Global, Regional, and National Cancer Incidence, Mortality, Years
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

World Bank Document
CONFORMED COPY Public Disclosure Authorized LOAN NUMBER 7616-CN Loan Agreement Public Disclosure Authorized (Wenchuan Earthquake Recovery Project) between PEOPLE’S REPUBLIC OF CHINA Public Disclosure Authorized and INTERNATIONAL BANK FOR RECONSTRUCTION AND DEVELOPMENT Dated March 20, 2009 Public Disclosure Authorized LOAN AGREEMENT AGREEMENT dated March 20, 2009, between PEOPLE’S REPUBLIC OF CHINA (“Borrower”) and INTERNATIONAL BANK FOR RECONSTRUCTION AND DEVELOPMENT (“Bank”). The Borrower and the Bank hereby agree as follows: ARTICLE I – GENERAL CONDITIONS; DEFINITIONS 1.01. The General Conditions (as defined in the Appendix to this Agreement) constitute an integral part of this Agreement. 1.02. Unless the context requires otherwise, the capitalized terms used in the Loan Agreement have the meanings ascribed to them in the General Conditions or in the Appendix to this Agreement. ARTICLE II – LOAN 2.01. The Bank agrees to lend to the Borrower, on the terms and conditions set forth or referred to in this Agreement, an amount equal to seven hundred ten million Dollars ($710,000,000), as such amount may be converted from time to time through a Currency Conversion in accordance with the provisions of Section 2.07 of this Agreement (“Loan”), to assist in financing the project described in Schedule 1 to this Agreement (“Project”). 2.02. The Borrower may withdraw the proceeds of the Loan in accordance with Section IV of Schedule 2 to this Agreement. 2.03. The Front-end Fee payable by the Borrower shall be equal to one quarter of one percent (0.25%) of the Loan amount. The Borrower shall pay the Front-end Fee not later than sixty (60) days after the Effective Date. -

Analysis of Accumulation Models of Middle Permian in Northwest Sichuan Basin
EARTH SCIENCES RESEARCH JOURNAL GEOLOGY Earth Sci. Res. J. Vol. 24, No. 4 (December, 2020): 419-428 Analysis of accumulation models of middle Permian in Northwest Sichuan Basin Bin Li1,2,*, Qiqi Li3, Wenhua Mei2, Qingong Zhuo4, Xuesong Lu4 1State Key Laboratory of Shale Oil and Gas Enrichment Mechanisms and Effective Development, Wuxi, 214126 2School of Geoscience and Technology, Southwest Petroleum University, Chengdu, 610500 3School of Earth Resources, China University of Geosciences, Wuhan, 430074; 4PetroChina Research Institute of Petroleum Exploration and Development, Beijing 100083, China *Corresponding author: [email protected] ABSTRACT Keywords: Northwest of Sichuan Basin; Great progress has been made in middle Permian exploration in Northwest Sichuan in recent years, but there are still Structure Evolution of Foreland Basin; Middle many questions in understanding the hydrocarbon accumulation conditions. Due to the abundance of source rocks and Permian; Paleozoic; Accumulation conditions; the multi-term tectonic movements in this area, the hydrocarbon accumulation model is relatively complex, which has Accumulation Model. become the main problem to be solved urgently in oil and gas exploration. Based on the different tectonic backgrounds of the middle Permian in northwest Sichuan Basin, the thrust nappe belt, the hidden front belt, and the depression belt are taken as the research units to comb and compare the geologic conditions of the middle Permian reservoir. The evaluation of source rocks and the comparison of hydrocarbon sources suggest that the middle Permian hydrocarbon mainly comes from the bottom of the lower Cambrian and middle Permian, and the foreland orogeny promoted the thermal evolution of Paleozoic source rocks in northwest Sichuan to high maturity and over maturity stage. -

Research on the Space Transformation of Yungang Grottoes Art Heritage and the Design of Wisdom Museum from the Perspective of Digital Humanity
E3S Web of Conferences 23 6 , 05023 (2021) https://doi.org/10.1051/e3sconf/202123605023 ICERSD 2020 Research on the Space transformation of Yungang Grottoes Art Heritage and the Design of wisdom Museum from the perspective of digital Humanity LiuXiaoDan1, 2, XiaHuiWen3 1Jinzhong college, Academy of fine arts, Jinzhong ShanXi, 030619 2Nanjing University, School of History, Nanjing JiangSu, 210023 3Taiyuan University of Technology Academy of arts, Jinzhong ShanXi 030606 *Corresponding author: LiuXiaoDan, No. 199 Wenhua Street, Yuci District, Jinzhong City, Shanxi Province, 030619, China. ABSTRACT: The pace transformation and innovative design of Yungang art heritage should keep pace with times on the setting of “digital humanity” times, make the most of new research approaches which is given by “digital humanity” and explore a new way of pace transformation of art heritage actively. The research object of this topic is lineage master in space and form and the transformation of promotion and Creativity of Yungang Grotto art heritage. The goal is taking advantage of big data basics and the information sample collection and integration in the context of the full media era. Transferring Yungang Grotto art materially and creatively by making use of the world's advanced "art + science and technology" means, building a new type of modern sapiential museum and explore the construction mode of it and the upgraded version of modern educational functions. approaches which is given by “digital humanity” and explore a new way of pace transformation of art heritage 1 Introduction actively. During the long race against the "disappearance" Yungang Grotto represent the highest level of north royal of cultural relics, the design of the wise exhibition grotto art in the 5th century AD. -

Research on the Organizational Forms and Development Status of Anhui's Agricultural Industrialization Management
Academic Journal of Humanities & Social Sciences ISSN 2616-5783 Vol.4, Issue 3: 67-70, DOI: 10.25236/AJHSS.2021.040314 Research on the Organizational Forms and Development Status of Anhui's Agricultural Industrialization Management Qiuxuan Wan Anhui University of Finance and Economics, Bengbu 233000, China Abstract: Agricultural industrialization is an important measure to promote the modernization of agriculture and promote the effective supply of agricultural products in the society, revealing the relationship between agricultural industrialization and the relationship between agriculture and related industries. Based on the statistical data of agricultural industrialization in the 16 prefecture-level cities in Anhui Province, this article analyzes and evaluates the development of agricultural industrialization based on the form of agricultural industrialization management organization, and puts forward corresponding suggestions for the existing development pain points, with the intention to provide policy enlightenment for the sustainable development of agricultural industrialization in Anhui Province. Keywords: Agricultural industrialization, Organizational business form, Cooperative, Agricultural industry chain 1. Introduction Agricultural industrialization is essentially the implementation of technological changes to traditional agriculture, through the advancement of agricultural science and technology to promote the progress of agriculture and its related industries, and the self-development of agriculture in modern business methods and industrial organization forms. In the early 1990s, the business model of agricultural industrialization emerged in China, and now it has basically realized the form of intensive production. Domestic scholars' research on agricultural industrialization is mostly based on the consideration of theories and systems, and the reference of foreign research results, combined with current political methods to conduct more in-depth research on theoretical issues. -

World Bank-Financed Anhui Aged Care
SFG3798 REV Zhongzi Huayu REV RR RREV Public Disclosure Authorized G. H. P. Z. J. Zi No. 1051 World Bank-financed Anhui Aged Care System Demonstration Project Public Disclosure Authorized Environment and Social Management Framework (ESMF) Public Disclosure Authorized Commissioned by: Department of Civil Affairs of Anhui Public Disclosure Authorized Province Prepared by: Beijing Zhongzi Huayu Environmental Protection Technology Co., Ltd. Prepared in: December 2017 Table of Contents I. Introduction and Objectives........................................................................................ 3 II. Project Overview ....................................................................................................... 3 III. Policy Framework of Environmental and Social Problems ..................................... 5 IV. Paths of Solving Environmental and Social Problems ........................................... 11 4.1 The first step: identify sub-projects according to project selection criteria ... 11 4.2 The second step: screen potential environmental and social impacts ............ 11 4.3 The third step: review the screening results ................................................... 13 4.4 The fourth step: prepare the safeguard documents and have public consultation and information disclosure .............................................................. 14 4.5 The fifth step: review and approve the safeguard documents ........................ 15 4.6 The sixth step: implement, supervise, monitor and assess the approved actions -

ATTACHMENT 1 Barcode:3800584-02 C-570-107 INV - Investigation
ATTACHMENT 1 Barcode:3800584-02 C-570-107 INV - Investigation - Chinese Producers of Wooden Cabinets and Vanities Company Name Company Information Company Name: A Shipping A Shipping Street Address: Room 1102, No. 288 Building No 4., Wuhua Road, Hongkou City: Shanghai Company Name: AA Cabinetry AA Cabinetry Street Address: Fanzhong Road Minzhong Town City: Zhongshan Company Name: Achiever Import and Export Co., Ltd. Street Address: No. 103 Taihe Road Gaoming Achiever Import And Export Co., City: Foshan Ltd. Country: PRC Phone: 0757-88828138 Company Name: Adornus Cabinetry Street Address: No.1 Man Xing Road Adornus Cabinetry City: Manshan Town, Lingang District Country: PRC Company Name: Aershin Cabinet Street Address: No.88 Xingyuan Avenue City: Rugao Aershin Cabinet Province/State: Jiangsu Country: PRC Phone: 13801858741 Website: http://www.aershin.com/i14470-m28456.htmIS Company Name: Air Sea Transport Street Address: 10F No. 71, Sung Chiang Road Air Sea Transport City: Taipei Country: Taiwan Company Name: All Ways Forwarding (PRe) Co., Ltd. Street Address: No. 268 South Zhongshan Rd. All Ways Forwarding (China) Co., City: Huangpu Ltd. Zip Code: 200010 Country: PRC Company Name: All Ways Logistics International (Asia Pacific) LLC. Street Address: Room 1106, No. 969 South, Zhongshan Road All Ways Logisitcs Asia City: Shanghai Country: PRC Company Name: Allan Street Address: No.188, Fengtai Road City: Hefei Allan Province/State: Anhui Zip Code: 23041 Country: PRC Company Name: Alliance Asia Co Lim Street Address: 2176 Rm100710 F Ho King Ctr No 2 6 Fa Yuen Street Alliance Asia Co Li City: Mongkok Country: PRC Company Name: ALMI Shipping and Logistics Street Address: Room 601 No. -

Yunnan Sustainable Road Maintenance (Sector) Project
Environmental Monitoring Report Project Number: 45030-002 July 2016 PRC: Yunnan Sustainable Road Maintenance (Sector) Project Prepared by the Yunnan Highway Administration Bureau for the People’s Republic of China and the Asian Development Bank This environmental monitoring report is a document of the borrower. The views expressed herein do not necessarily represent those of ADB's Board of Directors, Management, or staff, and may be preliminary in nature. In preparing any country program or strategy, financing any project, or by making any designation of or reference to a particular territory or geographic area in this document, the Asian Development Bank does not intend to make any judgments as to the legal or other status of any territory or area. Asian Development Bank PRC-3074: Yunnan Sustainable Road Maintenance (Sector) Project The 2nd Quarterly Environmental Monitoring Report in 2016 Drafted in July 2016 For phase I and phase II-Maintenance Components Prepared by the Yunnan Highway Administration Bureau for the Asian Development Bank Table of Contents LIST OF FIGURES, TABLES AND PICTURES II EXECUTIVE SUMMARY VII I. BACKGROUND AND PROJECT PROGRESS 1 II. REPORT PREPARATION 4 A. Purposes and Objectives 4 B. Policy, Legal, and Administrative Framework 4 C. Report Period 5 D. Scope of Environment Monitoring 5 IV. IMPLEMENTATION OF EMP 6 A. Detailed design drawings and bidding documents 6 B. Institutional Arrangements 6 C. Construction Supervisors 6 D. Implementation status of the mitigation measures 7 E. Analysis of Environmental Monitoring Results 10 F. Institutional Capacity Building and Training 16 V. PUBLIC CONSULTATION, INFORMATION DISCLOSURE AND GRIEVANCE REDRESS MECHANISM 18 A. -
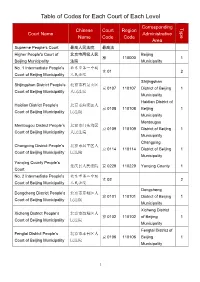
Table of Codes for Each Court of Each Level
Table of Codes for Each Court of Each Level Corresponding Type Chinese Court Region Court Name Administrative Name Code Code Area Supreme People’s Court 最高人民法院 最高法 Higher People's Court of 北京市高级人民 Beijing 京 110000 1 Beijing Municipality 法院 Municipality No. 1 Intermediate People's 北京市第一中级 京 01 2 Court of Beijing Municipality 人民法院 Shijingshan Shijingshan District People’s 北京市石景山区 京 0107 110107 District of Beijing 1 Court of Beijing Municipality 人民法院 Municipality Haidian District of Haidian District People’s 北京市海淀区人 京 0108 110108 Beijing 1 Court of Beijing Municipality 民法院 Municipality Mentougou Mentougou District People’s 北京市门头沟区 京 0109 110109 District of Beijing 1 Court of Beijing Municipality 人民法院 Municipality Changping Changping District People’s 北京市昌平区人 京 0114 110114 District of Beijing 1 Court of Beijing Municipality 民法院 Municipality Yanqing County People’s 延庆县人民法院 京 0229 110229 Yanqing County 1 Court No. 2 Intermediate People's 北京市第二中级 京 02 2 Court of Beijing Municipality 人民法院 Dongcheng Dongcheng District People’s 北京市东城区人 京 0101 110101 District of Beijing 1 Court of Beijing Municipality 民法院 Municipality Xicheng District Xicheng District People’s 北京市西城区人 京 0102 110102 of Beijing 1 Court of Beijing Municipality 民法院 Municipality Fengtai District of Fengtai District People’s 北京市丰台区人 京 0106 110106 Beijing 1 Court of Beijing Municipality 民法院 Municipality 1 Fangshan District Fangshan District People’s 北京市房山区人 京 0111 110111 of Beijing 1 Court of Beijing Municipality 民法院 Municipality Daxing District of Daxing District People’s 北京市大兴区人 京 0115 -

Annual Report 2019
HAITONG SECURITIES CO., LTD. 海通證券股份有限公司 Annual Report 2019 2019 年度報告 2019 年度報告 Annual Report CONTENTS Section I DEFINITIONS AND MATERIAL RISK WARNINGS 4 Section II COMPANY PROFILE AND KEY FINANCIAL INDICATORS 8 Section III SUMMARY OF THE COMPANY’S BUSINESS 25 Section IV REPORT OF THE BOARD OF DIRECTORS 33 Section V SIGNIFICANT EVENTS 85 Section VI CHANGES IN ORDINARY SHARES AND PARTICULARS ABOUT SHAREHOLDERS 123 Section VII PREFERENCE SHARES 134 Section VIII DIRECTORS, SUPERVISORS, SENIOR MANAGEMENT AND EMPLOYEES 135 Section IX CORPORATE GOVERNANCE 191 Section X CORPORATE BONDS 233 Section XI FINANCIAL REPORT 242 Section XII DOCUMENTS AVAILABLE FOR INSPECTION 243 Section XIII INFORMATION DISCLOSURES OF SECURITIES COMPANY 244 IMPORTANT NOTICE The Board, the Supervisory Committee, Directors, Supervisors and senior management of the Company warrant the truthfulness, accuracy and completeness of contents of this annual report (the “Report”) and that there is no false representation, misleading statement contained herein or material omission from this Report, for which they will assume joint and several liabilities. This Report was considered and approved at the seventh meeting of the seventh session of the Board. All the Directors of the Company attended the Board meeting. None of the Directors or Supervisors has made any objection to this Report. Deloitte Touche Tohmatsu (Deloitte Touche Tohmatsu and Deloitte Touche Tohmatsu Certified Public Accountants LLP (Special General Partnership)) have audited the annual financial reports of the Company prepared in accordance with PRC GAAP and IFRS respectively, and issued a standard and unqualified audit report of the Company. All financial data in this Report are denominated in RMB unless otherwise indicated. -

11D Yunnan Yuxi Pu-Er Xishuang-Banna Tengchong Ruili
1 Park Road, #03-09 People’s Park Complex, Singapore 059108 Tel: 6438 7478 Fax: 6438 9794 www.newaytravel.com.sg Exclusive of Neway Travel 11D Yunnan Yuxi Pu-Er Xishuang-Banna Tengchong Ruili Colorful Ethnic Experience Tour TOUR CODE: KMG11M 【Colorful Ethnic Experience】: To enter deeply into Dai,Naxi,Bai,Bulang,Wa Peoples Culture,Language, Costumes and ect. 【High Light】:Yuxi,Pu-Er,Xishuang Banna,Cangyuan,Tengchong,Ruili 【Local Flavor】:Steam Chicken+ Guoqiao Rice Noodle,Tengchong Local Style,Dai Flavor 【Shopping】: Yunnan Puer Tea Shop,Jade Shop Tengchong. D1:SIN KMG/YUXI (L/D) Accommodation:Gaodeng Hotel or similar 4Star Assemble at Singapore Changi Airport for our flight to Kunming, capital of Yunnan Province. Upon arrival, transfer to Yuxi. Yuxi Cigarette Company (Hongta Group) has become a famous cigarette making company. Its product brands such as Hongtashan, Ashima and Hongmei have been popular for many years in China. Upon arrival, the area also has beautiful natural sceneries and colorful cultures. Upon arrival, visit Hongta District. Next proceed to Pu-Er. : ( ) : D2 Pu-Er/Xishuang-Banna B/L/D Accommodation King Land Hotel or similar 4Star Pu'er is the birthplace of Pu Er Tea, a very famous international tea brand. Tea culture is an attractive tourist factor of Puer City visit the Pr-Er Tea Exhibition Garden. The garden is built on the Yinpan Mountain,29 km from downtown Puer, Against the background of a vast tea garden, it is made up of Pu'er Tea Museum, Pu'er Tea Making workshop, tea-picking zone, tea appraisal garden and some other facilities to comprehensively show the history of Pu'er Tea, and how to plant and produce as well. -

Evaluating the Influence of Land Use and Land Cover Change on Fine
www.nature.com/scientificreports OPEN Evaluating the infuence of land use and land cover change on fne particulate matter Wei Yang1* & Xiaoli Jiang2 Fine particulate matter (i.e. particles with diameters smaller than 2.5 microns, PM2.5) has become a critical environmental issue in China. Land use and land cover (LULC) is recognized as one of the most important infuence factors, however very fewer investigations have focused on the impact of LULC on PM2.5. The infuences of diferent LULC types and diferent land use and land cover change (LULCC) types on PM2.5 are discussed. A geographically weighted regression model is used for the general analysis, and a spatial analysis method based on the geographic information system is used for a detailed analysis. The results show that LULCC has a stable infuence on PM2.5 concentration. For diferent LULC types, construction lands have the highest PM2.5 concentration and woodlands have the lowest. The order of PM2.5 concentration for the diferent LULC types is: construction lands > unused lands > water > farmlands >grasslands > woodlands. For diferent LULCC types, when high-grade land types are converted to low-grade types, the PM2.5 concentration decreases; otherwise, the PM2.5 concentration increases. The result of this study can provide a decision basis for regional environmental protection and regional ecological security agencies. With the rapid development of China’s economy and society, its rate of urbanization is accelerating. China’s industrial scale is also expanding rapidly, and the problem of air pollution is becoming increasingly serious, which has a tremendous impact on the environment, economic development, and even people’s health 1. -

Land Use Change in Hongta District in Yuxi City, China Based on Archived Landsat Data of the Past 30 Years
Journal of Computer and Communications, 2018, 6, 138-145 http://www.scirp.org/journal/jcc ISSN Online: 2327-5227 ISSN Print: 2327-5219 Land Use Change in Hongta District in Yuxi City, China Based on Archived Landsat Data of the Past 30 Years Shanyu Zhou, Chaonan Ji, Ran Kang, Hermann Kaufmann Navigation and Remote Sensing Group, Institute of Space Science, Shandong University, Weihai, China How to cite this paper: Zhou, S.Y., Ji, Abstract C.N., Kang, R. and Kaufmann, H. (2018) Land Use Change in Hongta District in Urban expansion, which has affected ecosystem functioning and services at Yuxi City, China Based on Archived Land- local to global scale, is projected to have aggravating impacts on landscape in sat Data of the Past 30 Years. Journal of the future. A detailed study on spatiotemporal patterns of Hongta District in Computer and Communications, 6, 138-145. https://doi.org/10.4236/jcc.2018.611013 Yuxi City over a relatively long timeframe was conducted using mul- ti-temporal Landsat TM, ETM+ and OLI data of 1987-2017 associated with Received: October 27, 2018 Geographic Information System techniques and landscape analysis ap- Accepted: November 12, 2018 proaches. Results showed that sealed surfaces (urban area) have expanded Published: November 19, 2018 from 1.4% to 11.8%, with an annually increasing rate of 7.63%. Trees, farm areas and barren lands all underwent a slight reduction during this time pe- riod, whereby 23.8% of farms and 9.8% of bare lands transferred into sealed surfaces. Focusing on the urban expansion pattern, it presents a uninuclear polarizing polygon mode illustrating the urbanization sprawl over time to all directions.