University of Cincinnati
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Role of Amylase in Ovarian Cancer Mai Mohamed University of South Florida, [email protected]
University of South Florida Scholar Commons Graduate Theses and Dissertations Graduate School July 2017 Role of Amylase in Ovarian Cancer Mai Mohamed University of South Florida, [email protected] Follow this and additional works at: http://scholarcommons.usf.edu/etd Part of the Pathology Commons Scholar Commons Citation Mohamed, Mai, "Role of Amylase in Ovarian Cancer" (2017). Graduate Theses and Dissertations. http://scholarcommons.usf.edu/etd/6907 This Dissertation is brought to you for free and open access by the Graduate School at Scholar Commons. It has been accepted for inclusion in Graduate Theses and Dissertations by an authorized administrator of Scholar Commons. For more information, please contact [email protected]. Role of Amylase in Ovarian Cancer by Mai Mohamed A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy Department of Pathology and Cell Biology Morsani College of Medicine University of South Florida Major Professor: Patricia Kruk, Ph.D. Paula C. Bickford, Ph.D. Meera Nanjundan, Ph.D. Marzenna Wiranowska, Ph.D. Lauri Wright, Ph.D. Date of Approval: June 29, 2017 Keywords: ovarian cancer, amylase, computational analyses, glycocalyx, cellular invasion Copyright © 2017, Mai Mohamed Dedication This dissertation is dedicated to my parents, Ahmed and Fatma, who have always stressed the importance of education, and, throughout my education, have been my strongest source of encouragement and support. They always believed in me and I am eternally grateful to them. I would also like to thank my brothers, Mohamed and Hussien, and my sister, Mariam. I would also like to thank my husband, Ahmed. -

Role and Regulation of the P53-Homolog P73 in the Transformation of Normal Human Fibroblasts
Role and regulation of the p53-homolog p73 in the transformation of normal human fibroblasts Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Bayerischen Julius-Maximilians-Universität Würzburg vorgelegt von Lars Hofmann aus Aschaffenburg Würzburg 2007 Eingereicht am Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Dr. Martin J. Müller Gutachter: Prof. Dr. Michael P. Schön Gutachter : Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am Erklärung Hiermit erkläre ich, dass ich die vorliegende Arbeit selbständig angefertigt und keine anderen als die angegebenen Hilfsmittel und Quellen verwendet habe. Diese Arbeit wurde weder in gleicher noch in ähnlicher Form in einem anderen Prüfungsverfahren vorgelegt. Ich habe früher, außer den mit dem Zulassungsgesuch urkundlichen Graden, keine weiteren akademischen Grade erworben und zu erwerben gesucht. Würzburg, Lars Hofmann Content SUMMARY ................................................................................................................ IV ZUSAMMENFASSUNG ............................................................................................. V 1. INTRODUCTION ................................................................................................. 1 1.1. Molecular basics of cancer .......................................................................................... 1 1.2. Early research on tumorigenesis ................................................................................. 3 1.3. Developing -

A Workbench for Interactive Exploratory Data Analysis of Large Expression Datasets
bioRxiv preprint doi: https://doi.org/10.1101/698969; this version posted July 14, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. MetaOmGraph: a workbench for interactive exploratory data analysis of large expression datasets Urminder Singh1,2, Manhoi Hur2, Karin Dorman1,2,3, and Eve Wurtele1,2* 1Bioinformatics and Computational Biology Program, Iowa State University, Ames, IA 50011, USA 2Department of Genetics Development and Cell Biology, Iowa State University, Ames, IA 50011, USA 3Department of Statistics, Iowa State University, Ames, IA 50011, USA The diverse and growing omics data in public domains provide can be attained with bigger datasets and the wide variety researchers with a tremendous opportunity to extract hidden of biological conditions can reveal the diverse transcription knowledge. However, the challenge of providing domain ex- spectrum of genes. Yet, despite the availability of such vast perts with easy access to these big data has resulted in the vast data resources, most bioinformatic studies use only a limited majority of archived data remaining unused. Here, we present amount of the available data. MetaOmGraph (MOG), a free, open-source, standalone soft- A common goal of analyzing omics data is to infer func- ware for exploratory data analysis of massive datasets by sci- entific researchers. Using MOG, a researcher can interactively tional roles of particular features by investigating differen- visualize and statistically analyze the data, in the context of its tial expression and coexpression patterns. A wide variety metadata. Researchers can interactively hone-in on groups of of R-platform tools can provide specific analyses (9–16). -

Downregulation of SNRPG Induces Cell Cycle Arrest and Sensitizes Human Glioblastoma Cells to Temozolomide by Targeting Myc Through a P53-Dependent Signaling Pathway
Cancer Biol Med 2020. doi: 10.20892/j.issn.2095-3941.2019.0164 ORIGINAL ARTICLE Downregulation of SNRPG induces cell cycle arrest and sensitizes human glioblastoma cells to temozolomide by targeting Myc through a p53-dependent signaling pathway Yulong Lan1,2*, Jiacheng Lou2*, Jiliang Hu1, Zhikuan Yu1, Wen Lyu1, Bo Zhang1,2 1Department of Neurosurgery, Shenzhen People’s Hospital, Second Clinical Medical College of Jinan University, The First Affiliated Hospital of Southern University of Science and Technology, Shenzhen 518020, China;2 Department of Neurosurgery, The Second Affiliated Hospital of Dalian Medical University, Dalian 116023, China ABSTRACT Objective: Temozolomide (TMZ) is commonly used for glioblastoma multiforme (GBM) chemotherapy. However, drug resistance limits its therapeutic effect in GBM treatment. RNA-binding proteins (RBPs) have vital roles in posttranscriptional events. While disturbance of RBP-RNA network activity is potentially associated with cancer development, the precise mechanisms are not fully known. The SNRPG gene, encoding small nuclear ribonucleoprotein polypeptide G, was recently found to be related to cancer incidence, but its exact function has yet to be elucidated. Methods: SNRPG knockdown was achieved via short hairpin RNAs. Gene expression profiling and Western blot analyses were used to identify potential glioma cell growth signaling pathways affected by SNRPG. Xenograft tumors were examined to determine the carcinogenic effects of SNRPG on glioma tissues. Results: The SNRPG-mediated inhibitory effect on glioma cells might be due to the targeted prevention of Myc and p53. In addition, the effects of SNRPG loss on p53 levels and cell cycle progression were found to be Myc-dependent. Furthermore, SNRPG was increased in TMZ-resistant GBM cells, and downregulation of SNRPG potentially sensitized resistant cells to TMZ, suggesting that SNRPG deficiency decreases the chemoresistance of GBM cells to TMZ via the p53 signaling pathway. -

An Integrated Pan-Cancer Analysis and Structure-Based Virtual Screening of GPR15
International Journal of Molecular Sciences Article An Integrated Pan-Cancer Analysis and Structure-Based Virtual Screening of GPR15 1, 1, 1 1 1 2 Yanjing Wang y , Xiangeng Wang y , Yi Xiong , Cheng-Dong Li , Qin Xu , Lu Shen , Aman Chandra Kaushik 3,* and Dong-Qing Wei 1,4,* 1 State Key Laboratory of Microbial Metabolism, School of Life Sciences and Biotechnology, and Joint Laboratory of International Cooperation in Metabolic and Developmental Sciences, Ministry of Education, Shanghai Jiao Tong University, Shanghai 200240, China; [email protected] (Y.W.); [email protected] (X.W.); [email protected] (Y.X.); [email protected] (C.-D.L.); [email protected] (Q.X.) 2 Bio-X Institutes, Key Laboratory for the Genetics of Developmental and Neuropsychiatric Disorders, Ministry of Education, Shanghai Jiao Tong University, Shanghai 200030, China; [email protected] 3 Wuxi School of Medicine, Jiangnan University, Wuxi 214122, China 4 Peng Cheng Laboratory, Vanke Cloud City Phase I Building 8, Xili Street, Nanshan District, Shenzhen 518055, China * Correspondence: [email protected] (A.C.K.); [email protected] (D.-Q.W.) These authors contributed equally to this work. y Received: 21 July 2019; Accepted: 4 December 2019; Published: 10 December 2019 Abstract: G protein-coupled receptor 15 (GPR15, also known as BOB) is an extensively studied orphan G protein-coupled receptors (GPCRs) involving human immunodeficiency virus (HIV) infection, colonic inflammation, and smoking-related diseases. Recently, GPR15 was deorphanized and its corresponding natural ligand demonstrated an ability to inhibit cancer cell growth. However, no study reported the potential role of GPR15 in a pan-cancer manner. -

A Protocol for Constructing Gene Targeting Vectors: Generating Knockout Mice for the Cadherin Family and Beyond
PROTOCOL A protocol for constructing gene targeting vectors: generating knockout mice for the cadherin family and beyond Sen Wu1, Guoxin Ying2, Qiang Wu2 & Mario R Capecchi1,2 1Howard Hughes Medical Institute and 2Department of Human Genetics, University of Utah, Salt Lake City, Utah 84112, USA. Correspondence should be addressed to M.R.C. ([email protected]). Published online 29 May 2008; doi:10.1038/nprot.2008.70 s We describe here a streamlined procedure for targeting vector construction, which often is a limiting factor for gene targeting (knockout) technology. This procedure combines various highly efficient recombination-based cloning methods in bacteria, consisting of three steps. First step is the use of Red-pathway-mediated recombination (recombineering) to capture a genomic fragment into a Gateway-compatible vector. Second, the vector is modified by recombineering to include a positive selection gene neo,fromavariety natureprotocol / of modular reagents. Finally, through a simple in vitro Gateway recombination, the modified genomic fragment is switched into a m o c vector that contains negative selection cassettes, as well as unique sites for linearization. To demonstrate the usefulness of this . e r protocol, we report targeted disruptions of members of the cadherin gene family, focusing on those that have not been previously u t B a studied at the molecular genetic level. This protocol needs 2 weeks to construct a targeting vector, and several vectors can be n . easily handled simultaneously using common laboratory setup. w w w / / : p t INTRODUCTION t h Gene targeting, the use of homologous recombination in mouse this procedure is reduced when used for generating large DNA p 1–5 19,20 u embryonic stem (ES) cells to modify mouse genes precisely , constructs, required for gene targeting vector construction . -

1111111111111111111Inuuu11
1111111111111111111inuuu1111111111u~ (12) United States Patent (io) Patent No.: US 9,896,681 B2 Goodwin et al. (45) Date of Patent: *Feb. 20, 2018 (54) GENETIC REGULATION OF BONE AND 4,993,413 A 2/1991 McLeod CELLS BY ELECTROMAGNETIC 5,002,890 A 3/1991 Morrison STIMULATION FIELDS AND USES 5,026,650 A 6/1991 Schwarz 5,153,132 A 10/1992 Goodwin THEREOF 5,153,133 A 10/1992 Schwarz 5,155,034 A 10/1992 Wolf (71) Applicant: The United States of America as 5,155,035 A 10/1992 Schwarz Represented by the Administrator of 5,308,764 A 5/1994 Goodwin the National Aeronautics and Space 5,627,021 A 5/1997 Goodwin 5,846,807 A 12/1998 Goodwin Administration, Washington, DC (US) 6,485,963 B1 11/2002 Wolf 6,673,597 B2 1/2004 Wolf (72) Inventors: Thomas J. Goodwin, Kemah, TX 6,730,498 B1 5/2004 Goodwin (US); Linda C. Shackelford, Webster, 6,919,205 B2 7/2005 Brighton TX (US) 7,160,024 B2 1/2007 Dougherty, Sr. 7,179,217 B2 2/2007 Goodwin 7,456,019 B2 11/2008 Goodwin (73) Assignee: The United States of America as 2006/0229487 Al 10/2006 Goodwin represented by the National 2007/0105769 Al 5/2007 Simon Aeronautics and Space 2008/0138415 Al 6/2008 Hussain Administration, Washington, DC (US) 2009/0234417 Al 9/2009 Pastena 2011/0105959 Al 5/2011 OConnor (*) Notice: Subject to any disclaimer, the term of this patent is extended or adjusted under 35 OTHER PUBLICATIONS U.S.C. -
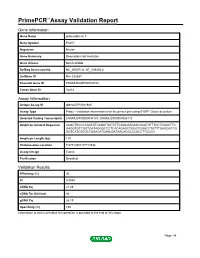
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name protocadherin 7 Gene Symbol Pcdh7 Organism Mouse Gene Summary Description Not Available Gene Aliases Not Available RefSeq Accession No. NC_000071.6, NT_039305.8 UniGene ID Mm.332387 Ensembl Gene ID ENSMUSG00000029108 Entrez Gene ID 54216 Assay Information Unique Assay ID qMmuCEP0041945 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENSMUST00000094783, ENSMUST00000068110 Amplicon Context Sequence AAGCTGCCCCAATGTCAGATGATCTTCGACGAGAACGAATGTTTCCTGGACTTC GAGGTGTCGGTGATAGGGCCCTCACAGAGCTGGGTGGACCTGTTTGAGGGTCG GGTCATCGTGCTGGACATCAACGATAACACGCCCACCTTCCCG Amplicon Length (bp) 120 Chromosome Location 5:57719387-57719536 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 93 R2 0.9994 cDNA Cq 21.89 cDNA Tm (Celsius) 86 gDNA Cq 26.19 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report Pcdh7, Mouse Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Mouse Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. Detected Coding Transcript(s) This is a list of the Ensembl transcript ID(s) that this assay will detect. -

Human C10orf99 Alexa Fluor® 488‑Conjugated Antibody
Human C10orf99 Alexa Fluor® 488‑conjugated Antibody Monoclonal Mouse IgG2B Clone # 1020904 Catalog Number: IC104161G 100 µg DESCRIPTION Species Reactivity Human Specificity Detects human C10orf99 in direct ELISAs. Source Monoclonal Mouse IgG2B Clone # 1020904 Purification Protein A or G purified from cell culture supernatant Immunogen E. coli-derived human C10orf99 Conjugate Alexa Fluor 488 Excitation Wavelength: 488 nm Emission Wavelength: 515-545 nm Formulation Supplied 0.2 mg/mL in a saline solution containing BSA and Sodium Azid *Contains <0.1% Sodium Azide, which is not hazardous at this concentration according to GHS classifications. Refer to the Safety Data Sheet (SDS) for additional information and handling instructions. APPLICATIONS Please Note: Optimal dilutions should be determined by each laboratory for each application. General Protocols are available in the Technical Information section on our website. Recommended Sample Concentration Intracellular Staining by Flow Cytometry 0.25-1 µg/106 cells SW40 human cell line fixed with Flow Cytometry Fixation Buffer (Catalog # FC004) and permeabilized with Flow Cytometry Permeabilization/Wash Buffer I (Catalog # FC005) PREPARATION AND STORAGE Shipping The product is shipped with polar packs. Upon receipt, store it immediately at the temperature recommended below. Stability & Storage Protect from light. Do not freeze. 12 months from date of receipt, 2 to 8 °C as supplied. BACKGROUND Human Chromosome 10 open reading frame 99 is a 9.2 KDa protein encoded by the C10orf99 gene. C10orf99 has been described as a potential cytokine and a ligand Sushi Domain Containing 2 (SUSD2), inducing SUSD2 internalization upon binding. It has also been proposed to be a chemotactic factor that mediates lymphocytes recruitment to epithelia through binding and activation of the G-protein coupled receptor GPR15. -

Genome-Wide Association Study Identifies Candidate Genes
animals Article Genome-Wide Association Study Identifies Candidate Genes Associated with Feet and Leg Conformation Traits in Chinese Holstein Cattle Ismail Mohamed Abdalla 1, Xubin Lu 1 , Mudasir Nazar 1, Abdelaziz Adam Idriss Arbab 1,2, Tianle Xu 3 , Mohammed Husien Yousif 4, Yongjiang Mao 1 and Zhangping Yang 1,* 1 College of Animal Science and Technology, Yangzhou University, Yangzhou 225009, China; [email protected] (I.M.A.); [email protected] (X.L.); [email protected] (M.N.); [email protected] (A.A.I.A.); [email protected] (Y.M.) 2 Biomedical Research Institute, Darfur College, Nyala 63313, Sudan 3 Joint International Research Laboratory of Agriculture and Agri-Product Safety, Yangzhou University, Yangzhou 225009, China; [email protected] 4 Faculty of Animal Production, West Kordufan University, Alnuhud City 12942, Sudan; [email protected] * Correspondence: [email protected]; Tel.: +86-0514-87979269 Simple Summary: Feet and leg problems are among the major reasons for dairy cows leaving the herd, as well as having direct association with production and reproduction efficiency, health (e.g., claw disorders and lameness) and welfare. Hence, understanding the genetic architecture underlying feet and conformation traits in dairy cattle offers new opportunities toward the genetic improvement and long-term selection. Through a genome-wide association study on Chinese Holstein cattle, we identified several candidate genes associated with feet and leg conformation traits. These results could provide useful information about the molecular breeding basis of feet and leg traits, thus Citation: Abdalla, I.M.; Lu, X.; Nazar, improving the longevity and productivity of dairy cattle. -

Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands That Promote Axonal Growth
Research Article: New Research Development Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands that Promote Axonal Growth Jeremy S. Toma,1 Konstantina Karamboulas,1,ª Matthew J. Carr,1,2,ª Adelaida Kolaj,1,3 Scott A. Yuzwa,1 Neemat Mahmud,1,3 Mekayla A. Storer,1 David R. Kaplan,1,2,4 and Freda D. Miller1,2,3,4 https://doi.org/10.1523/ENEURO.0066-20.2020 1Program in Neurosciences and Mental Health, Hospital for Sick Children, 555 University Avenue, Toronto, Ontario M5G 1X8, Canada, 2Institute of Medical Sciences University of Toronto, Toronto, Ontario M5G 1A8, Canada, 3Department of Physiology, University of Toronto, Toronto, Ontario M5G 1A8, Canada, and 4Department of Molecular Genetics, University of Toronto, Toronto, Ontario M5G 1A8, Canada Abstract Peripheral nerves provide a supportive growth environment for developing and regenerating axons and are es- sential for maintenance and repair of many non-neural tissues. This capacity has largely been ascribed to paracrine factors secreted by nerve-resident Schwann cells. Here, we used single-cell transcriptional profiling to identify ligands made by different injured rodent nerve cell types and have combined this with cell-surface mass spectrometry to computationally model potential paracrine interactions with peripheral neurons. These analyses show that peripheral nerves make many ligands predicted to act on peripheral and CNS neurons, in- cluding known and previously uncharacterized ligands. While Schwann cells are an important ligand source within injured nerves, more than half of the predicted ligands are made by nerve-resident mesenchymal cells, including the endoneurial cells most closely associated with peripheral axons. At least three of these mesen- chymal ligands, ANGPT1, CCL11, and VEGFC, promote growth when locally applied on sympathetic axons.