Evolutionary Genomics of Human Intellectual Disability Bernard Crespi,1 Kyle Summers2 and Steve Dorus3
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Structure and Function of the Human Recq DNA Helicases
Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2005 Structure and function of the human RecQ DNA helicases Garcia, P L Posted at the Zurich Open Repository and Archive, University of Zurich ZORA URL: https://doi.org/10.5167/uzh-34420 Dissertation Published Version Originally published at: Garcia, P L. Structure and function of the human RecQ DNA helicases. 2005, University of Zurich, Faculty of Science. Structure and Function of the Human RecQ DNA Helicases Dissertation zur Erlangung der naturwissenschaftlichen Doktorw¨urde (Dr. sc. nat.) vorgelegt der Mathematisch-naturwissenschaftlichen Fakultat¨ der Universitat¨ Z ¨urich von Patrick L. Garcia aus Unterseen BE Promotionskomitee Prof. Dr. Josef Jiricny (Vorsitz) Prof. Dr. Ulrich H ¨ubscher Dr. Pavel Janscak (Leitung der Dissertation) Z ¨urich, 2005 For my parents ii Summary The RecQ DNA helicases are highly conserved from bacteria to man and are required for the maintenance of genomic stability. All unicellular organisms contain a single RecQ helicase, whereas the number of RecQ homologues in higher organisms can vary. Mu- tations in the genes encoding three of the five human members of the RecQ family give rise to autosomal recessive disorders called Bloom syndrome, Werner syndrome and Rothmund-Thomson syndrome. These diseases manifest commonly with genomic in- stability and a high predisposition to cancer. However, the genetic alterations vary as well as the types of tumours in these syndromes. Furthermore, distinct clinical features are observed, like short stature and immunodeficiency in Bloom syndrome patients or premature ageing in Werner Syndrome patients. Also, the biochemical features of the human RecQ-like DNA helicases are diverse, pointing to different roles in the mainte- nance of genomic stability. -

Open Full Page
CCR PEDIATRIC ONCOLOGY SERIES CCR Pediatric Oncology Series Recommendations for Childhood Cancer Screening and Surveillance in DNA Repair Disorders Michael F. Walsh1, Vivian Y. Chang2, Wendy K. Kohlmann3, Hamish S. Scott4, Christopher Cunniff5, Franck Bourdeaut6, Jan J. Molenaar7, Christopher C. Porter8, John T. Sandlund9, Sharon E. Plon10, Lisa L. Wang10, and Sharon A. Savage11 Abstract DNA repair syndromes are heterogeneous disorders caused by around the world to discuss and develop cancer surveillance pathogenic variants in genes encoding proteins key in DNA guidelines for children with cancer-prone disorders. Herein, replication and/or the cellular response to DNA damage. The we focus on the more common of the rare DNA repair dis- majority of these syndromes are inherited in an autosomal- orders: ataxia telangiectasia, Bloom syndrome, Fanconi ane- recessive manner, but autosomal-dominant and X-linked reces- mia, dyskeratosis congenita, Nijmegen breakage syndrome, sive disorders also exist. The clinical features of patients with DNA Rothmund–Thomson syndrome, and Xeroderma pigmento- repair syndromes are highly varied and dependent on the under- sum. Dedicated syndrome registries and a combination of lying genetic cause. Notably, all patients have elevated risks of basic science and clinical research have led to important in- syndrome-associated cancers, and many of these cancers present sights into the underlying biology of these disorders. Given the in childhood. Although it is clear that the risk of cancer is rarity of these disorders, it is recommended that centralized increased, there are limited data defining the true incidence of centers of excellence be involved directly or through consulta- cancer and almost no evidence-based approaches to cancer tion in caring for patients with heritable DNA repair syn- surveillance in patients with DNA repair disorders. -

FANCJ Regulates the Stability of FANCD2/FANCI Proteins and Protects Them from Proteasome and Caspase-3 Dependent Degradation
FANCJ regulates the stability of FANCD2/FANCI proteins and protects them from proteasome and caspase-3 dependent degradation Komaraiah Palle, Ph.D. (Kumar) Assistant Professor of Oncologic Sciences Abraham Mitchell Cancer Research Scholar Mitchell Cancer Institute University of South Alabama Outline • Fanconi anemia (FA) pathway • Role of FA pathway in Genome maintenance • FANCJ and FANCD2 functional relationship • FANCJ-mediated DDR in response to Fork-stalling Fanconi Anemia • Rare, inherited blood disorder. • 1:130,000 births Guido Fanconi 1892-1979 • Affects men and women equally. • Affects all racial and ethnic groups – higher incidence in Ashkenazi Jews and Afrikaners Birth Defects Fanconi anemia pathway • FA is a rare chromosome instability syndrome • Autosomal recessive disorder (or X-linked) • Developmental abnormalities • 17 complementation groups identified to date • FA pathway is involved in DNA repair • Increased cancer susceptibility - many patients develop AML - in adults solid tumors Fanconi Anemia is an aplastic anemia FA patients are prone to multiple types of solid tumors • Increased incidence and earlier onset cancers: oral cavity, GI and genital and reproductive tract head and neck breast esophagus skin liver brain Why? FA is a DNA repair disorder • FA caused by mutations in 17 genes: FANCA FANCF FANCM FANCB FANCG/XRCC9 FANCN/PALB2 FANCC FANCI RAD51C/FANCO FANCD1/BRCA2 FANCJ SLX4/FANCP FANCD2 FANCL ERCC2/XPF/FANCQ FANCE BRCA1/FANCS • FA genes function in DNA repair processes • FA patient cells are highly sensitive -

Fanconi Anemia, Bloom Syndrome and Breast Cancer
A multiprotein complex in DNA damage response network of Fanconi anemia, Bloom syndrome and Breast cancer Weidong Wang Lab of Genetics, NIA A Multi-protein Complex Connects Two Genomic Instability Diseases: Bloom Syndrome and Fanconi Anemia Bloom Syndrome . Genomic Instability: -sister-chromatid exchange . Cancer predisposition . Mutation in BLM, a RecQ DNA Helicase . BLM participates in: HR-dependent DSB repair Recovery of stalled replication forks . BLM works with Topo IIIa and RMI to Suppress crossover recombination Courtesy of Dr. Ian Hickson A Multi-protein Complex Connects Two Genomic Instability Diseases: Bloom Syndrome and Fanconi Anemia P I l o r t n o BLM IP kDa C HeLa BLAP 250 Nuclear Extract 200- BLM* FANCA* 116- TOPO IIIα* 97- BLAP 100 MLH1* BLM IP BLAP 75 * 66- RPA 70 IgG H 45- * 30- RPA32 IgG L 20- * 12- RPA14 Meetei et al. MCB 2003 A Multi-protein Complex Connects Two Genomic Instability Diseases: Bloom Syndrome and Fanconi Anemia P I A C N A F BLM IP HeLa FANCM= FAAP 250 BLAP 250 Nuclear Extract BLM* BLM* * FANCA* FANCA TOPO IIIα* TOPO IIIα* FAAP 100 BLAP 100 FANCB= FAAP 95 MLH1 FANCA IP BLM IP BLAP 75 BLAP 75 RPA70*/FANCG* RPA 70* FANCC*/FANCE* IgG H FANCL= FAAP 43 FANCF* RPA32* IgG L Meetei et al. MCB 2003 Meetei et al. Nat Genet. 2003, 2004, 2005 BRAFT-a Multisubunit Machine that Maintains Genome Stability and is defective in Fanconi anemia and Bloom syndrome BRAFT Super-complex Fanconi Anemia Bloom Syndrome Core Complex Complex 12 polypeptides 7 polypeptides FANCA BLM Helicase (HJ, fork, D-loop), fork FANCC regression, dHJ dissolution Topo IIIα Topoisomerase, FANCE dHJ dissolution FANCF BLAP75 RMI1 FANCG Stimulates dHJ dissolution. -

HEREDITARY CANCER PANELS Part I
Pathology and Laboratory Medicine Clinic Building, K6, Core Lab, E-655 2799 W. Grand Blvd. HEREDITARY CANCER PANELS Detroit, MI 48202 855.916.4DNA (4362) Part I- REQUISITION Required Patient Information Ordering Physician Information Name: _________________________________________________ Gender: M F Name: _____________________________________________________________ MRN: _________________________ DOB: _______MM / _______DD / _______YYYY Address: ___________________________________________________________ ICD10 Code(s): _________________/_________________/_________________ City: _______________________________ State: ________ Zip: __________ ICD-10 Codes are required for billing. When ordering tests for which reimbursement will be sought, order only those tests that are medically necessary for the diagnosis and treatment of the patient. Phone: _________________________ Fax: ___________________________ Billing & Collection Information NPI: _____________________________________ Patient Demographic/Billing/Insurance Form is required to be submitted with this form. Most genetic testing requires insurance prior authorization. Due to high insurance deductibles and member policy benefits, patients may elect to self-pay. Call for more information (855.916.4362) Bill Client or Institution Client Name: ______________________________________________________ Client Code/Number: _____________ Bill Insurance Prior authorization or reference number: __________________________________________ Patient Self-Pay Call for pricing and payment options Toll -

MECHANISMS in ENDOCRINOLOGY: Novel Genetic Causes of Short Stature
J M Wit and others Genetics of short stature 174:4 R145–R173 Review MECHANISMS IN ENDOCRINOLOGY Novel genetic causes of short stature 1 1 2 2 Jan M Wit , Wilma Oostdijk , Monique Losekoot , Hermine A van Duyvenvoorde , Correspondence Claudia A L Ruivenkamp2 and Sarina G Kant2 should be addressed to J M Wit Departments of 1Paediatrics and 2Clinical Genetics, Leiden University Medical Center, PO Box 9600, 2300 RC Leiden, Email The Netherlands [email protected] Abstract The fast technological development, particularly single nucleotide polymorphism array, array-comparative genomic hybridization, and whole exome sequencing, has led to the discovery of many novel genetic causes of growth failure. In this review we discuss a selection of these, according to a diagnostic classification centred on the epiphyseal growth plate. We successively discuss disorders in hormone signalling, paracrine factors, matrix molecules, intracellular pathways, and fundamental cellular processes, followed by chromosomal aberrations including copy number variants (CNVs) and imprinting disorders associated with short stature. Many novel causes of GH deficiency (GHD) as part of combined pituitary hormone deficiency have been uncovered. The most frequent genetic causes of isolated GHD are GH1 and GHRHR defects, but several novel causes have recently been found, such as GHSR, RNPC3, and IFT172 mutations. Besides well-defined causes of GH insensitivity (GHR, STAT5B, IGFALS, IGF1 defects), disorders of NFkB signalling, STAT3 and IGF2 have recently been discovered. Heterozygous IGF1R defects are a relatively frequent cause of prenatal and postnatal growth retardation. TRHA mutations cause a syndromic form of short stature with elevated T3/T4 ratio. Disorders of signalling of various paracrine factors (FGFs, BMPs, WNTs, PTHrP/IHH, and CNP/NPR2) or genetic defects affecting cartilage extracellular matrix usually cause disproportionate short stature. -

Epigenetic Regulation of DNA Repair Genes and Implications for Tumor Therapy ⁎ ⁎ Markus Christmann , Bernd Kaina
Mutation Research-Reviews in Mutation Research xxx (xxxx) xxx–xxx Contents lists available at ScienceDirect Mutation Research-Reviews in Mutation Research journal homepage: www.elsevier.com/locate/mutrev Review Epigenetic regulation of DNA repair genes and implications for tumor therapy ⁎ ⁎ Markus Christmann , Bernd Kaina Department of Toxicology, University of Mainz, Obere Zahlbacher Str. 67, D-55131 Mainz, Germany ARTICLE INFO ABSTRACT Keywords: DNA repair represents the first barrier against genotoxic stress causing metabolic changes, inflammation and DNA repair cancer. Besides its role in preventing cancer, DNA repair needs also to be considered during cancer treatment Genotoxic stress with radiation and DNA damaging drugs as it impacts therapy outcome. The DNA repair capacity is mainly Epigenetic silencing governed by the expression level of repair genes. Alterations in the expression of repair genes can occur due to tumor formation mutations in their coding or promoter region, changes in the expression of transcription factors activating or Cancer therapy repressing these genes, and/or epigenetic factors changing histone modifications and CpG promoter methylation MGMT Promoter methylation or demethylation levels. In this review we provide an overview on the epigenetic regulation of DNA repair genes. GADD45 We summarize the mechanisms underlying CpG methylation and demethylation, with de novo methyl- TET transferases and DNA repair involved in gain and loss of CpG methylation, respectively. We discuss the role of p53 components of the DNA damage response, p53, PARP-1 and GADD45a on the regulation of the DNA (cytosine-5)- methyltransferase DNMT1, the key enzyme responsible for gene silencing. We stress the relevance of epigenetic silencing of DNA repair genes for tumor formation and tumor therapy. -
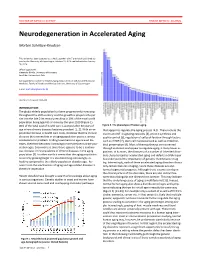
Neurodegeneration in Accelerated Aging
DOCTOR OF MEDICAL SCIENCE DANISH MEDICAL JOURNAL Neurodegeneration in Accelerated Aging Morten Scheibye-Knudsen This review has been accepted as a thesis together with 7 previously published pa- pers by the University of Copenhagen, October 16, 2014 and defended on January 14, 2016 Official opponents: Alexander Bürkle, University of Konstanz Lars Eide, University of Oslo Correspondence: Center for Healthy Aging, Department of Cellular and Molecular Medicine, Faculty of Health and Medical Sciences, University of Copenhagen E-mail: [email protected] Dan Med J 2016;63(11):B5308 INTRODUCTION The global elderly population has been progressively increasing throughout the 20th century and this growth is projected to per- sist into the late 21st century resulting in 20% of the total world population being aged 65 or more by the year 2100 (Figure 1). 80% of the total cost of health care is accrued after 40 years of Figure 2. The phenotype of human aging. age where chronic diseases become prevalent [1, 2]. With an ex- that appear to regulate the aging process [4,5]. These include the ponential increase in health care costs, it follows that the chronic insulin and IGF-1 signaling cascades [4], protein synthesis and diseases that accumulate in an aging population poses a serious quality control [6], regulation of cell proliferation through factors socioeconomic problem. Finding treatments to age related dis- such as mTOR [7], stem cell maintenance 8 as well as mitochon- eases, therefore becomes increasingly more pertinent as the pop- drial preservation [9]. Most of these pathways are conserved ulation ages. Even more so since there appears to be a continu- through evolution and appear to regulate aging in many lower or- ous increase in the prevalence of chronic diseases in the aging ganisms. -

Posttranslational Regulation of the Fanconi Anemia Cancer Susceptibility Pathway
DNA repair interest group 04/21/15 Posttranslational Regulation of the Fanconi Anemia Cancer Susceptibility Pathway Assistant professor Department of Pharmacological Sciences Stony Brook University Hyungjin Kim, Ph.D. DNA Repair System Preserves Genome Stability DNA DAMAGE X-rays DNA DAMAGE Oxygen radicals UV light X-rays RESPONSE Alkylating agents Polycyclic aromatic Anti-tumor agents Replication Spontaneous reactions hydrocarbons (cisplatin, MMC) errors Cell-cycle arrest Cell death Uracil DNA REPAIR Abasic site (6-4)PP Interstrand cross-link Base pair mismatch DEFECT 8-Oxiguanine Bulky adduct Double-strand break Insertion/deletion Single-strand break CPD Cancer • Mutations • Chromosome Base- Nucleotide- Fanconi anemia (FA) aberrations Mismatch excision repair excision repair pathway repair (BER) (NER) HR/NHEJ DNA REPAIR Adapted from Hoeijmakers (2001) Nature DNA Repair Mechanism as a Tumor Suppressor Network Cancer is a disease of DNA repair Environmental Mutagen Oncogenic Stress • Hereditary cancers: mutations in DNA repair genes (BRCA1, BRCA2, MSH2, etc) DNA damage response Checkpoint, • Sporadic cancers: DNA repair - Oncogene-induced replication stress - Inaccurate DNA repair caused by selection of somatic mutations in DNA repair genes Tumorigenesis Genome instability Germ-line and Somatic Disruption of DNA Repair Is Prevalent in Cancer The Cancer Genome Atlas (TCGA): Genomic characterization and sequence analysis of various tumors from patient samples TCGA network, (2011) Nature Clinical and Cellular Features of Fanconi Anemia (FA) -

Deletion and Reduced Expression of the Fanconi Anemia FANCA Gene in Sporadic Acute Myeloid Leukemia
Leukemia (2004) 18, 420–425 & 2004 Nature Publishing Group All rights reserved 0887-6924/04 $25.00 www.nature.com/leu Deletion and reduced expression of the Fanconi anemia FANCA gene in sporadic acute myeloid leukemia MD Tischkowitz1, NV Morgan1, D Grimwade1,2, C Eddy1, S Ball3, I Vorechovsky4, S Langabeer2, R Sto¨ger1, SV Hodgson1 and CG Mathew1 1Department of Medical and Molecular Genetics, Division of Genetics and Development Guy’s, King’s and St Thomas’ School of Medicine, King’s College London, Guy’s Hospital, London, UK; 2Department of Haematology, University College London, London, UK; 3Department of Haematology, St George’s Hospital, London, UK; and 4Human Genetics Division, School of Medicine, Southampton University Hospital, Tremona Road, Southampton, UK Fanconi anemia (FA) is an autosomal recessive chromosomal Since the incidence of AML is highly elevated in FA patients, instability disorder caused by mutations in one of seven known it is possible that inherited or acquired mutations in the FA genes genes (FANCA,C,D2,E,F,G and BRCA2). Mutations in the FANCA gene are the most prevalent, accounting for two-thirds contribute to the etiology of sporadic AML. Epidemiological of FA cases. Affected individuals have greatly increased risks studies assessing incidence of malignancies (including AML) in of acute myeloid leukemia (AML). This raises the question as to FA families have failed to demonstrate an increased incidence in whether inherited or acquired mutations in FA genes might be heterozygous individuals.14,15 However, it is conceivable that involved in the development of sporadic AML. Quantitative genomic instability may result from a second somatic mutation fluorescent PCR was used to screen archival DNA from in an FA carrier or from one or two somatic mutations in a sporadic AML cases for FANCA deletions, which account for 16 40% of FANCA mutations in FA homozygotes. -

Overexpression of the Fanconi Anemia a Gene in Hela and MCF10A Cells
Korean J Hematol Vol. 41, No. 1, March, 2006 □ Original Article □ Overexpression of the Fanconi Anemia A Gene in Hela and MCF10A Cells Woo Hyun Park, Ph.D. Department of Physiology, Chonbuk National University Medical School, Jeonju, Korea Background: Fanconi Anemia (FA) is an autosomal recessive inherited disease, which is characterized by developmental abnormalities, progressive bone marrow failure and a predisposition to cancer. The phenotypes of FA cells show extreme sensitivities towards oxygen and DNA cross linking agents, such as diepoxybutane and mitomycin C (MMC). Methods: In the current study, retroviruses expressing the FANCA gene were prepared to create the stable cell lines, Hela (cervical carcinoma) and MCF10A (breast). The expression of FANCA protein in the Hela and MCF10A stable cells, following puromycin selection, was checked using Western blot. The difference in the cell growth between the parent and FANCA expressing cells following MMC treatment was checked using the MTT assay. Results: The expression of exogenous FANCA protein in the Hela and MCF10A stable cells was observed using Western blot. The MCF10A cells expressing exogenous FANCA were resistant to MMC concentrations with the range 0.01~1µM compared with the MCF10 parent cells. However, at an MMC concentration of 10µM, there was no difference in the susceptibility between the parent and FANCA expressing MCF10 cells. The Hela cells expressing FANCA showed no resistance at any MMC con- centration (0.01~10µM). Conclusion: FANCA protein is an important factor for -

Distinct Roles of BRCA2 in Replication Fork Protection in Response to Hydroxyurea and DNA Interstrand Cross-Links
Downloaded from genesdev.cshlp.org on October 1, 2021 - Published by Cold Spring Harbor Laboratory Press Distinct roles of BRCA2 in replication fork protection in response to hydroxyurea and DNA interstrand cross-links Kimberly A. Rickman,1 Raymond J. Noonan,1 Francis P. Lach,1 Sunandini Sridhar,1 Anderson T. Wang,1,5 Avinash Abhyankar,2 Athena Huang,1 Michael Kelly,3 Arleen D. Auerbach,4 and Agata Smogorzewska1 1Laboratory of Genome Maintenance, The Rockefeller University, New York, New York 10065, USA; 2New York Genome Center, New York, New York 10013, USA; 3Tufts Medical Center, Boston, Massachusetts 02111, USA; 4Human Genetics and Hematology, The Rockefeller University, New York, New York 10065, USA DNA interstrand cross-links (ICLs) are a form of DNA damage that requires the interplay of a number of repair proteins including those of the Fanconi anemia (FA) and the homologous recombination (HR) pathways. Pathogenic variants in the essential gene BRCA2/FANCD1, when monoallelic, predispose to breast and ovarian cancer, and when biallelic, result in a severe subtype of Fanconi anemia. BRCA2 function in the FA pathway is attributed to its role as a mediator of the RAD51 recombinase in HR repair of programmed DNA double-strand breaks (DSB). BRCA2 and RAD51 functions are also required to protect stalled replication forks from nucleolytic degradation during re- sponse to hydroxyurea (HU). While RAD51 has been shown to be necessary in the early steps of ICL repair to prevent aberrant nuclease resection, the role of BRCA2 in this process has not been described. Here, based on the analysis of BRCA2 DNA-binding domain (DBD) mutants (c.8488-1G>A and c.8524C>T) discovered in FA patients presenting with atypical FA-like phenotypes, we establish that BRCA2 is necessary for the protection of DNA at ICLs.