Whole Egg Consumption Increases Gene Expression Within the Glutathione Pathway in the Liver of Zucker Diabetic Fatty Rats
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

Acetyl Group Coordinated Progression Through the Catalytic Cycle of an Arylalkylamine N-Acetyltransferase
RESEARCH ARTICLE Acetyl group coordinated progression through the catalytic cycle of an arylalkylamine N-acetyltransferase Adam A. Aboalroub, Ashleigh B. Bachman, Ziming Zhang, Dimitra Keramisanou, David J. Merkler, Ioannis Gelis* Department of Chemistry, University of South Florida, Tampa, Florida, United States of America * [email protected] a1111111111 a1111111111 a1111111111 Abstract a1111111111 a1111111111 The transfer of an acetyl group from acetyl-CoA to an acceptor amine is a ubiquitous bio- chemical transformation catalyzed by Gcn5-related N-acetyltransferases (GNATs). Although it is established that the reaction proceeds through a sequential ordered mecha- nism, the role of the acetyl group in driving the ordered formation of binary and ternary com- OPEN ACCESS plexes remains elusive. Herein, we show that CoA and acetyl-CoA alter the conformation of the substrate binding site of an arylalkylamine N-acetyltransferase (AANAT) to facilitate Citation: Aboalroub AA, Bachman AB, Zhang Z, Keramisanou D, Merkler DJ, Gelis I (2017) Acetyl interaction with acceptor substrates. However, it is the presence of the acetyl group within group coordinated progression through the the catalytic funnel that triggers high affinity binding. Acetyl group occupancy is relayed catalytic cycle of an arylalkylamine N- through a conserved salt bridge between the P-loop and the acceptor binding site, and is acetyltransferase. PLoS ONE 12(5): e0177270. manifested as differential dynamics in the CoA and acetyl-CoA-bound states. The capacity https://doi.org/10.1371/journal.pone.0177270 of the acetyl group carried by an acceptor to promote its tight binding even in the absence of Editor: Viswanathan V. Krishnan, California State CoA, but also its mutually exclusive position to the acetyl group of acetyl-CoA underscore its University Fresno, UNITED STATES importance in coordinating the progression of the catalytic cycle. -

PDF Document Created by Pdffiller
Patient: 1234567843314948-COtGx0053 CLIA ID#: 11D2066426 Larry Hung, MD, Laboratory Director GxTM Carrier Screen Testing Report Patient Information Provider Information Specimen Patient Name Haley Papevies Provider Harbin Clinic Women's Accession ID 1234567843314948 Center Cartersville Date of Birth Apr 16, 1998 Sample ID COtGx0053XX Provider ID 1124488556 Age 19 Specimen Type Saliva Physician Vicki Yates Sex female Collection Date Jul 20, 2017 Ethnicity Report Date Aug 5, 2017 Test Ordered CF Patient Results: Negative - No Pathogenic or Likely-Pathogenic Variant(s) Detected Additional Comments This report is based on the analysis of CFTR gene included in the Carrier Screen. No known pathogenic or likely pathogenic variant(s) detected in the coding sequences of CFTR gene. Followup Recommendations Follow up with physicians for updated carrier screen information. The sequencing for CFTR gene was carried out with the other genes included in the Carrier Screen Testing (listed below). The analysis of the other genes in the Carrier Screen could be ordered through your physicians. Genes Tested Targeted regions for “Carrier Screen Testing” includes the exonic regions of the following genes: ABCC8, ABCD1, ABCD4, ACAD8, ACADM, ACADS, ACADSB, ACADVL, ACAT1, ACSF3, ACTA2, ACTC1, ADA, ADAMTS2, AGXT, AHCY, APC, APOB, ARG1, ASL, ASPA, ASS1, ATP7B, AUH, BCKDHA, BBS2, BCKDHB, BLM, BTD, CBS, COL3A1, COL4A3, CD320, CFTR, CLRN1, CPT1A, CPT2, CYP1B1, CYP21A2, DBT, DHCR7, DHDDS, DLD, DMD, DNAJC19, DSC2, DSG2, DSP, DUOX2, ETFA, ETFB, ETFDH, FAH, FANCC, FBN1, -

Supplement 1 Overview of Dystonia Genes
Supplement 1 Overview of genes that may cause dystonia in children and adolescents Gene (OMIM) Disease name/phenotype Mode of inheritance 1: (Formerly called) Primary dystonias (DYTs): TOR1A (605204) DYT1: Early-onset generalized AD primary torsion dystonia (PTD) TUBB4A (602662) DYT4: Whispering dystonia AD GCH1 (600225) DYT5: GTP-cyclohydrolase 1 AD deficiency THAP1 (609520) DYT6: Adolescent onset torsion AD dystonia, mixed type PNKD/MR1 (609023) DYT8: Paroxysmal non- AD kinesigenic dyskinesia SLC2A1 (138140) DYT9/18: Paroxysmal choreoathetosis with episodic AD ataxia and spasticity/GLUT1 deficiency syndrome-1 PRRT2 (614386) DYT10: Paroxysmal kinesigenic AD dyskinesia SGCE (604149) DYT11: Myoclonus-dystonia AD ATP1A3 (182350) DYT12: Rapid-onset dystonia AD parkinsonism PRKRA (603424) DYT16: Young-onset dystonia AR parkinsonism ANO3 (610110) DYT24: Primary focal dystonia AD GNAL (139312) DYT25: Primary torsion dystonia AD 2: Inborn errors of metabolism: GCDH (608801) Glutaric aciduria type 1 AR PCCA (232000) Propionic aciduria AR PCCB (232050) Propionic aciduria AR MUT (609058) Methylmalonic aciduria AR MMAA (607481) Cobalamin A deficiency AR MMAB (607568) Cobalamin B deficiency AR MMACHC (609831) Cobalamin C deficiency AR C2orf25 (611935) Cobalamin D deficiency AR MTRR (602568) Cobalamin E deficiency AR LMBRD1 (612625) Cobalamin F deficiency AR MTR (156570) Cobalamin G deficiency AR CBS (613381) Homocysteinuria AR PCBD (126090) Hyperphelaninemia variant D AR TH (191290) Tyrosine hydroxylase deficiency AR SPR (182125) Sepiaterine reductase -

Supplemental Table S1
Entrez Gene Symbol Gene Name Affymetrix EST Glomchip SAGE Stanford Literature HPA confirmed Gene ID Profiling profiling Profiling Profiling array profiling confirmed 1 2 A2M alpha-2-macroglobulin 0 0 0 1 0 2 10347 ABCA7 ATP-binding cassette, sub-family A (ABC1), member 7 1 0 0 0 0 3 10350 ABCA9 ATP-binding cassette, sub-family A (ABC1), member 9 1 0 0 0 0 4 10057 ABCC5 ATP-binding cassette, sub-family C (CFTR/MRP), member 5 1 0 0 0 0 5 10060 ABCC9 ATP-binding cassette, sub-family C (CFTR/MRP), member 9 1 0 0 0 0 6 79575 ABHD8 abhydrolase domain containing 8 1 0 0 0 0 7 51225 ABI3 ABI gene family, member 3 1 0 1 0 0 8 29 ABR active BCR-related gene 1 0 0 0 0 9 25841 ABTB2 ankyrin repeat and BTB (POZ) domain containing 2 1 0 1 0 0 10 30 ACAA1 acetyl-Coenzyme A acyltransferase 1 (peroxisomal 3-oxoacyl-Coenzyme A thiol 0 1 0 0 0 11 43 ACHE acetylcholinesterase (Yt blood group) 1 0 0 0 0 12 58 ACTA1 actin, alpha 1, skeletal muscle 0 1 0 0 0 13 60 ACTB actin, beta 01000 1 14 71 ACTG1 actin, gamma 1 0 1 0 0 0 15 81 ACTN4 actinin, alpha 4 0 0 1 1 1 10700177 16 10096 ACTR3 ARP3 actin-related protein 3 homolog (yeast) 0 1 0 0 0 17 94 ACVRL1 activin A receptor type II-like 1 1 0 1 0 0 18 8038 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 1 0 0 0 0 19 8751 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 1 0 0 0 0 20 8728 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 1 0 0 0 0 21 81792 ADAMTS12 ADAM metallopeptidase with thrombospondin type 1 motif, 12 1 0 0 0 0 22 9507 ADAMTS4 ADAM metallopeptidase with thrombospondin type 1 -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -

4-6 Weeks Old Female C57BL/6 Mice Obtained from Jackson Labs Were Used for Cell Isolation
Methods Mice: 4-6 weeks old female C57BL/6 mice obtained from Jackson labs were used for cell isolation. Female Foxp3-IRES-GFP reporter mice (1), backcrossed to B6/C57 background for 10 generations, were used for the isolation of naïve CD4 and naïve CD8 cells for the RNAseq experiments. The mice were housed in pathogen-free animal facility in the La Jolla Institute for Allergy and Immunology and were used according to protocols approved by the Institutional Animal Care and use Committee. Preparation of cells: Subsets of thymocytes were isolated by cell sorting as previously described (2), after cell surface staining using CD4 (GK1.5), CD8 (53-6.7), CD3ε (145- 2C11), CD24 (M1/69) (all from Biolegend). DP cells: CD4+CD8 int/hi; CD4 SP cells: CD4CD3 hi, CD24 int/lo; CD8 SP cells: CD8 int/hi CD4 CD3 hi, CD24 int/lo (Fig S2). Peripheral subsets were isolated after pooling spleen and lymph nodes. T cells were enriched by negative isolation using Dynabeads (Dynabeads untouched mouse T cells, 11413D, Invitrogen). After surface staining for CD4 (GK1.5), CD8 (53-6.7), CD62L (MEL-14), CD25 (PC61) and CD44 (IM7), naïve CD4+CD62L hiCD25-CD44lo and naïve CD8+CD62L hiCD25-CD44lo were obtained by sorting (BD FACS Aria). Additionally, for the RNAseq experiments, CD4 and CD8 naïve cells were isolated by sorting T cells from the Foxp3- IRES-GFP mice: CD4+CD62LhiCD25–CD44lo GFP(FOXP3)– and CD8+CD62LhiCD25– CD44lo GFP(FOXP3)– (antibodies were from Biolegend). In some cases, naïve CD4 cells were cultured in vitro under Th1 or Th2 polarizing conditions (3, 4). -

Oral Presentations
Journal of Inherited Metabolic Disease (2018) 41 (Suppl 1):S37–S219 https://doi.org/10.1007/s10545-018-0233-9 ABSTRACTS Oral Presentations PARALLEL SESSION 1A: Clycosylation and cardohydrate disorders O-002 Link between glycemia and hyperlipidemia in Glycogen Storage O-001 Disease type Ia Hoogerland J A1, Hijmans B S1, Peeks F1, Kooijman S3, 4, Bos T2, Fertility in classical galactosaemia, N-glycan, hormonal and inflam- Bleeker A1, Van Dijk T H2, Wolters H1, Havinga R1,PronkACM3, 4, matory gene expression interactions Rensen P C N3, 4,MithieuxG5, 6, Rajas F5, 6, Kuipers F1, 2,DerksTGJ1, Reijngoud D1,OosterveerMH1 Colhoun H O1,Rubio-GozalboME2,BoschAM3, Knerr I4,DawsonC5, Brady J J6,GalliganM8,StepienKM9, O'Flaherty R O7,MossC10, 1Dep Pediatrics, CLDM, Univ of Groningen, Groningen, Barker P11, Fitzgibbon M C6, Doran P8,TreacyEP1, 4, 9 Netherlands, 2Lab Med, CLDM, Univ of Groningen, Groningen, Netherlands, 3Dep of Med, Div of Endocrinology, LUMC, Leiden, 1Dept Paediatrics, Trinity College Dublin, Dublin, Ireland, 2Dept Paeds and Netherlands, 4Einthoven Lab Exp Vasc Med, LUMC, Leiden, Clin Genetics, UMC, Maastricht, Netherlands, 3Dept Paediatrics, AMC, Netherlands, 5Institut Nat Sante et Recherche Med, Lyon, Amsterdam, Netherlands, 4NCIMD, TSCUH, Dublin, Ireland, 5Dept France, 6Univ Lyon 1, Villeurbanne, France Endocrinology, NHS Foundation Trust, Birmingham, United Kingdom, 6Dept Clin Biochem, MMUH, Dublin, Ireland, 7NIBRT Glycoscience, Background: Glycogen Storage Disease type Ia (GSD Ia) is an NIBRT, Dublin, Ireland, 8UCDCRC,UCD,Dublin,Ireland,9NCIMD, inborn error of glucose metabolism characterized by fasting hypo- MMUH, Dublin, Ireland, 10Conway Institute, UCD, Dublin, Ireland, glycemia, hyperlipidemia and fatty liver disease. We have previ- 11CBAL, NHS Foundation, Cambridge, United Kingdom ously reported considerable heterogeneity in circulating triglycer- ide levels between individual GSD Ia patients, a phenomenon that Background: Classical Galactosaemia (CG) is caused by deficiency of is poorly understood. -

Balance Between Senescence and Apoptosis Is Regulated by Telomere Damage–Induced Association Between P16 and Caspase-3
JBC Papers in Press. Published on May 10, 2018 as Manuscript RA118.003506 The latest version is at http://www.jbc.org/cgi/doi/10.1074/jbc.RA118.003506 Balance between senescence and apoptosis is regulated by telomere damage–induced association between p16 and caspase-3 Shanmugam Panneer Selvam1,2, Braden M. Roth1,2, Rose Nganga1,2, Jisun Kim1,2, Marion A. Cooley3,#, Kristi Helke4, Charles D. Smith5, and Besim Ogretmen1,2* 1Department of Biochemistry and Molecular Biology, 2Hollings Cancer Center, 3Department of Regenerative Medicine, 4Department of Comparative Medicine, Medical University of South Carolina, 86 Jonathan Lucas Street, Charleston, SC 29425; 5Department of Pharmacology, Pennsylvania State University, 500 University Drive, Hershey, PA 17033. #Current Address: Department of Oral Biology, Augusta University, Augusta, GA 30912. *Address correspondence to: [email protected] Running title: p16-caspase-3 complex and senescence Key words: apoptosis; senescence; sphingolipid; sphingosine 1-phosphate; telomere damage Downloaded from Abstract damage–induced apoptosis, indicating that an Telomerase activation protects cells from telomere association between p16 and caspase-3 damage by delaying senescence and inducing cell proteinsforces senescence induction by inhibiting immortalization, whereas telomerase inhibition caspase- 3 activation and apoptosis. These results http://www.jbc.org/ mediates rapid senescence or apoptosis. However, suggest that p16 plays a direct role in telomere the cellular mechanisms that determine telomere damage–dependent senescence by limiting damage–dependent senescence versus apoptosis apoptosis via binding to caspase-3, revealing a induction are largely unknown. Here, we direct link between telomere damage–dependent demonstrate that telomerase instability mediated senescence and apoptosis with regards to aging by guest on May 31, 2018 by silencing of sphingosine kinase 2 (SPHK2) and and cancer. -

SLC45A3-ELK4 Is a Novel and Frequent Erythroblast Transformation–Specific Fusion Transcript in Prostate Cancer
Published OnlineFirst March 17, 2009; DOI: 10.1158/0008-5472.CAN-08-4926 Priority Report SLC45A3-ELK4 Is a Novel and Frequent Erythroblast Transformation–Specific Fusion Transcript in Prostate Cancer David S. Rickman,1 Dorothee Pflueger,1 Benjamin Moss,1 Vanessa E. VanDoren,1 Chen X. Chen,1 Alexandre de la Taille,4,5 Rainer Kuefer,6 Ashutosh K. Tewari,2 Sunita R. Setlur,7 Francesca Demichelis,1,3 and Mark A. Rubin1 Departments of 1Pathology and Laboratory Medicine and 2Urology, and 3Institute for Computational Biomedicine, Weill Cornell Medical College, New York, New York; 4Department of Urology, CHU Mondor and 5Institut National de la Sante et de la Recherche Medicale, Unite´841, Cre´teil,France; 6Department of Urology, University Hospital Ulm, Ulm, Germany; and 7Department of Pathology, Brigham and Women’s Hospital and Harvard Medical School, Boston, Massachusetts Abstract (2, 3), and a specific expression profile (4, 5). Androgen-regulated ¶ Chromosomal rearrangements account for all erythroblast genes account for the majority of the 5 genomic regulatory transformation–specific (ETS) family member gene fusions promoters elements fused with ETS genes in prostate cancer (6). The promoter of the androgen-regulated transmembrane protease, that have been reported in prostate cancer and have clinical, serine 2 TMPRSS2 diagnostic, and prognostic implications. Androgen-regulated ( ) gene is fused to the coding region of members genes account for the majority of the 5¶ genomic regulatory of the ETS family of transcription factors, most commonly v-ets TMPRSS2-ERG, erythroblastosis virus E26 oncogene homologue (avian; ERG; ref. 7). promoter elements fused with ETS genes. Solute carrier family 45, member 3 SLC45A3 TMPRSS2-ETV1, and SLC45A3-ERG rearrangements account ( ), also referred to as for roughly 90% of ETS fusion prostate cancer. -
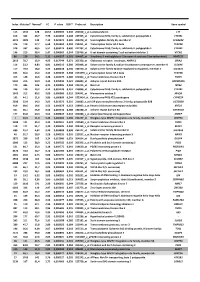
Index Mutated* Normal* FC P Value FDR** Probe Set Description Gene Symbol
Index Mutated* Normal* FC P value FDR** Probe set Description Gene symbol 113 1342 128 10,52 0,000503 0,240 202018_s_at Lactotransferrin LTF 616 362 46,7 7,76 0,003493 0,308 237395_at Cytochrome P450, family 4, subfamily Z, polypeptide 1 CYP4Z1 3073 1009 142 7,10 0,021674 0,385 206378_at Secretoglobin, family 2A, member 2 SCGB2A2 376 119 17,7 6,68 0,001962 0,284 214451_at Transcription factor AP-2 beta TFAP2B 578 337 60,5 5,57 0,003174 0,300 227702_at Cytochrome P450, family 4, subfamily X, polypeptide 1 CYP4X1 146 210 38,4 5,47 0,000669 0,244 219768_at V-set domain containing T cell activation inhibitor 1 VTCN1 86 129 24,2 5,31 0,000327 0,201 204607_at 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 2 (mitochondrial) HMGCS2 2613 78,7 15,9 4,95 0,017744 0,371 205358_at Glutamate receptor, ionotropic, AMPA 2 GRIA2 116 23,2 4,83 4,81 0,000513 0,240 203908_at Solute carrier family 4, sodium bicarbonate cotransporter, member 4 SLC4A4 117 79,4 18,0 4,42 0,000518 0,240 239723_at Solute carrier family 40 (iron-regulated transporter), member 1 SLC40A1 625 56,6 13,0 4,34 0,003549 0,308 1553394_a_atTranscription factor AP-2 beta TFAP2B 413 148 34,6 4,28 0,002175 0,286 240304_s_at Transmembrane channel-like 5 TMC5 5181 216 50,9 4,25 0,039946 0,421 223864_at Ankyrin repeat domain 30A ANKRD30A 179 446 106 4,21 0,000826 0,244 223315_at Netrin 4 NTN4 246 120 29,2 4,12 0,001128 0,251 210096_at Cytochrome P450, family 4, subfamily B, polypeptide 1 CYP4B1 1043 312 80,2 3,89 0,006080 0,319 204041_at Monoamine oxidase B MAOB 192 44,1 11,6 3,80 0,000899 0,244