2073-4425 www.mdpi.com/journal/genes Article Epigenetic Variation in Monozygotic Twins: A Genome-Wide Analysis of DNA Methylation in Buccal Cells Jenny van Dongen 1,*, Erik A. Ehli 2,3, Roderick C. Slieker 4, Meike Bartels 1, Zachary M. Weber 2, Gareth E. Davies 2,3, P. Eline Slagboom 4, Bastiaan T. Heijmans 4 and Dorret I. Boomsma 1 1 Department of Biological Psychology, VU University Amsterdam, Van der Boechorststraat 1, 1081 BT Amsterdam, The Netherlands; E-Mails: [email protected] (M.B.); [email protected] (D.I.B.) 2 Avera Institute for Human Genetics, 3720 W. 69th Street, Sioux Falls, SD 57108, USA; E-Mails: [email protected] (E.A.E.); [email protected] (Z.M.W.); [email protected] (G.E.D.) 3 Department of Psychiatry, University of South Dakota, 4400 W. 69th Street, Sioux Falls, SD 57108, USA 4 Department of Molecular Epidemiology, Leiden University Medical Center, P.O. Box 9600, 2300 RC Leiden, The Netherlands; E-Mails: [email protected] (R.C.S.); [email protected] (P.E.S.); [email protected] (B.T.H.) * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +31-20-598-3570; Fax: +31-20-598-8832. Received: 7 February 2014; in revised form: 31 March 2014 / Accepted: 16 April 2014 / Published: 5 May 2014 Abstract: DNA methylation is one of the most extensively studied epigenetic marks in humans. -

Genetic, Epidemiological, Biomarker, and 'Omics'
Twin Research and Human Genetics Volume 21 Number 3 pp. 239–252 C The Author(s) 2018. This is an Open Access article, distributed under the terms of the Creative Commons Attribution licence (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted re-use, distribution, and reproduction in any medium, provided the original work is properly cited. doi:10.1017/thg.2018.23 Establishing a Twin Register: An Invaluable Resource for (Behavior) Genetic, Epidemiological, Biomarker, and ‘Omics’ Studies Veronika V. Odintsova,1,2 Gonneke Willemsen,1 Conor V. Dolan,1 Jouke-Jan Hottenga,1 Nicholas G. Martin,3 P. Eline Slagboom,4 Juan R. Ordoñana,5,∗ and Dorret I. Boomsma1,∗ 1Netherlands Twin Register, Department of Biological Psychology, Vrije Universiteit Amsterdam, and Amsterdam Public Health (APH), Amsterdam, the Netherlands 2National Medical Research Center for Obstetrics, Gynecology and Perinatology, Ministry of Healthcare of the Russian Federation, Moscow, Russia 3Genetic Epidemiology, QIMR Berghofer Medical Research Institute, Brisbane, Australia 4Department of Molecular Epidemiology, Leiden University Medical Centre, Leiden, the Netherlands 5Department of Human Anatomy and Psychobiology, University of Murcia, and Murcia Institute for BioHealth Research (IMIB-Arrixaca-UMU), Murcia, Spain Twin registers are wonderful research resources for research applications in medical and behavioral genet- ics, epidemiology, psychology, molecular genetics, and other areas of research. New registers continue to be launched all over the world as researchers from different disciplines recognize the potential to boost and widen their research agenda. In this article, we discuss multiple aspects that need to be taken into account when initiating a register, from its preliminary sketch to its actual development.