In Vitro Functional Studies of the Tumour Vascular Target PCDH7 And
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Cerebral Autosomal Dominant Arteriopathy with Subcortical Infarcts and Leukoencephalopathy Revisited Genotype-Phenotype Correlations of All Published Cases
VIEWS AND REVIEWS OPEN ACCESS Cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy revisited Genotype-phenotype correlations of all published cases Georgia Xiromerisiou, MD, PhD, Chrysoula Marogianni, MD, MSc, Katerina Dadouli, MSc, Correspondence Christina Zompola, MD, Despoina Georgouli, MD, MSc, Antonios Provatas, MD, PhD, Dr. Xiromerisiou [email protected] Aikaterini Theodorou, MD, Paschalis Zervas, MD, Christina Nikolaidou, MD, Stergios Stergiou, MD, Panagiotis Ntellas, MD, Maria Sokratous, MD, MSc, Pantelis Stathis, MD, PhD, Georgios P. Paraskevas, MD, PhD, Anastasios Bonakis, MD, PhD, Konstantinos Voumvourakis, MD, PhD, Christos Hadjichristodoulou, MD, PhD, Georgios M. Hadjigeorgiou, MD, PhD, and Georgios Tsivgoulis, MD, PhD Neurol Genet 2020;6:e434. doi:10.1212/NXG.0000000000000434 Abstract Objective The aim of this study was to evaluate the correlation between the various NOTCH3 mutations and their clinical and genetic profile, along with the presentation of a novel mutation in a patient. Methods Here, we describe the phenotype of a patient with cerebral autosomal dominant arteriopathy with subcortical infarcts and leukoencephalopathy (CADASIL) harboring a novel mutation. We also performed an extensive literature research for NOTCH3 mutations published since the identification of the gene and performed a systematic review of all published cases with NOTCH3 mutations. We evaluated the mutation pathogenicity in a great number of patients with detailed clinical and genetic evaluation and investigated the possible phenotype-genotype correlations. Results Our patient harbored a novel mutation in the NOTCH3 gene, the c.3084 G > C, corresponding to the aminoacidic substitution p.Trp1028Cys, presenting with seizures as the first neurologic manifestation. We managed to find a correlation between the pathogenicity of mutations, severity of the phenotype, and age at onset of CADASIL. -

Role and Regulation of the P53-Homolog P73 in the Transformation of Normal Human Fibroblasts
Role and regulation of the p53-homolog p73 in the transformation of normal human fibroblasts Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Bayerischen Julius-Maximilians-Universität Würzburg vorgelegt von Lars Hofmann aus Aschaffenburg Würzburg 2007 Eingereicht am Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Dr. Martin J. Müller Gutachter: Prof. Dr. Michael P. Schön Gutachter : Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am Erklärung Hiermit erkläre ich, dass ich die vorliegende Arbeit selbständig angefertigt und keine anderen als die angegebenen Hilfsmittel und Quellen verwendet habe. Diese Arbeit wurde weder in gleicher noch in ähnlicher Form in einem anderen Prüfungsverfahren vorgelegt. Ich habe früher, außer den mit dem Zulassungsgesuch urkundlichen Graden, keine weiteren akademischen Grade erworben und zu erwerben gesucht. Würzburg, Lars Hofmann Content SUMMARY ................................................................................................................ IV ZUSAMMENFASSUNG ............................................................................................. V 1. INTRODUCTION ................................................................................................. 1 1.1. Molecular basics of cancer .......................................................................................... 1 1.2. Early research on tumorigenesis ................................................................................. 3 1.3. Developing -

Genome-Wide Analysis of LRR-RLK Gene Family in Four Gossypium Species and Expression Analysis During Cotton Development and Stress Responses
G C A T T A C G G C A T genes Article Genome-Wide Analysis of LRR-RLK Gene Family in Four Gossypium Species and Expression Analysis during Cotton Development and Stress Responses Ruibin Sun 1, Shaohui Wang 1, Dan Ma 1 and Chuanliang Liu 1,2,* 1 State Key Laboratory of Cotton Biology, Institute of Cotton Research, Chinese Academy of Agricultural Sciences, Anyang 455000, China; [email protected] (R.S.); [email protected] (S.W.); [email protected] (D.M.) 2 Zhengzhou Research Base, State Key Laboratory of Cotton Biology, Zhengzhou University, Zhengzhou 450066, China * Correspondence: [email protected] Received: 30 September 2018; Accepted: 26 November 2018; Published: 29 November 2018 Abstract: Leucine-rich repeat receptor-like kinases (LRR-RLKs) have been reported to play important roles in plant growth, development, and stress responses. However, no comprehensive analysis of this family has been performed in cotton (Gossypium spp.), which is an important economic crop that suffers various stresses in growth and development. Here we conducted a comprehensive analysis of LRR-RLK family in four Gossypium species (Gossypium arboreum, Gossypium barbadense, Gossypium hirsutum, and Gossypium raimondii). A total of 1641 LRR-RLK genes were identified in the four Gossypium species involved in our study. The maximum-likelihood phylogenetic tree revealed that all the LRR-RLK genes were divided into 21 subgroups. Exon-intron organization structure of LRR-RLK genes kept relatively conserved within subfamilies and between Arabidopsis and Gossypium genomes. Notably, subfamilies XI and XII were found dramatically expanded in Gossypium species. Tandem duplication acted as an important mechanism in expansion of the Gossypium LRR-RLK gene family. -

How Chaotic Is Genome Chaos?
cancers Review How Chaotic Is Genome Chaos? James A. Shapiro Department of Biochemistry and Molecular Biology, University of Chicago, Chicago, IL 60637, USA; [email protected] Simple Summary: Cancer genomes can undergo major restructurings involving many chromosomal locations at key stages in tumor development. This restructuring process has been designated “genome chaos” by some authors. In order to examine how chaotic cancer genome restructuring may be, the cell and molecular processes for DNA restructuring are reviewed. Examination of the action of these processes in various cancers reveals a degree of specificity that indicates genome restructuring may be sufficiently reproducible to enable possible therapies that interrupt tumor progression to more lethal forms. Abstract: Cancer genomes evolve in a punctuated manner during tumor evolution. Abrupt genome restructuring at key steps in this evolution has been called “genome chaos.” To answer whether widespread genome change is truly chaotic, this review (i) summarizes the limited number of cell and molecular systems that execute genome restructuring, (ii) describes the characteristic signatures of DNA changes that result from activity of those systems, and (iii) examines two cases where genome restructuring is determined to a significant degree by cell type or viral infection. The conclusion is that many restructured cancer genomes display sufficiently unchaotic signatures to identify the cellular systems responsible for major oncogenic transitions, thereby identifying possible targets for therapies to inhibit tumor progression to greater aggressiveness. Keywords: DNA break repair; alternative end-joining (alt-EJ); chromothripsis; chromoplexy; chro- Citation: Shapiro, J.A. How Chaotic moanasynthesis; retrotransposition; target-primed reverse transcription (TPRT); immunoglobulin Is Genome Chaos? Cancers 2021, 13, VDJ joining; class switch recombination (CSR); human papillomavirus (HPV) 1358. -

On the Role of Chromosomal Rearrangements in Evolution
On the role of chromosomal rearrangements in evolution: Reconstruction of genome reshuffling in rodents and analysis of Robertsonian fusions in a house mouse chromosomal polymorphism zone by Laia Capilla Pérez A thesis submitted for the degree of Doctor of Philosophy in Animal Biology Supervisors: Dra. Aurora Ruiz-Herrera Moreno and Dr. Jacint Ventura Queija Institut de Biotecnologia i Biomedicina (IBB) Departament de Biologia Cel·lular, Fisiologia i Immunologia Departament de Biologia Animal, Biologia Vegetal i Ecologia Universitat Autònoma de Barcelona Supervisor Supervisor PhD candidate Aurora Ruiz-Herrera Moreno Jacint Ventura Queija Laia Capilla Pérez Bellaterra, 2015 A la mare Al pare Al mano “Visto a la luz de la evolución, la biología es, quizás, la ciencia más satisfactoria e inspiradora. Sin esa luz, se convierte en un montón de hechos varios, algunos de ellos interesantes o curiosos, pero sin formar ninguna visión conjunta.” Theodosius Dobzhansky “La evolución es tan creativa. Por eso tenemos jirafas.” Kurt Vonnegut This thesis was supported by grants from: • Ministerio de Economía y Competitividad (CGL2010-15243 and CGL2010- 20170). • Generalitat de Catalunya, GRQ 1057. • Ministerio de Economía y Competitividad. Beca de Formación de Personal Investigador (FPI) (BES-2011-047722). • Ministerio de Economía y Competitividad. Beca para la realización de estancias breves (EEBB-2011-07350). Covers designed by cintamontserrat.blogspot.com INDEX Abstract 15-17 Acronyms 19-20 1. GENERAL INTRODUCTION 21-60 1.1 Chromosomal rearrangements -

Downregulation of SNRPG Induces Cell Cycle Arrest and Sensitizes Human Glioblastoma Cells to Temozolomide by Targeting Myc Through a P53-Dependent Signaling Pathway
Cancer Biol Med 2020. doi: 10.20892/j.issn.2095-3941.2019.0164 ORIGINAL ARTICLE Downregulation of SNRPG induces cell cycle arrest and sensitizes human glioblastoma cells to temozolomide by targeting Myc through a p53-dependent signaling pathway Yulong Lan1,2*, Jiacheng Lou2*, Jiliang Hu1, Zhikuan Yu1, Wen Lyu1, Bo Zhang1,2 1Department of Neurosurgery, Shenzhen People’s Hospital, Second Clinical Medical College of Jinan University, The First Affiliated Hospital of Southern University of Science and Technology, Shenzhen 518020, China;2 Department of Neurosurgery, The Second Affiliated Hospital of Dalian Medical University, Dalian 116023, China ABSTRACT Objective: Temozolomide (TMZ) is commonly used for glioblastoma multiforme (GBM) chemotherapy. However, drug resistance limits its therapeutic effect in GBM treatment. RNA-binding proteins (RBPs) have vital roles in posttranscriptional events. While disturbance of RBP-RNA network activity is potentially associated with cancer development, the precise mechanisms are not fully known. The SNRPG gene, encoding small nuclear ribonucleoprotein polypeptide G, was recently found to be related to cancer incidence, but its exact function has yet to be elucidated. Methods: SNRPG knockdown was achieved via short hairpin RNAs. Gene expression profiling and Western blot analyses were used to identify potential glioma cell growth signaling pathways affected by SNRPG. Xenograft tumors were examined to determine the carcinogenic effects of SNRPG on glioma tissues. Results: The SNRPG-mediated inhibitory effect on glioma cells might be due to the targeted prevention of Myc and p53. In addition, the effects of SNRPG loss on p53 levels and cell cycle progression were found to be Myc-dependent. Furthermore, SNRPG was increased in TMZ-resistant GBM cells, and downregulation of SNRPG potentially sensitized resistant cells to TMZ, suggesting that SNRPG deficiency decreases the chemoresistance of GBM cells to TMZ via the p53 signaling pathway. -

A Protocol for Constructing Gene Targeting Vectors: Generating Knockout Mice for the Cadherin Family and Beyond
PROTOCOL A protocol for constructing gene targeting vectors: generating knockout mice for the cadherin family and beyond Sen Wu1, Guoxin Ying2, Qiang Wu2 & Mario R Capecchi1,2 1Howard Hughes Medical Institute and 2Department of Human Genetics, University of Utah, Salt Lake City, Utah 84112, USA. Correspondence should be addressed to M.R.C. ([email protected]). Published online 29 May 2008; doi:10.1038/nprot.2008.70 s We describe here a streamlined procedure for targeting vector construction, which often is a limiting factor for gene targeting (knockout) technology. This procedure combines various highly efficient recombination-based cloning methods in bacteria, consisting of three steps. First step is the use of Red-pathway-mediated recombination (recombineering) to capture a genomic fragment into a Gateway-compatible vector. Second, the vector is modified by recombineering to include a positive selection gene neo,fromavariety natureprotocol / of modular reagents. Finally, through a simple in vitro Gateway recombination, the modified genomic fragment is switched into a m o c vector that contains negative selection cassettes, as well as unique sites for linearization. To demonstrate the usefulness of this . e r protocol, we report targeted disruptions of members of the cadherin gene family, focusing on those that have not been previously u t B a studied at the molecular genetic level. This protocol needs 2 weeks to construct a targeting vector, and several vectors can be n . easily handled simultaneously using common laboratory setup. w w w / / : p t INTRODUCTION t h Gene targeting, the use of homologous recombination in mouse this procedure is reduced when used for generating large DNA p 1–5 19,20 u embryonic stem (ES) cells to modify mouse genes precisely , constructs, required for gene targeting vector construction . -

1111111111111111111Inuuu11
1111111111111111111inuuu1111111111u~ (12) United States Patent (io) Patent No.: US 9,896,681 B2 Goodwin et al. (45) Date of Patent: *Feb. 20, 2018 (54) GENETIC REGULATION OF BONE AND 4,993,413 A 2/1991 McLeod CELLS BY ELECTROMAGNETIC 5,002,890 A 3/1991 Morrison STIMULATION FIELDS AND USES 5,026,650 A 6/1991 Schwarz 5,153,132 A 10/1992 Goodwin THEREOF 5,153,133 A 10/1992 Schwarz 5,155,034 A 10/1992 Wolf (71) Applicant: The United States of America as 5,155,035 A 10/1992 Schwarz Represented by the Administrator of 5,308,764 A 5/1994 Goodwin the National Aeronautics and Space 5,627,021 A 5/1997 Goodwin 5,846,807 A 12/1998 Goodwin Administration, Washington, DC (US) 6,485,963 B1 11/2002 Wolf 6,673,597 B2 1/2004 Wolf (72) Inventors: Thomas J. Goodwin, Kemah, TX 6,730,498 B1 5/2004 Goodwin (US); Linda C. Shackelford, Webster, 6,919,205 B2 7/2005 Brighton TX (US) 7,160,024 B2 1/2007 Dougherty, Sr. 7,179,217 B2 2/2007 Goodwin 7,456,019 B2 11/2008 Goodwin (73) Assignee: The United States of America as 2006/0229487 Al 10/2006 Goodwin represented by the National 2007/0105769 Al 5/2007 Simon Aeronautics and Space 2008/0138415 Al 6/2008 Hussain Administration, Washington, DC (US) 2009/0234417 Al 9/2009 Pastena 2011/0105959 Al 5/2011 OConnor (*) Notice: Subject to any disclaimer, the term of this patent is extended or adjusted under 35 OTHER PUBLICATIONS U.S.C. -
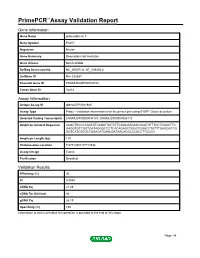
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name protocadherin 7 Gene Symbol Pcdh7 Organism Mouse Gene Summary Description Not Available Gene Aliases Not Available RefSeq Accession No. NC_000071.6, NT_039305.8 UniGene ID Mm.332387 Ensembl Gene ID ENSMUSG00000029108 Entrez Gene ID 54216 Assay Information Unique Assay ID qMmuCEP0041945 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENSMUST00000094783, ENSMUST00000068110 Amplicon Context Sequence AAGCTGCCCCAATGTCAGATGATCTTCGACGAGAACGAATGTTTCCTGGACTTC GAGGTGTCGGTGATAGGGCCCTCACAGAGCTGGGTGGACCTGTTTGAGGGTCG GGTCATCGTGCTGGACATCAACGATAACACGCCCACCTTCCCG Amplicon Length (bp) 120 Chromosome Location 5:57719387-57719536 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 93 R2 0.9994 cDNA Cq 21.89 cDNA Tm (Celsius) 86 gDNA Cq 26.19 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report Pcdh7, Mouse Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Mouse Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. Detected Coding Transcript(s) This is a list of the Ensembl transcript ID(s) that this assay will detect. -

Role of ECDYSONELESS in ERBB2/HER2 Mediated Breast Oncogenesis
University of Nebraska Medical Center DigitalCommons@UNMC Theses & Dissertations Graduate Studies Spring 5-7-2016 Role of ECDYSONELESS in ERBB2/HER2 Mediated Breast Oncogenesis Shalis A. Ammons University of Nebraska Medical Center Follow this and additional works at: https://digitalcommons.unmc.edu/etd Part of the Biology Commons, and the Cancer Biology Commons Recommended Citation Ammons, Shalis A., "Role of ECDYSONELESS in ERBB2/HER2 Mediated Breast Oncogenesis" (2016). Theses & Dissertations. 107. https://digitalcommons.unmc.edu/etd/107 This Dissertation is brought to you for free and open access by the Graduate Studies at DigitalCommons@UNMC. It has been accepted for inclusion in Theses & Dissertations by an authorized administrator of DigitalCommons@UNMC. For more information, please contact [email protected]. ROLE OF ECDYSONELESS IN ERBB2/HER2 MEDIATED BREAST ONCOGENESIS By Shalis Ammons A DISSERTATION Presented to the Faculty of the University of Nebraska Graduate College in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy Genetics, Cell Biology & Anatomy Graduate Program Under the supervision of Professor Vimla Band University of Nebraska Medical Center Omaha, Nebraska April, 2016 Supervisory Committee: Karen Gould, Ph.D Kay-Uwe Wagner, Ph.D Hamid Band, Ph.D Kaustubh Datta, Ph.D ROLE OF ECDYSONELESS IN ERBB2/HER2 MEDIATED BREAST ONCOGENESIS By Shalis Ammons University of Nebraska Medical Center, 2016 Advisor: Vimla Band, PhD Breast cancer is the second leading cause of cancer related deaths in women in the United States. The human Epidermal Growth Factor 2 (ErbB2) gene amplification and/or receptor overexpression subtype of breast cancer accounts for 25% of all breast cancers. -

Genes of Innate Immunity and Their Significance in Evolutionary Ecology of Free Livings Rodents Alena Fornuskova
Genes of innate immunity and their significance in evolutionary ecology of free livings rodents Alena Fornuskova To cite this version: Alena Fornuskova. Genes of innate immunity and their significance in evolutionary ecology of free livings rodents. Populations and Evolution [q-bio.PE]. Université Montpellier II - Sciences et Tech- niques du Languedoc; Masarykova univerzita (Brno, République tchèque), 2013. English. NNT : 2013MON20103. tel-01021258 HAL Id: tel-01021258 https://tel.archives-ouvertes.fr/tel-01021258 Submitted on 9 Jul 2014 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. UNIVERSITE•MONTPELLIER•II•• SCIENCES•ET•TECHNIQUES•DU•LANGUEDOC•• FACULTE•DES•SCIENCES• • and• • MASARYK•UNIVERSITY,•BRNO• FACULTY•OF•SCIENCE• • THESIS•• • To•obtain•doctoral•degree• • Formation•doctorale :•Biologie•de•l'évolution•et•écologie• • Ecole•Doctorale :•Systèmes•Intégrés•en•Biologie,•Agronomie,•Géosciences,•Hydrosciences, • Environnement,•SIBAGHE• • • Presented•and•defended•publicly• • AUTHOR:•Alena•Fornuskova• • • GENES•OF•INNATE•IMMUNITY•AND•THEIR•SIGNIFICANCE•IN•