CPTC-MAPK3-1 (CAB079934) Immunohistochemistry
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Information Material and Methods
MCT-11-0474 BKM120: a potent and specific pan-PI3K inhibitor Supplementary Information Material and methods Chemicals The EGFR inhibitor NVP-AEE788 (Novartis), the Jak inhibitor I (Merck Calbiochem, #420099) and anisomycin (Alomone labs, # A-520) were prepared as 50 mM stock solutions in 100% DMSO. Doxorubicin (Adriablastin, Pfizer), EGF (Sigma Ref: E9644), PDGF (Sigma, Ref: P4306) and IL-4 (Sigma, Ref: I-4269) stock solutions were prepared as recommended by the manufacturer. For in vivo administration: Temodal (20 mg Temozolomide capsules, Essex Chemie AG, Luzern) was dissolved in 4 mL KZI/glucose (20/80, vol/vol); Taxotere was bought as 40 mg/mL solution (Sanofi Aventis, France), and prepared in KZI/glucose. Antibodies The primary antibodies used were as follows: anti-S473P-Akt (#9271), anti-T308P-Akt (#9276,), anti-S9P-GSK3β (#9336), anti-T389P-p70S6K (#9205), anti-YP/TP-Erk1/2 (#9101), anti-YP/TP-p38 (#9215), anti-YP/TP-JNK1/2 (#9101), anti-Y751P-PDGFR (#3161), anti- p21Cip1/Waf1 (#2946), anti-p27Kip1 (#2552) and anti-Ser15-p53 (#9284) antibodies were from Cell Signaling Technologies; anti-Akt (#05-591), anti-T32P-FKHRL1 (#06-952) and anti- PDGFR (#06-495) antibodies were from Upstate; anti-IGF-1R (#SC-713) and anti-EGFR (#SC-03) antibodies were from Santa Cruz; anti-GSK3α/β (#44610), anti-Y641P-Stat6 (#611566), anti-S1981P-ATM (#200-301), anti-T2609 DNA-PKcs (#GTX24194) and anti- 1 MCT-11-0474 BKM120: a potent and specific pan-PI3K inhibitor Y1316P-IGF-1R were from Bio-Source International, Becton-Dickinson, Rockland, GenTex and internal production, respectively. The 4G10 antibody was from Millipore (#05-321MG). -
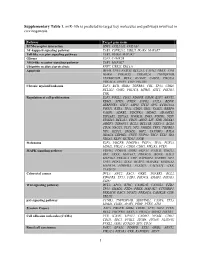
1 Supplementary Table 1. Mir-10B Is Predicted to Target Key Molecules
Supplementary Table 1. miR-10b is predicted to target key molecules and pathways involved in carcinogenesis. Pathway Target gene name ECM-receptor interaction SDC1, COL24A1, COL4A4 NF-kappa B signaling pathway TAB1, CSNK2A2, UBE2I, IRAK4, MAP3K7 Toll-like receptor signaling pathway TAB1, IRAK4, MAP3K7 Glioma E2F3, CAMK2B NOD-like receptor signaling pathway TAB1, MAP3K7 Ubiquitin mediated proteolysis RNF7, UBE2I, ERCC8 Apoptosis DFFB, TP53, FASLG, BCL2L1, CAPN2, PRKX, ATM, IRAK4, PRKACG, PRKAR2A, TNFRSF10B, TNFRSF10D, BCL2, IL1RAP, CASP8, PIK3CA, PRKACA, APAF1, CHP, PIK3R3 Chronic myeloid leukemia E2F3, BCR, GRB2, TGFBR1, CBL, TP53, CDK6, BCL2L1, GAB2, PIK3CA, MDM2, SHC1, PIK3R3, CRK Regulation of cell proliferation E2F3, FOSL2, CDX2, PDGFB, OSMR, E2F7, ARNT2, RBM5, STRN, PTEN, S1PR2, CUL3, BDNF, SERPINE1, SHC1, ASPH, ITCH, SPN, CCDC88A, FOXJ1, RXRA, TP53, CDK6, IRS1, VASH2, RBBP9, VASH1, ADRB2, PDGFRA, MDM2, ADAMTS1, EIF2AK2, EIF5A2, ICOSLG, ING5, FGFR3, NDN, ST8SIA1, BCL2L1, CDH5, ARNT, LIF, VDR, HOXA3, AGGF1, TSPAN31, BCL2, BCL11B, NKX3-1, BCL6, CD28, NACC1, FLT1, NF2, JARID2, TBX5, TGFBR1, NF1, KLF11, SMAD2, IGF2, TAX1BP3, BTLA, HDAC4, LEPRE1, CNTF, NUP62, TSC1, ETS1, ID4, NR5A2, KLF4, KCTD11, NFIB Melanoma E2F3, PDGFB, PDGFRA, FGF11, TP53, FGF23, MDM2, PIK3CA, CDK6, CDH1, PIK3R3, PTEN MAPK signaling pathway FGFR3, PDGFB, GRB2, FGF11, FASLG, GNG12, SRF, PRKX, MAP3K7, PRKACG, BDNF, RAC3, MAP3K2, PRKACA, CHP, RAPGEF2, TGFBR1, NF1, TP53, FGF23, STK4, DUSP5, MAP4K4, RPS6KA2, MAPK14, PDGFRA, PLA2G3, CACNA1C, CRK, PLA2G2F Colorectal cancer -

IER3 Is a Crucial Mediator of Tap73b-Induced Apoptosis In
OPEN IER3 is a crucial mediator of SUBJECT AREAS: TAp73b-induced apoptosis in cervical CELL DEATH TUMOUR SUPPRESSORS cancer and confers etoposide sensitivity Hanyong Jin1*, Dae-Shik Suh2*, Tae-Hyoung Kim3, Ji-Hyun Yeom4, Kangseok Lee4 & Jeehyeon Bae5 Received 14 September 2014 1Department of Pharmacy, CHA University, Seongnam, 463-836, Korea, 2Department of Obstetrics and Gynecology, Asan Accepted Medical Center, University of Ulsan College of Medicine, 3Department of Biochemistry, Chosun University School of Medicine, 4 5 9 January 2015 Gwangju 501-759, Korea, Department of Life Science, Chung-Ang University, Seoul, 156-756, Korea, School of Pharmacy, Chung-Ang University, Seoul, 156-756, Korea. Published 10 February 2015 Infection with high-risk human papillomaviruses (HPVs) causes cervical cancer. E6 oncoprotein, an HPV gene product, inactivates the major gatekeeper p53. In contrast, its isoform, TAp73b, has become increasingly important, as it is resistant to E6. However, the intracellular signaling mechanisms that account Correspondence and for TAp73b tumor suppressor activity in cervix are poorly understood. Here, we identified that IER3 is a requests for materials novel target gene of TAp73b. In particular, TAp73b exclusively transactivated IER3 in cervical cancer cells, should be addressed to whereas p53 and TAp63 failed to do. IER3 efficiently induced apoptosis, and its knockdown promoted K.L. (kangseok@cau. survival of HeLa cells. In addition, TAp73b-induced cell death, but not p53-induced cell death, was inhibited ac.kr) or J.B. upon IER3 silencing. Moreover, etoposide, a DNA-damaging chemotherapeutics, upregulated TAp73b and IER3 in a c-Abl tyrosine kinase-dependent manner, and the etoposide chemosensitivity of HeLa cells was ([email protected]) largely determined by TAp73b-induced IER3. -

S41467-020-18249-3.Pdf
ARTICLE https://doi.org/10.1038/s41467-020-18249-3 OPEN Pharmacologically reversible zonation-dependent endothelial cell transcriptomic changes with neurodegenerative disease associations in the aged brain Lei Zhao1,2,17, Zhongqi Li 1,2,17, Joaquim S. L. Vong2,3,17, Xinyi Chen1,2, Hei-Ming Lai1,2,4,5,6, Leo Y. C. Yan1,2, Junzhe Huang1,2, Samuel K. H. Sy1,2,7, Xiaoyu Tian 8, Yu Huang 8, Ho Yin Edwin Chan5,9, Hon-Cheong So6,8, ✉ ✉ Wai-Lung Ng 10, Yamei Tang11, Wei-Jye Lin12,13, Vincent C. T. Mok1,5,6,14,15 &HoKo 1,2,4,5,6,8,14,16 1234567890():,; The molecular signatures of cells in the brain have been revealed in unprecedented detail, yet the ageing-associated genome-wide expression changes that may contribute to neurovas- cular dysfunction in neurodegenerative diseases remain elusive. Here, we report zonation- dependent transcriptomic changes in aged mouse brain endothelial cells (ECs), which pro- minently implicate altered immune/cytokine signaling in ECs of all vascular segments, and functional changes impacting the blood–brain barrier (BBB) and glucose/energy metabolism especially in capillary ECs (capECs). An overrepresentation of Alzheimer disease (AD) GWAS genes is evident among the human orthologs of the differentially expressed genes of aged capECs, while comparative analysis revealed a subset of concordantly downregulated, functionally important genes in human AD brains. Treatment with exenatide, a glucagon-like peptide-1 receptor agonist, strongly reverses aged mouse brain EC transcriptomic changes and BBB leakage, with associated attenuation of microglial priming. We thus revealed tran- scriptomic alterations underlying brain EC ageing that are complex yet pharmacologically reversible. -

HIV-1 TAR Mirna Protects Against Apoptosis by Altering Cellular Gene
Retrovirology BioMed Central Research Open Access HIV-1 TAR miRNA protects against apoptosis by altering cellular gene expression Zachary Klase1, Rafael Winograd1, Jeremiah Davis1, Lawrence Carpio1, Richard Hildreth1, Mohammad Heydarian2, Sidney Fu2, Timothy McCaffrey2, Eti Meiri3, Mila Ayash-Rashkovsky3, Shlomit Gilad3, Zwi Bentwich3 and Fatah Kashanchi*1 Address: 1The Department of Microbiology, Immunology and Tropical Medicine program, The George Washington University School of Medicine, Washington, District of Columbia 20037, USA, 2The Department of Biochemistry and Molecular Biology, The George Washington University School of Medicine, Washington, District of Columbia 20037, USA and 3Rosetta Genomics Ltd., Rehovot, Israel Email: Zachary Klase - [email protected]; Rafael Winograd - [email protected]; Jeremiah Davis - [email protected]; Lawrence Carpio - [email protected]; Richard Hildreth - [email protected]; Mohammad Heydarian - [email protected]; Sidney Fu - [email protected]; Timothy McCaffrey - [email protected]; Eti Meiri - [email protected]; Mila Ayash- Rashkovsky - [email protected]; Shlomit Gilad - [email protected]; Zwi Bentwich - [email protected]; Fatah Kashanchi* - [email protected] * Corresponding author Published: 16 February 2009 Received: 15 August 2008 Accepted: 16 February 2009 Retrovirology 2009, 6:18 doi:10.1186/1742-4690-6-18 This article is available from: http://www.retrovirology.com/content/6/1/18 © 2009 Klase et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: RNA interference is a gene regulatory mechanism that employs small RNA molecules such as microRNA. -

Supplementary Materials
Supplementary materials Supplementary Table S1: MGNC compound library Ingredien Molecule Caco- Mol ID MW AlogP OB (%) BBB DL FASA- HL t Name Name 2 shengdi MOL012254 campesterol 400.8 7.63 37.58 1.34 0.98 0.7 0.21 20.2 shengdi MOL000519 coniferin 314.4 3.16 31.11 0.42 -0.2 0.3 0.27 74.6 beta- shengdi MOL000359 414.8 8.08 36.91 1.32 0.99 0.8 0.23 20.2 sitosterol pachymic shengdi MOL000289 528.9 6.54 33.63 0.1 -0.6 0.8 0 9.27 acid Poricoic acid shengdi MOL000291 484.7 5.64 30.52 -0.08 -0.9 0.8 0 8.67 B Chrysanthem shengdi MOL004492 585 8.24 38.72 0.51 -1 0.6 0.3 17.5 axanthin 20- shengdi MOL011455 Hexadecano 418.6 1.91 32.7 -0.24 -0.4 0.7 0.29 104 ylingenol huanglian MOL001454 berberine 336.4 3.45 36.86 1.24 0.57 0.8 0.19 6.57 huanglian MOL013352 Obacunone 454.6 2.68 43.29 0.01 -0.4 0.8 0.31 -13 huanglian MOL002894 berberrubine 322.4 3.2 35.74 1.07 0.17 0.7 0.24 6.46 huanglian MOL002897 epiberberine 336.4 3.45 43.09 1.17 0.4 0.8 0.19 6.1 huanglian MOL002903 (R)-Canadine 339.4 3.4 55.37 1.04 0.57 0.8 0.2 6.41 huanglian MOL002904 Berlambine 351.4 2.49 36.68 0.97 0.17 0.8 0.28 7.33 Corchorosid huanglian MOL002907 404.6 1.34 105 -0.91 -1.3 0.8 0.29 6.68 e A_qt Magnogrand huanglian MOL000622 266.4 1.18 63.71 0.02 -0.2 0.2 0.3 3.17 iolide huanglian MOL000762 Palmidin A 510.5 4.52 35.36 -0.38 -1.5 0.7 0.39 33.2 huanglian MOL000785 palmatine 352.4 3.65 64.6 1.33 0.37 0.7 0.13 2.25 huanglian MOL000098 quercetin 302.3 1.5 46.43 0.05 -0.8 0.3 0.38 14.4 huanglian MOL001458 coptisine 320.3 3.25 30.67 1.21 0.32 0.9 0.26 9.33 huanglian MOL002668 Worenine -

The P90 RSK Family Members: Common Functions and Isoform Specificity
Published OnlineFirst August 22, 2013; DOI: 10.1158/0008-5472.CAN-12-4448 Cancer Review Research The p90 RSK Family Members: Common Functions and Isoform Specificity Romain Lara, Michael J. Seckl, and Olivier E. Pardo Abstract The p90 ribosomal S6 kinases (RSK) are implicated in various cellular processes, including cell proliferation, survival, migration, and invasion. In cancer, RSKs modulate cell transformation, tumorigenesis, and metastasis. Indeed, changes in the expression of RSK isoforms have been reported in several malignancies, including breast, prostate, and lung cancers. Four RSK isoforms have been identified in humans on the basis of their high degree of sequence homology. Although this similarity suggests some functional redundancy between these proteins, an increasing body of evidence supports the existence of isoform-based specificity among RSKs in mediating particular cellular processes. This review briefly presents the similarities between RSK family members before focusing on the specific function of each of the isoforms and their involvement in cancer progression. Cancer Res; 73(17); 1–8. Ó2013 AACR. Introduction subsequently cloned throughout the Metazoan kingdom (2). The extracellular signal–regulated kinase (ERK)1/2 pathway The genomic analysis of several cancer types suggests that fi is involved in key cellular processes, including cell prolifera- these genes are not frequently ampli ed or mutated, with some tion, differentiation, survival, metabolism, and migration. notable exceptions (e.g., in the case of hepatocellular carcino- More than 30% of all human cancers harbor mutations within ma; ref. 6). Table 1 summarizes reported genetic changes in this pathway, mostly resulting in gain of function and conse- RSK genes. -

ERBB2 Influences the Subcellular Localization of the Estrogen
Endocrine-Related Cancer (2008) 15 985–1002 ERBB2 influences the subcellular localization of the estrogen receptor in tamoxifen-resistant MCF-7 cells leading to the activation of AKT and RPS6KA2 Sunil Pancholi, Anne E Lykkesfeldt1, Caroline Hilmi, Susana Banerjee, Alexandra Leary, Suzanne Drury, Stephen Johnston 2,MitchDowsett and Lesley-Ann Martin Institute of Cancer Research, The Breakthrough Breast Cancer Research Centre, 237 Fulham Road, London SW3 6JB, UK 1Institute of Cancer Biology, Danish Cancer Society, Copenhagen, Denmark 2Department of Medicine, Royal Marsden Hospital, Fulham Road, London, UK (Correspondence should be addressed to L-A Martin; Email: [email protected]) Abstract Acquired resistance to endocrine therapies remains a major clinical obstacle in hormone-sensitive breast tumors. We used an MCF-7 breast tumor cell line (TamR-1) resistant to tamoxifen to investigate this mechanism. We demonstrate that TamR-1 express elevated levels of phosphorylated AKT and MAPK3/1-activated RPS6KA2 compared with the parental MCF-7 cell line (MCF-7). There was no change in the level of total ESR between the two cell lines; however, the TamR-1 cells had increased phosphorylation of ESR1 ser167. SiRNA blockade of AKT or MAPK3/1 had little effect on ESR1 ser167 phosphorylation, but a combination of the two siRNAs abrogated this. Co-localization studies revealed an association between ERBB2 and ESR1 in the TamR-1 but not MCF-7 cells. ESR1 was redistributed to extranuclear sites in TamR-1 and was less transcriptionally competent compared with MCF-7 suggesting that nuclear ESR1 activity was suppressed in TamR-1. Tamoxifen resistance in the TamR-1 cells could be partially overcome by the ERBB2 inhibitor AG825 in combination with tamoxifen, and this was associated with re-localization of ESR1 to the nucleus. -

WO 2017/015637 Al 26 January 2017 (26.01.2017) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2017/015637 Al 26 January 2017 (26.01.2017) P O P C T (51) International Patent Classification: (81) Designated States (unless otherwise indicated, for every A61K 48/00 (2006.01) C12N 15/63 (2006.01) kind of national protection available): AE, AG, AL, AM, C07K 19/00 (2006.01) C12N 15/65 (2006.01) AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, C12N 5/22 (2006.0 1) C12N 15/867 (2006.0 1) BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, (21) International Application Number: HN, HR, HU, ID, IL, IN, IR, IS, JP, KE, KG, KN, KP, KR, PCT/US2016/043756 KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, (22) International Filing Date: MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, 22 July 2016 (22.07.2016) PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, (25) Filing Language: English TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (26) Publication Language: English (84) Designated States (unless otherwise indicated, for every (30) Priority Data: kind of regional protection available): ARIPO (BW, GH, 62/195,680 22 July 2015 (22.07.2015) US GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, 62/293,3 13 9 February 2016 (09.02.2016) US TZ, UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, TJ, TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, (71) Applicant: DUKE UNIVERSITY [US/US]; 2812 Erwin DK, EE, ES, FI, FR, GB, GR, HR, HU, IE, IS, IT, LT, LU, Road, Suite 306, Durham, NC 27705 (US). -

Transcriptome Sequencing of Peripheral Blood Mononuclear
www.nature.com/scientificreports OPEN Transcriptome Sequencing of Peripheral Blood Mononuclear Cells from Elite Controller-Long Term Received: 8 February 2019 Accepted: 12 September 2019 Non Progressors Published: xx xx xxxx Francisco Díez-Fuertes1,2, Humberto Erick De La Torre-Tarazona1, Esther Calonge1, Maria Pernas3, María del Mar Alonso-Socas4, Laura Capa1, Javier García-Pérez1, Anavaj Sakuntabhai 5 & José Alcamí 1,2 The elite controller (EC)-long term non-progressor (LTNP) phenotype represent a spontaneous and advantageous model of HIV-1 control in the absence of therapy. The transcriptome of peripheral blood mononuclear cells (PBMCs) collected from EC-LTNPs was sequenced by RNA-Seq and compared with the transcriptomes from other phenotypes of disease progression. The transcript abundance estimation combined with the use of supervised classifcation algorithms allowed the selection of 20 genes and pseudogenes, mainly involved in interferon-regulated antiviral mechanisms and cell machineries of transcription and translation, as the best predictive genes of disease progression. Diferential expression analyses between phenotypes showed an altered calcium homeostasis in EC-LTNPs evidenced by the upregulation of several membrane receptors implicated in calcium-signaling cascades and intracellular calcium-mobilization and by the overrepresentation of NFAT1/Elk-1-binding sites in the promoters of the genes diferentially expressed in these individuals. A coordinated upregulation of host genes associated with HIV-1 reverse transcription and viral transcription was also observed in EC-LTNPs –i.e. p21/CDKN1A, TNF, IER3 and GADD45B. We also found an upregulation of ANKRD54 in EC-LTNPs and viremic LTNPs in comparison with typical progressors and a clear alteration of type-I interferon signaling as a consequence of viremia in typical progressors before and after receiving antiretroviral therapy. -

Microarray Gene Expression Analysis in Ovine Ductus Arteriosus During Fetal Development and Birth Transition
nature publishing group Articles Basic Science Investigation Microarray gene expression analysis in ovine ductus arteriosus during fetal development and birth transition Ravi Goyal1, Dipali Goyal1, Lawrence D. Longo1 and Ronald I. Clyman2 BACKGROUND: Patent ductus arteriosus (PDA) in the new- (PDA) persists after birth, the blood shunts from the high- born is the most common congenital heart anomaly and is pressure aorta to the low-pressure pulmonary artery (left to significantly more common in preterm infants. Contemporary right shunt). This can lead to pulmonary hyperemia and edema pharmacological treatment is effective in only 70–80% of the and decreases renal, mesenteric, and cerebral perfusion result- cases. Moreover, indomethacin or ibuprofen, which are used ing in pulmonary engorgement with an increase in pulmonary to close a PDA may be accompanied by serious side effects in vascular resistance and congestive cardiac failure. In those premature infants. To explore the novel molecular pathways, cases in which the pulmonary vascular resistance exceeds which may be involved in the maturation and closure of the the systemic vascular resistance, the blood shunts from right ductus arteriosus (DA), we used fetal and neonatal sheep to test to left. The main therapeutic option for a PDA is to treat the the hypothesis that maturational development of DA is associ- neonate with indomethacin or ibuprofen to inhibit the enzyme ated with significant alterations in specific mRNA expression. cyclooxygenase and thus inhibit prostaglandin synthesis. This METHODS: We conducted oligonucleotide microarray exper- is successful in about 70–80% of infants. Its use, however, may iments on the isolated mRNA from DA and ascending aorta lead to undesirable side effects such as gastrointestinal bleed- from three study groups (premature fetus—97 ± 0 d, near-term ing, perforation, and/or necrotizing enterocolitis. -

Supplementary Figure 1. Network Map Associated with Upregulated Canonical Pathways Shows Interferon Alpha As a Key Regulator
Supplementary Figure 1. Network map associated with upregulated canonical pathways shows interferon alpha as a key regulator. IPA core analysis determined interferon-alpha as an upstream regulator in the significantly upregulated genes from RNAseq data from nasopharyngeal swabs of COVID-19 patients (GSE152075). Network map was generated in IPA, overlaid with the Coronavirus Replication Pathway. Supplementary Figure 2. Network map associated with Cell Cycle, Cellular Assembly and Organization, DNA Replication, Recombination, and Repair shows relationships among significant canonical pathways. Significant pathways were identified from pathway analysis of RNAseq from PBMCs of COVID-19 patients. Coronavirus Pathogenesis Pathway was also overlaid on the network map. The orange and blue colors in indicate predicted activation or predicted inhibition, respectively. Supplementary Figure 3. Significant biological processes affected in brochoalveolar lung fluid of severe COVID-19 patients. Network map was generated by IPA core analysis of differentially expressed genes for severe vs mild COVID-19 patients in bronchoalveolar lung fluid (BALF) from scRNA-seq profile of GSE145926. Orange color represents predicted activation. Red boxes highlight important cytokines involved. Supplementary Figure 4. 10X Genomics Human Immunology Panel filtered differentially expressed genes in each immune subset (NK cells, T cells, B cells, and Macrophages) of severe versus mild COVID-19 patients. Three genes (HLA-DQA2, IFIT1, and MX1) were found significantly and consistently differentially expressed. Gene expression is shown per the disease severity (mild, severe, recovered) is shown on the top row and expression across immune cell subsets are shown on the bottom row. Supplementary Figure 5. Network map shows interactions between differentially expressed genes in severe versus mild COVID-19 patients.