The Response of C1 and C13 Cysteine Proteases in Soybean Nodules to Drought
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Babela Massiliensis, a Representative of a Widespread Bacterial
Babela massiliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae Isabelle Pagnier, Natalya Yutin, Olivier Croce, Kira S Makarova, Yuri I Wolf, Samia Benamar, Didier Raoult, Eugene V. Koonin, Bernard La Scola To cite this version: Isabelle Pagnier, Natalya Yutin, Olivier Croce, Kira S Makarova, Yuri I Wolf, et al.. Babela mas- siliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae. Biology Direct, BioMed Central, 2015, 10 (13), 10.1186/s13062-015-0043-z. hal-01217089 HAL Id: hal-01217089 https://hal-amu.archives-ouvertes.fr/hal-01217089 Submitted on 19 Oct 2015 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Pagnier et al. Biology Direct (2015) 10:13 DOI 10.1186/s13062-015-0043-z RESEARCH Open Access Babela massiliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae Isabelle Pagnier1, Natalya Yutin2, Olivier Croce1, Kira S Makarova2, Yuri I Wolf2, Samia Benamar1, Didier Raoult1, Eugene V Koonin2 and Bernard La Scola1* Abstract Background: Only a small fraction of bacteria and archaea that are identifiable by metagenomics can be grown on standard media. -

Enzymatic Encoding Methods for Efficient Synthesis Of
(19) TZZ__T (11) EP 1 957 644 B1 (12) EUROPEAN PATENT SPECIFICATION (45) Date of publication and mention (51) Int Cl.: of the grant of the patent: C12N 15/10 (2006.01) C12Q 1/68 (2006.01) 01.12.2010 Bulletin 2010/48 C40B 40/06 (2006.01) C40B 50/06 (2006.01) (21) Application number: 06818144.5 (86) International application number: PCT/DK2006/000685 (22) Date of filing: 01.12.2006 (87) International publication number: WO 2007/062664 (07.06.2007 Gazette 2007/23) (54) ENZYMATIC ENCODING METHODS FOR EFFICIENT SYNTHESIS OF LARGE LIBRARIES ENZYMVERMITTELNDE KODIERUNGSMETHODEN FÜR EINE EFFIZIENTE SYNTHESE VON GROSSEN BIBLIOTHEKEN PROCEDES DE CODAGE ENZYMATIQUE DESTINES A LA SYNTHESE EFFICACE DE BIBLIOTHEQUES IMPORTANTES (84) Designated Contracting States: • GOLDBECH, Anne AT BE BG CH CY CZ DE DK EE ES FI FR GB GR DK-2200 Copenhagen N (DK) HU IE IS IT LI LT LU LV MC NL PL PT RO SE SI • DE LEON, Daen SK TR DK-2300 Copenhagen S (DK) Designated Extension States: • KALDOR, Ditte Kievsmose AL BA HR MK RS DK-2880 Bagsvaerd (DK) • SLØK, Frank Abilgaard (30) Priority: 01.12.2005 DK 200501704 DK-3450 Allerød (DK) 02.12.2005 US 741490 P • HUSEMOEN, Birgitte Nystrup DK-2500 Valby (DK) (43) Date of publication of application: • DOLBERG, Johannes 20.08.2008 Bulletin 2008/34 DK-1674 Copenhagen V (DK) • JENSEN, Kim Birkebæk (73) Proprietor: Nuevolution A/S DK-2610 Rødovre (DK) 2100 Copenhagen 0 (DK) • PETERSEN, Lene DK-2100 Copenhagen Ø (DK) (72) Inventors: • NØRREGAARD-MADSEN, Mads • FRANCH, Thomas DK-3460 Birkerød (DK) DK-3070 Snekkersten (DK) • GODSKESEN, -

B Number Gene Name Mrna Intensity Mrna
sample) total list predicted B number Gene name assignment mRNA present mRNA intensity Gene description Protein detected - Membrane protein membrane sample detected (total list) Proteins detected - Functional category # of tryptic peptides # of tryptic peptides # of tryptic peptides detected (membrane b0002 thrA 13624 P 39 P 18 P(m) 2 aspartokinase I, homoserine dehydrogenase I Metabolism of small molecules b0003 thrB 6781 P 9 P 3 0 homoserine kinase Metabolism of small molecules b0004 thrC 15039 P 18 P 10 0 threonine synthase Metabolism of small molecules b0008 talB 20561 P 20 P 13 0 transaldolase B Metabolism of small molecules chaperone Hsp70; DNA biosynthesis; autoregulated heat shock b0014 dnaK 13283 P 32 P 23 0 proteins Cell processes b0015 dnaJ 4492 P 13 P 4 P(m) 1 chaperone with DnaK; heat shock protein Cell processes b0029 lytB 1331 P 16 P 2 0 control of stringent response; involved in penicillin tolerance Global functions b0032 carA 9312 P 14 P 8 0 carbamoyl-phosphate synthetase, glutamine (small) subunit Metabolism of small molecules b0033 carB 7656 P 48 P 17 0 carbamoyl-phosphate synthase large subunit Metabolism of small molecules b0048 folA 1588 P 7 P 1 0 dihydrofolate reductase type I; trimethoprim resistance Metabolism of small molecules peptidyl-prolyl cis-trans isomerase (PPIase), involved in maturation of b0053 surA 3825 P 19 P 4 P(m) 1 GenProt outer membrane proteins (1st module) Cell processes b0054 imp 2737 P 42 P 5 P(m) 5 GenProt organic solvent tolerance Cell processes b0071 leuD 4770 P 10 P 9 0 isopropylmalate -

Product Sheet Info
Master Clone List for NR-19279 ® Vibrio cholerae Gateway Clone Set, Recombinant in Escherichia coli, Plates 1-46 Catalog No. NR-19279 Table 1: Vibrio cholerae Gateway® Clones, Plate 1 (NR-19679) Clone ID Well ORF Locus ID Symbol Product Accession Position Length Number 174071 A02 367 VC2271 ribD riboflavin-specific deaminase NP_231902.1 174346 A03 336 VC1877 lpxK tetraacyldisaccharide 4`-kinase NP_231511.1 174354 A04 342 VC0953 holA DNA polymerase III, delta subunit NP_230600.1 174115 A05 388 VC2085 sucC succinyl-CoA synthase, beta subunit NP_231717.1 174310 A06 506 VC2400 murC UDP-N-acetylmuramate--alanine ligase NP_232030.1 174523 A07 132 VC0644 rbfA ribosome-binding factor A NP_230293.2 174632 A08 322 VC0681 ribF riboflavin kinase-FMN adenylyltransferase NP_230330.1 174930 A09 433 VC0720 phoR histidine protein kinase PhoR NP_230369.1 174953 A10 206 VC1178 conserved hypothetical protein NP_230823.1 174976 A11 213 VC2358 hypothetical protein NP_231988.1 174898 A12 369 VC0154 trmA tRNA (uracil-5-)-methyltransferase NP_229811.1 174059 B01 73 VC2098 hypothetical protein NP_231730.1 174075 B02 82 VC0561 rpsP ribosomal protein S16 NP_230212.1 174087 B03 378 VC1843 cydB-1 cytochrome d ubiquinol oxidase, subunit II NP_231477.1 174099 B04 383 VC1798 eha eha protein NP_231433.1 174294 B05 494 VC0763 GTP-binding protein NP_230412.1 174311 B06 314 VC2183 prsA ribose-phosphate pyrophosphokinase NP_231814.1 174603 B07 108 VC0675 thyA thymidylate synthase NP_230324.1 174474 B08 466 VC1297 asnS asparaginyl-tRNA synthetase NP_230942.2 174933 B09 198 -

Downloading the Nucleotide Sequences and Scanning Them Against the Database
An in silico analysis, purification and partial kinetic characterisation of a serine protease from Gelidium pristoides A dissertation submitted in fulfilment of the requirement for the degree of Master of Science (MSc) Biochemistry by Zolani Ntsata Supervisor: Prof. Graeme Bradley 2020 Department of Biochemistry and Microbiology Declaration I, Zolani Ntsata (201106067), declare that this dissertation, entitled ‘An in silico analysis and kinetic characterisation of proteases from red algae’ submitted to the University of Fort Hare for the Master’s degree (Biochemistry) award, is my original work and has NOT been submitted to any other university. Signature: __________________ I, Zolani Ntsata (201106067), declare that I am highly cognisant of the University of Fort Hare policy on plagiarism and I have been careful to comply with these regulations. Furthermore, all the sources of information have been cited as indicated in the bibliography. Signature: __________________ I, Zolani Ntsata (201106067), declare that I am fully aware of the University of Fort Hare’s policy on research ethics, and I have taken every precaution to comply with these regulations. There was no need for ethical clearance. Signature: _________________ i Dedication I dedicate this work to my grandmother, Nyameka Mabi. ii Acknowledgements Above all things, I would like to give thanks to God for the opportunity to do this project and for the extraordinary strength to persevere in spite of the challenges that came along. I am thankful to my family, especially my grandmother, for her endless support. I would also like to acknowledge Prof Graeme Bradley for his supervision and guidance. Thanks to my friends and colleagues, especially Yanga Gogela and Ntombekhaya Nqumla, and the plant stress group for their help and support. -

WO 2013/185113 Al 12 December 2013 (12.12.2013) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2013/185113 Al 12 December 2013 (12.12.2013) P O P C T (51) International Patent Classification: (81) Designated States (unless otherwise indicated, for every A61K 38/02 (2006.01) A61P 7/02 (2006.01) kind of national protection available): AE, AG, AL, AM, A61K 38/36 (2006.01) AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, (21) Number: International Application DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, PCT/US20 13/044841 HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KN, KP, KR, (22) International Filing Date: KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, ME, 7 June 2013 (07.06.2013) MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SC, (25) Filing Language: English SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, (26) Publication Language: English TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (30) Priority Data: (84) Designated States (unless otherwise indicated, for every 61/657,688 8 June 2012 (08.06.2012) kind of regional protection available): ARIPO (BW, GH, 61/800,626 15 March 2013 (15.03.2013) GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, SZ, TZ, UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, TJ, (71) Applicant: BIOGEN IDEC MA INC. -

Next Generation Sequencing Identifies Five Major Classes of Potentially Therapeutic Enzymes Secreted by Lucilia Sericata Medical Maggots
Hindawi Publishing Corporation BioMed Research International Volume 2016, Article ID 8285428, 27 pages http://dx.doi.org/10.1155/2016/8285428 Research Article Next Generation Sequencing Identifies Five Major Classes of Potentially Therapeutic Enzymes Secreted by Lucilia sericata Medical Maggots Zdenjk Franta,1 Heiko Vogel,2 Rüdiger Lehmann,1 Oliver Rupp,3 Alexander Goesmann,3 and Andreas Vilcinskas1,4 1 Department of Bioresources, Fraunhofer Institute for Molecular Biology and Applied Ecology, Winchesterstraße 2, 35394 Giessen, Germany 2Department of Entomology, Max Planck Institute for Chemical Ecology, Hans-Knoll-Straße¨ 8, 07745 Jena, Germany 3Justus-Liebig-University of Giessen, Bioinformatics and System Biology, Heinrich-Buff-Ring 58, 35392 Giessen, Germany 4Justus-Liebig-University of Giessen, Institute for Insect Biotechnology, Heinrich-Buff-Ring 26-32, 35392 Giessen, Germany Correspondence should be addressed to Andreas Vilcinskas; [email protected] Received 29 January 2016; Accepted 7 March 2016 Academic Editor: Yudong Cai Copyright © 2016 Zdenekˇ Franta et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Lucilia sericata larvae are used as an alternative treatment for recalcitrant and chronic wounds. Their excretions/secretions contain molecules that facilitate tissue debridement, disinfect, or accelerate wound healing and have therefore -

Galhenage Thanusha Ranithri Abeynayake
Development of hydrolysates with antioxidant effects from brewers’ spent grain proteins by Galhenage Thanusha Ranithri Abeynayake A thesis submitted in partial fulfillment of the requirements for the degree of Master of Science in Food Science and Technology Department of Agricultural, Food and Nutritional Science University of Alberta © Galhenage Thanusha Ranithri Abeynayake, 2020 Abstract Brewers’ spent grain (BSG), the most abundant brewing by-product contains up to 24% (w/w) of protein on dry basis. Despite of a rich protein source, it is still mainly used as low value animal feed. This study was conducted to develop hydrolysates with antioxidant effects from BSG proteins by enzymatic hydrolysis, and study BSG peptide antioxidant activities in relation to molecular structures. The BSG proteins were extracted by alkaline method with a protein content of 62.6% (w/w) and a protein recovery of 46.3%. The proteins were then hydrolyzed by selected proteases or protease combinations having different specificities. Resulting hydrolysates were analyzed for their molecular structures such as hydrolysis degree, amino acid composition, molecular weight and conformation, and their antioxidant effects were measured by reducing power, DPPH radical scavenging, ferrous chelating and superoxide radical scavenging assays. Hydrolysates prepared by alcalase and alcalase combined treatments with neutrase, flavourzyme and everlase showed the highest DPPH radical scavenging activities ranged between 72.6-74.9% at the hydrolysate concentration of 1.0 mg/mL. It could be due to specific activity of alcalase to produce amino acid sequences with greater DPPH radical scavenging potential. Similarly, high proton donation ability of the imidazole ring present in the side chain of histidine may have strong ability to scavenge the DPPH radical. -

SUMO Protease 1 (GST-Tagged) from Yeast, Recombinant Cat
SUMO Protease 1 (GST-tagged) from Yeast, Recombinant Cat. No. NATE-1708 Lot. No. (See product label) Introduction Description SUMO (Small Ubiquitin-like MOdifiers) Protease 1 (Ulp1, Ubl-specific protease 1 from Saccharomyces cerevisiae) is a highly active cysteine protease. It is highly specific as it recognizes the tertiary structure of the ubiquitin-like (UBL) protein, SUMO (Smt3), rather than its amino acid sequence. SUMO fusion tag, as an N-terminal fusion partner, has been shown to enhance functional protein production in prokaryotic and eukaryotic expression systems with significantly improved protein stability and solubility. The SUMO protease 1 can be used to cleave SUMO protein tag from recombinant SUMO-fusion proteins. The optimal temperature for cleavage is 30°C; however, the enzyme is active over wide ranges of temperature and pH. After the completion of the cleavage reaction, the protease can be easily removed from the reaction by affinity chromatography using the Glutathione resin. Synonyms Ulp1 peptidase; SUMO Protease; SUMO Protease Product Information Species Yeast Source E. coli Form Liquid EC Number EC 3.4.22.68 Molecular Weight 52.6 kDa (403-621 aa + N-terminal GST). Purity > 90% by SDS-PAGE Activity >10,000 units/mg Concentration 1 mg/ml Unit Definition One unit is defined as the amount of SUMO Protease 1 required to cleave >90% of 5 µg a control protein substrate (SUMO-GFP) in 1 h at 37°C. Creative Enzymes. All rights reserved. 45-1 Ramsey Road, Shirley, NY 11967, USA Tel:1-631-562-8517 1-516-512-3133 Fax:1-631-938-8127 E-mail: [email protected] http://www.creative-enzymes.com Notes INTENDED FOR RESEARCH USE ONLY, NOT FOR USE IN HUMAN, THERAPEUTIC OR DIAGNOSTIC APPLICATIONS. -
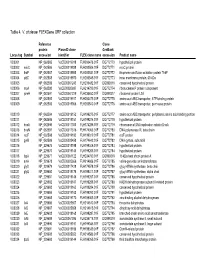
Table 4. V. Cholerae Flexgene ORF Collection
Table 4. V. cholerae FLEXGene ORF collection Reference Clone protein PlasmID clone GenBank Locus tag Symbol accession identifier FLEX clone name accession Product name VC0001 NP_062585 VcCD00019918 FLH200476.01F DQ772770 hypothetical protein VC0002 mioC NP_062586 VcCD00019938 FLH200506.01F DQ772771 mioC protein VC0003 thdF NP_062587 VcCD00019958 FLH200531.01F DQ772772 thiophene and furan oxidation protein ThdF VC0004 yidC NP_062588 VcCD00019970 FLH200545.01F DQ772773 inner membrane protein, 60 kDa VC0005 NP_062589 VcCD00061243 FLH236482.01F DQ899316 conserved hypothetical protein VC0006 rnpA NP_062590 VcCD00025697 FLH214799.01F DQ772774 ribonuclease P protein component VC0007 rpmH NP_062591 VcCD00061229 FLH236450.01F DQ899317 ribosomal protein L34 VC0008 NP_062592 VcCD00019917 FLH200475.01F DQ772775 amino acid ABC transporter, ATP-binding protein VC0009 NP_062593 VcCD00019966 FLH200540.01F DQ772776 amino acid ABC transproter, permease protein VC0010 NP_062594 VcCD00019152 FLH199275.01F DQ772777 amino acid ABC transporter, periplasmic amino acid-binding portion VC0011 NP_062595 VcCD00019151 FLH199274.01F DQ772778 hypothetical protein VC0012 dnaA NP_062596 VcCD00017363 FLH174286.01F DQ772779 chromosomal DNA replication initiator DnaA VC0013 dnaN NP_062597 VcCD00017316 FLH174063.01F DQ772780 DNA polymerase III, beta chain VC0014 recF NP_062598 VcCD00019182 FLH199319.01F DQ772781 recF protein VC0015 gyrB NP_062599 VcCD00025458 FLH174642.01F DQ772782 DNA gyrase, subunit B VC0016 NP_229675 VcCD00019198 FLH199346.01F DQ772783 hypothetical protein -

Biochemical Investigation of the Ubiquitin Carboxyl-Terminal Hydrolase Family" (2015)
Purdue University Purdue e-Pubs Open Access Dissertations Theses and Dissertations Spring 2015 Biochemical investigation of the ubiquitin carboxyl- terminal hydrolase family Joseph Rashon Chaney Purdue University Follow this and additional works at: https://docs.lib.purdue.edu/open_access_dissertations Part of the Biochemistry Commons, Biophysics Commons, and the Molecular Biology Commons Recommended Citation Chaney, Joseph Rashon, "Biochemical investigation of the ubiquitin carboxyl-terminal hydrolase family" (2015). Open Access Dissertations. 430. https://docs.lib.purdue.edu/open_access_dissertations/430 This document has been made available through Purdue e-Pubs, a service of the Purdue University Libraries. Please contact [email protected] for additional information. *UDGXDWH6FKRRO)RUP 8SGDWHG PURDUE UNIVERSITY GRADUATE SCHOOL Thesis/Dissertation Acceptance 7KLVLVWRFHUWLI\WKDWWKHWKHVLVGLVVHUWDWLRQSUHSDUHG %\ Joseph Rashon Chaney (QWLWOHG BIOCHEMICAL INVESTIGATION OF THE UBIQUITIN CARBOXYL-TERMINAL HYDROLASE FAMILY Doctor of Philosophy )RUWKHGHJUHHRI ,VDSSURYHGE\WKHILQDOH[DPLQLQJFRPPLWWHH Chittaranjan Das Angeline Lyon Christine A. Hrycyna George M. Bodner To the best of my knowledge and as understood by the student in the Thesis/Dissertation Agreement, Publication Delay, and Certification/Disclaimer (Graduate School Form 32), this thesis/dissertation adheres to the provisions of Purdue University’s “Policy on Integrity in Research” and the use of copyrighted material. Chittaranjan Das $SSURYHGE\0DMRU3URIHVVRU V BBBBBBBBBBBBBBBBBBBBBBBBBBBBBBBBBBBB BBBBBBBBBBBBBBBBBBBBBBBBBBBBBBBBBBBB $SSURYHGE\R. E. Wild 04/24/2015 +HDGRIWKH'HSDUWPHQW*UDGXDWH3URJUDP 'DWH BIOCHEMICAL INVESTIGATION OF THE UBIQUITIN CARBOXYL-TERMINAL HYDROLASE FAMILY Dissertation Submitted to the Faculty of Purdue University by Joseph Rashon Chaney In Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy May 2015 Purdue University West Lafayette, Indiana ii All of this I dedicate wife, Millicent, to my faithful and beautiful children, Josh and Caleb. -

Babela Massiliensis, a Representative of A
Pagnier et al. Biology Direct (2015) 10:13 DOI 10.1186/s13062-015-0043-z RESEARCH Open Access Babela massiliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae Isabelle Pagnier1, Natalya Yutin2, Olivier Croce1, Kira S Makarova2, Yuri I Wolf2, Samia Benamar1, Didier Raoult1, Eugene V Koonin2 and Bernard La Scola1* Abstract Background: Only a small fraction of bacteria and archaea that are identifiable by metagenomics can be grown on standard media. Recent efforts on deep metagenomics sequencing, single-cell genomics and the use of specialized culture conditions (culturomics) increasingly yield novel microbes some of which represent previously uncharacterized phyla and possess unusual biological traits. Results: We report isolation and genome analysis of Babela massiliensis, an obligate intracellular parasite of Acanthamoeba castellanii. B. massiliensis shows an unusual, fission mode of cell multiplication whereby large, polymorphic bodies accumulate in the cytoplasm of infected amoeba and then split into mature bacterial cells. This unique mechanism of cell division is associated with a deep degradation of the cell division machinery and delayed expression of the ftsZ gene. The genome of B. massiliensis consists of a circular chromosome approximately 1.12 megabase in size that encodes, 981 predicted proteins, 38 tRNAs and one typical rRNA operon. Phylogenetic analysis shows that B. massiliensis belongs to the putative bacterial phylum TM6 that so far was represented by the draft genome of the JCVI TM6SC1 bacterium obtained by single cell genomics and numerous environmental sequences. Conclusions: Currently, B. massiliensis is the only cultivated member of the putative TM6 phylum. Phylogenomic analysis shows diverse taxonomic affinities for B.