NIH Public Access Author Manuscript Nat Genet
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Seq2pathway Vignette
seq2pathway Vignette Bin Wang, Xinan Holly Yang, Arjun Kinstlick May 19, 2021 Contents 1 Abstract 1 2 Package Installation 2 3 runseq2pathway 2 4 Two main functions 3 4.1 seq2gene . .3 4.1.1 seq2gene flowchart . .3 4.1.2 runseq2gene inputs/parameters . .5 4.1.3 runseq2gene outputs . .8 4.2 gene2pathway . 10 4.2.1 gene2pathway flowchart . 11 4.2.2 gene2pathway test inputs/parameters . 11 4.2.3 gene2pathway test outputs . 12 5 Examples 13 5.1 ChIP-seq data analysis . 13 5.1.1 Map ChIP-seq enriched peaks to genes using runseq2gene .................... 13 5.1.2 Discover enriched GO terms using gene2pathway_test with gene scores . 15 5.1.3 Discover enriched GO terms using Fisher's Exact test without gene scores . 17 5.1.4 Add description for genes . 20 5.2 RNA-seq data analysis . 20 6 R environment session 23 1 Abstract Seq2pathway is a novel computational tool to analyze functional gene-sets (including signaling pathways) using variable next-generation sequencing data[1]. Integral to this tool are the \seq2gene" and \gene2pathway" components in series that infer a quantitative pathway-level profile for each sample. The seq2gene function assigns phenotype-associated significance of genomic regions to gene-level scores, where the significance could be p-values of SNPs or point mutations, protein-binding affinity, or transcriptional expression level. The seq2gene function has the feasibility to assign non-exon regions to a range of neighboring genes besides the nearest one, thus facilitating the study of functional non-coding elements[2]. Then the gene2pathway summarizes gene-level measurements to pathway-level scores, comparing the quantity of significance for gene members within a pathway with those outside a pathway. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Role and Regulation of the P53-Homolog P73 in the Transformation of Normal Human Fibroblasts
Role and regulation of the p53-homolog p73 in the transformation of normal human fibroblasts Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Bayerischen Julius-Maximilians-Universität Würzburg vorgelegt von Lars Hofmann aus Aschaffenburg Würzburg 2007 Eingereicht am Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Dr. Martin J. Müller Gutachter: Prof. Dr. Michael P. Schön Gutachter : Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am Erklärung Hiermit erkläre ich, dass ich die vorliegende Arbeit selbständig angefertigt und keine anderen als die angegebenen Hilfsmittel und Quellen verwendet habe. Diese Arbeit wurde weder in gleicher noch in ähnlicher Form in einem anderen Prüfungsverfahren vorgelegt. Ich habe früher, außer den mit dem Zulassungsgesuch urkundlichen Graden, keine weiteren akademischen Grade erworben und zu erwerben gesucht. Würzburg, Lars Hofmann Content SUMMARY ................................................................................................................ IV ZUSAMMENFASSUNG ............................................................................................. V 1. INTRODUCTION ................................................................................................. 1 1.1. Molecular basics of cancer .......................................................................................... 1 1.2. Early research on tumorigenesis ................................................................................. 3 1.3. Developing -

Genomic and Transcriptome Analysis Revealing an Oncogenic Functional Module in Meningiomas
Neurosurg Focus 35 (6):E3, 2013 ©AANS, 2013 Genomic and transcriptome analysis revealing an oncogenic functional module in meningiomas XIAO CHANG, PH.D.,1 LINGLING SHI, PH.D.,2 FAN GAO, PH.D.,1 JONATHAN RUssIN, M.D.,3 LIYUN ZENG, PH.D.,1 SHUHAN HE, B.S.,3 THOMAS C. CHEN, M.D.,3 STEVEN L. GIANNOTTA, M.D.,3 DANIEL J. WEISENBERGER, PH.D.,4 GAbrIEL ZADA, M.D.,3 KAI WANG, PH.D.,1,5,6 AND WIllIAM J. MAck, M.D.1,3 1Zilkha Neurogenetic Institute, Keck School of Medicine, University of Southern California, Los Angeles, California; 2GHM Institute of CNS Regeneration, Jinan University, Guangzhou, China; 3Department of Neurosurgery, Keck School of Medicine, University of Southern California, Los Angeles, California; 4USC Epigenome Center, Keck School of Medicine, University of Southern California, Los Angeles, California; 5Department of Psychiatry, Keck School of Medicine, University of Southern California, Los Angeles, California; and 6Division of Bioinformatics, Department of Preventive Medicine, Keck School of Medicine, University of Southern California, Los Angeles, California Object. Meningiomas are among the most common primary adult brain tumors. Although typically benign, roughly 2%–5% display malignant pathological features. The key molecular pathways involved in malignant trans- formation remain to be determined. Methods. Illumina expression microarrays were used to assess gene expression levels, and Illumina single- nucleotide polymorphism arrays were used to identify copy number variants in benign, atypical, and malignant me- ningiomas (19 tumors, including 4 malignant ones). The authors also reanalyzed 2 expression data sets generated on Affymetrix microarrays (n = 68, including 6 malignant ones; n = 56, including 3 malignant ones). -

Downregulation of SNRPG Induces Cell Cycle Arrest and Sensitizes Human Glioblastoma Cells to Temozolomide by Targeting Myc Through a P53-Dependent Signaling Pathway
Cancer Biol Med 2020. doi: 10.20892/j.issn.2095-3941.2019.0164 ORIGINAL ARTICLE Downregulation of SNRPG induces cell cycle arrest and sensitizes human glioblastoma cells to temozolomide by targeting Myc through a p53-dependent signaling pathway Yulong Lan1,2*, Jiacheng Lou2*, Jiliang Hu1, Zhikuan Yu1, Wen Lyu1, Bo Zhang1,2 1Department of Neurosurgery, Shenzhen People’s Hospital, Second Clinical Medical College of Jinan University, The First Affiliated Hospital of Southern University of Science and Technology, Shenzhen 518020, China;2 Department of Neurosurgery, The Second Affiliated Hospital of Dalian Medical University, Dalian 116023, China ABSTRACT Objective: Temozolomide (TMZ) is commonly used for glioblastoma multiforme (GBM) chemotherapy. However, drug resistance limits its therapeutic effect in GBM treatment. RNA-binding proteins (RBPs) have vital roles in posttranscriptional events. While disturbance of RBP-RNA network activity is potentially associated with cancer development, the precise mechanisms are not fully known. The SNRPG gene, encoding small nuclear ribonucleoprotein polypeptide G, was recently found to be related to cancer incidence, but its exact function has yet to be elucidated. Methods: SNRPG knockdown was achieved via short hairpin RNAs. Gene expression profiling and Western blot analyses were used to identify potential glioma cell growth signaling pathways affected by SNRPG. Xenograft tumors were examined to determine the carcinogenic effects of SNRPG on glioma tissues. Results: The SNRPG-mediated inhibitory effect on glioma cells might be due to the targeted prevention of Myc and p53. In addition, the effects of SNRPG loss on p53 levels and cell cycle progression were found to be Myc-dependent. Furthermore, SNRPG was increased in TMZ-resistant GBM cells, and downregulation of SNRPG potentially sensitized resistant cells to TMZ, suggesting that SNRPG deficiency decreases the chemoresistance of GBM cells to TMZ via the p53 signaling pathway. -

Transdifferentiation of Human Mesenchymal Stem Cells
Transdifferentiation of Human Mesenchymal Stem Cells Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Julius-Maximilians-Universität Würzburg vorgelegt von Tatjana Schilling aus San Miguel de Tucuman, Argentinien Würzburg, 2007 Eingereicht am: Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Martin J. Müller Gutachter: PD Dr. Norbert Schütze Gutachter: Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am: Hiermit erkläre ich ehrenwörtlich, dass ich die vorliegende Dissertation selbstständig angefertigt und keine anderen als die von mir angegebenen Hilfsmittel und Quellen verwendet habe. Des Weiteren erkläre ich, dass diese Arbeit weder in gleicher noch in ähnlicher Form in einem Prüfungsverfahren vorgelegen hat und ich noch keinen Promotionsversuch unternommen habe. Gerbrunn, 4. Mai 2007 Tatjana Schilling Table of contents i Table of contents 1 Summary ........................................................................................................................ 1 1.1 Summary.................................................................................................................... 1 1.2 Zusammenfassung..................................................................................................... 2 2 Introduction.................................................................................................................... 4 2.1 Osteoporosis and the fatty degeneration of the bone marrow..................................... 4 2.2 Adipose and bone -

A Protocol for Constructing Gene Targeting Vectors: Generating Knockout Mice for the Cadherin Family and Beyond
PROTOCOL A protocol for constructing gene targeting vectors: generating knockout mice for the cadherin family and beyond Sen Wu1, Guoxin Ying2, Qiang Wu2 & Mario R Capecchi1,2 1Howard Hughes Medical Institute and 2Department of Human Genetics, University of Utah, Salt Lake City, Utah 84112, USA. Correspondence should be addressed to M.R.C. ([email protected]). Published online 29 May 2008; doi:10.1038/nprot.2008.70 s We describe here a streamlined procedure for targeting vector construction, which often is a limiting factor for gene targeting (knockout) technology. This procedure combines various highly efficient recombination-based cloning methods in bacteria, consisting of three steps. First step is the use of Red-pathway-mediated recombination (recombineering) to capture a genomic fragment into a Gateway-compatible vector. Second, the vector is modified by recombineering to include a positive selection gene neo,fromavariety natureprotocol / of modular reagents. Finally, through a simple in vitro Gateway recombination, the modified genomic fragment is switched into a m o c vector that contains negative selection cassettes, as well as unique sites for linearization. To demonstrate the usefulness of this . e r protocol, we report targeted disruptions of members of the cadherin gene family, focusing on those that have not been previously u t B a studied at the molecular genetic level. This protocol needs 2 weeks to construct a targeting vector, and several vectors can be n . easily handled simultaneously using common laboratory setup. w w w / / : p t INTRODUCTION t h Gene targeting, the use of homologous recombination in mouse this procedure is reduced when used for generating large DNA p 1–5 19,20 u embryonic stem (ES) cells to modify mouse genes precisely , constructs, required for gene targeting vector construction . -

Dynamic Changes in the Epigenomic Landscape Regulate Human Organogenesis and Link to Developmental Disorders
ARTICLE https://doi.org/10.1038/s41467-020-17305-2 OPEN Dynamic changes in the epigenomic landscape regulate human organogenesis and link to developmental disorders Dave T. Gerrard 1,6, Andrew A. Berry1,6, Rachel E. Jennings1,2, Matthew J. Birket 1, Peyman Zarrineh1, Myles G. Garstang1, Sarah L. Withey1, Patrick Short 3, Sandra Jiménez-Gancedo4, Panos N. Firbas4, Ian Donaldson 1, Andrew D. Sharrocks1, Karen Piper Hanley 1,5, Matthew E. Hurles3, ✉ José Luis Gomez-Skarmeta4, Nicoletta Bobola1 & Neil A. Hanley 1,2 1234567890():,; How the genome activates or silences transcriptional programmes governs organ formation. Little is known in human embryos undermining our ability to benchmark the fidelity of stem cell differentiation or cell programming, or interpret the pathogenicity of noncoding variation. Here, we study histone modifications across thirteen tissues during human organogenesis. We integrate the data with transcription to build an overview of how the human genome differentially regulates alternative organ fates including by repression. Promoters from nearly 20,000 genes partition into discrete states. Key developmental gene sets are actively repressed outside of the appropriate organ without obvious bivalency. Candidate enhancers, functional in zebrafish, allow imputation of tissue-specific and shared patterns of transcrip- tion factor binding. Overlaying more than 700 noncoding mutations from patients with developmental disorders allows correlation to unanticipated target genes. Taken together, the data provide a comprehensive genomic framework for investigating normal and abnormal human development. 1 Faculty of Biology, Medicine & Health, Manchester Academic Health Sciences Centre, University of Manchester, Oxford Road, Manchester M13 9PT, UK. 2 Endocrinology Department, Manchester University NHS Foundation Trust, Grafton Street, Manchester M13 9WU, UK. -

1111111111111111111Inuuu11
1111111111111111111inuuu1111111111u~ (12) United States Patent (io) Patent No.: US 9,896,681 B2 Goodwin et al. (45) Date of Patent: *Feb. 20, 2018 (54) GENETIC REGULATION OF BONE AND 4,993,413 A 2/1991 McLeod CELLS BY ELECTROMAGNETIC 5,002,890 A 3/1991 Morrison STIMULATION FIELDS AND USES 5,026,650 A 6/1991 Schwarz 5,153,132 A 10/1992 Goodwin THEREOF 5,153,133 A 10/1992 Schwarz 5,155,034 A 10/1992 Wolf (71) Applicant: The United States of America as 5,155,035 A 10/1992 Schwarz Represented by the Administrator of 5,308,764 A 5/1994 Goodwin the National Aeronautics and Space 5,627,021 A 5/1997 Goodwin 5,846,807 A 12/1998 Goodwin Administration, Washington, DC (US) 6,485,963 B1 11/2002 Wolf 6,673,597 B2 1/2004 Wolf (72) Inventors: Thomas J. Goodwin, Kemah, TX 6,730,498 B1 5/2004 Goodwin (US); Linda C. Shackelford, Webster, 6,919,205 B2 7/2005 Brighton TX (US) 7,160,024 B2 1/2007 Dougherty, Sr. 7,179,217 B2 2/2007 Goodwin 7,456,019 B2 11/2008 Goodwin (73) Assignee: The United States of America as 2006/0229487 Al 10/2006 Goodwin represented by the National 2007/0105769 Al 5/2007 Simon Aeronautics and Space 2008/0138415 Al 6/2008 Hussain Administration, Washington, DC (US) 2009/0234417 Al 9/2009 Pastena 2011/0105959 Al 5/2011 OConnor (*) Notice: Subject to any disclaimer, the term of this patent is extended or adjusted under 35 OTHER PUBLICATIONS U.S.C. -
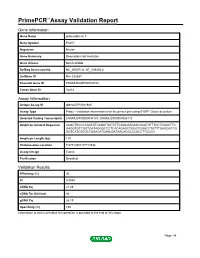
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name protocadherin 7 Gene Symbol Pcdh7 Organism Mouse Gene Summary Description Not Available Gene Aliases Not Available RefSeq Accession No. NC_000071.6, NT_039305.8 UniGene ID Mm.332387 Ensembl Gene ID ENSMUSG00000029108 Entrez Gene ID 54216 Assay Information Unique Assay ID qMmuCEP0041945 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENSMUST00000094783, ENSMUST00000068110 Amplicon Context Sequence AAGCTGCCCCAATGTCAGATGATCTTCGACGAGAACGAATGTTTCCTGGACTTC GAGGTGTCGGTGATAGGGCCCTCACAGAGCTGGGTGGACCTGTTTGAGGGTCG GGTCATCGTGCTGGACATCAACGATAACACGCCCACCTTCCCG Amplicon Length (bp) 120 Chromosome Location 5:57719387-57719536 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 93 R2 0.9994 cDNA Cq 21.89 cDNA Tm (Celsius) 86 gDNA Cq 26.19 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 1/4 PrimePCR™Assay Validation Report Pcdh7, Mouse Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 2/4 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Mouse Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. Detected Coding Transcript(s) This is a list of the Ensembl transcript ID(s) that this assay will detect. -

Genome-Wide Association Study Identifies Candidate Genes
animals Article Genome-Wide Association Study Identifies Candidate Genes Associated with Feet and Leg Conformation Traits in Chinese Holstein Cattle Ismail Mohamed Abdalla 1, Xubin Lu 1 , Mudasir Nazar 1, Abdelaziz Adam Idriss Arbab 1,2, Tianle Xu 3 , Mohammed Husien Yousif 4, Yongjiang Mao 1 and Zhangping Yang 1,* 1 College of Animal Science and Technology, Yangzhou University, Yangzhou 225009, China; [email protected] (I.M.A.); [email protected] (X.L.); [email protected] (M.N.); [email protected] (A.A.I.A.); [email protected] (Y.M.) 2 Biomedical Research Institute, Darfur College, Nyala 63313, Sudan 3 Joint International Research Laboratory of Agriculture and Agri-Product Safety, Yangzhou University, Yangzhou 225009, China; [email protected] 4 Faculty of Animal Production, West Kordufan University, Alnuhud City 12942, Sudan; [email protected] * Correspondence: [email protected]; Tel.: +86-0514-87979269 Simple Summary: Feet and leg problems are among the major reasons for dairy cows leaving the herd, as well as having direct association with production and reproduction efficiency, health (e.g., claw disorders and lameness) and welfare. Hence, understanding the genetic architecture underlying feet and conformation traits in dairy cattle offers new opportunities toward the genetic improvement and long-term selection. Through a genome-wide association study on Chinese Holstein cattle, we identified several candidate genes associated with feet and leg conformation traits. These results could provide useful information about the molecular breeding basis of feet and leg traits, thus Citation: Abdalla, I.M.; Lu, X.; Nazar, improving the longevity and productivity of dairy cattle. -

Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands That Promote Axonal Growth
Research Article: New Research Development Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands that Promote Axonal Growth Jeremy S. Toma,1 Konstantina Karamboulas,1,ª Matthew J. Carr,1,2,ª Adelaida Kolaj,1,3 Scott A. Yuzwa,1 Neemat Mahmud,1,3 Mekayla A. Storer,1 David R. Kaplan,1,2,4 and Freda D. Miller1,2,3,4 https://doi.org/10.1523/ENEURO.0066-20.2020 1Program in Neurosciences and Mental Health, Hospital for Sick Children, 555 University Avenue, Toronto, Ontario M5G 1X8, Canada, 2Institute of Medical Sciences University of Toronto, Toronto, Ontario M5G 1A8, Canada, 3Department of Physiology, University of Toronto, Toronto, Ontario M5G 1A8, Canada, and 4Department of Molecular Genetics, University of Toronto, Toronto, Ontario M5G 1A8, Canada Abstract Peripheral nerves provide a supportive growth environment for developing and regenerating axons and are es- sential for maintenance and repair of many non-neural tissues. This capacity has largely been ascribed to paracrine factors secreted by nerve-resident Schwann cells. Here, we used single-cell transcriptional profiling to identify ligands made by different injured rodent nerve cell types and have combined this with cell-surface mass spectrometry to computationally model potential paracrine interactions with peripheral neurons. These analyses show that peripheral nerves make many ligands predicted to act on peripheral and CNS neurons, in- cluding known and previously uncharacterized ligands. While Schwann cells are an important ligand source within injured nerves, more than half of the predicted ligands are made by nerve-resident mesenchymal cells, including the endoneurial cells most closely associated with peripheral axons. At least three of these mesen- chymal ligands, ANGPT1, CCL11, and VEGFC, promote growth when locally applied on sympathetic axons.