Microtubule Organization and Microtubule- Associated Proteins (Maps)
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

1 Evidence for Gliadin Antibodies As Causative Agents in Schizophrenia
1 Evidence for gliadin antibodies as causative agents in schizophrenia. C.J.Carter PolygenicPathways, 20 Upper Maze Hill, Saint-Leonard’s on Sea, East Sussex, TN37 0LG [email protected] Tel: 0044 (0)1424 422201 I have no fax Abstract Antibodies to gliadin, a component of gluten, have frequently been reported in schizophrenia patients, and in some cases remission has been noted following the instigation of a gluten free diet. Gliadin is a highly immunogenic protein, and B cell epitopes along its entire immunogenic length are homologous to the products of numerous proteins relevant to schizophrenia (p = 0.012 to 3e-25). These include members of the DISC1 interactome, of glutamate, dopamine and neuregulin signalling networks, and of pathways involved in plasticity, dendritic growth or myelination. Antibodies to gliadin are likely to cross react with these key proteins, as has already been observed with synapsin 1 and calreticulin. Gliadin may thus be a causative agent in schizophrenia, under certain genetic and immunological conditions, producing its effects via antibody mediated knockdown of multiple proteins relevant to the disease process. Because of such homology, an autoimmune response may be sustained by the human antigens that resemble gliadin itself, a scenario supported by many reports of immune activation both in the brain and in lymphocytes in schizophrenia. Gluten free diets and removal of such antibodies may be of therapeutic benefit in certain cases of schizophrenia. 2 Introduction A number of studies from China, Norway, and the USA have reported the presence of gliadin antibodies in schizophrenia 1-5. Gliadin is a component of gluten, intolerance to which is implicated in coeliac disease 6. -

Microtubule Nucleation Remote from Centrosomes May Explain
Microtubule nucleation remote from centrosomes may INAUGURAL ARTICLE explain how asters span large cells Keisuke Ishiharaa,b,1, Phuong A. Nguyena,b, Aaron C. Groena,b, Christine M. Fielda,b, and Timothy J. Mitchisona,b,1 aDepartment of Systems Biology, Harvard Medical School, Boston, MA 02115; and bMarine Biological Laboratory, Woods Hole, MA 02543 This contribution is part of the special series of Inaugural Articles by members of the National Academy of Sciences elected in 2014. Edited by Ronald D. Vale, Howard Hughes Medical Institute and University of California, San Francisco, CA, and approved November 13, 2014 (received for review October 6, 2014) A major challenge in cell biology is to understand how nanometer- aster growth in large cells, such as microtubule sliding, tread- sized molecules can organize micrometer-sized cells in space and milling, or nucleation remote from centrosomes. time. One solution in many animal cells is a radial array of Previously we developed a cell-free system to reconstitute microtubules called an aster, which is nucleated by a central cleavage furrow signaling where growing asters interacted (5, organizing center and spans the entire cytoplasm. Frog (here 12). Here, we combine cell-free reconstitution and quantitative Xenopus laevis) embryos are more than 1 mm in diameter and imaging to identify microtubule nucleation away from the cen- divide with a defined geometry every 30 min. Like smaller cells, trosome as the key biophysical mechanism underlying aster they are organized by asters, which grow, interact, and move to growth. We propose that aster growth in large cells should be precisely position the cleavage planes. -
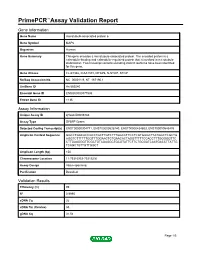
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name microtubule-associated protein 6 Gene Symbol MAP6 Organism Human Gene Summary This gene encodes a microtubule-associated protein. The encoded protein is a calmodulin-binding and calmodulin-regulated protein that is involved in microtubule stabilization. Two transcript variants encoding distinct isoforms have been identified for this gene. Gene Aliases FLJ41346, KIAA1878, MTAP6, N-STOP, STOP RefSeq Accession No. NC_000011.9, NT_167190.1 UniGene ID Hs.585540 Ensembl Gene ID ENSG00000171533 Entrez Gene ID 4135 Assay Information Unique Assay ID qHsaCID0006783 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000304771, ENST00000526740, ENST00000434603, ENST00000545476 Amplicon Context Sequence GGCCTGACACCGCCTGCTTGTCTTTGGCCTTCCTCGTGGGCTTATGGCTCGCTG AGGTCTTTTTTGGTTTGGAACTCTGAACACTAGGTTTTTCCACCTTTGGGGGTTC CTTGAAGGGTTCGCTGTAGAGGCTGCGTATTCTTCTGCGATCAATGACCTTATTG TCAGCTGTTGTTGGCT Amplicon Length (bp) 150 Chromosome Location 11:75316953-75319236 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 99 R2 0.9995 cDNA Cq 26 cDNA Tm (Celsius) 85 gDNA Cq 31.54 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report MAP6, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification Standard Curve Standard curve generated using 20 million copies of template diluted 10-fold to 20 copies Page 3/5 PrimePCR™Assay Validation Report Products used to generate validation data Real-Time PCR Instrument CFX384 Real-Time PCR Detection System Reverse Transcription Reagent iScript™ Advanced cDNA Synthesis Kit for RT-qPCR Real-Time PCR Supermix SsoAdvanced™ SYBR® Green Supermix Experimental Sample qPCR Human Reference Total RNA Data Interpretation Unique Assay ID This is a unique identifier that can be used to identify the assay in the literature and online. -

Role of Nucleation in Cortical Microtubule Array Organization: Variations on a Theme Erica A
Washington University in St. Louis Washington University Open Scholarship Biology Faculty Publications & Presentations Biology 7-2013 Role of nucleation in cortical microtubule array organization: variations on a theme Erica A. Fishel Washington University in St Louis Ram Dixit Washington University in St Louis, [email protected] Follow this and additional works at: https://openscholarship.wustl.edu/bio_facpubs Part of the Biochemistry Commons, Biology Commons, and the Plant Biology Commons Recommended Citation Fishel, Erica A. and Dixit, Ram, "Role of nucleation in cortical microtubule array organization: variations on a theme" (2013). Biology Faculty Publications & Presentations. 37. https://openscholarship.wustl.edu/bio_facpubs/37 This Article is brought to you for free and open access by the Biology at Washington University Open Scholarship. It has been accepted for inclusion in Biology Faculty Publications & Presentations by an authorized administrator of Washington University Open Scholarship. For more information, please contact [email protected]. 1 Title: Role of nucleation in cortical microtubule array organization: variations on a theme Authors: Erica A. Fishel and Ram Dixit Running title: Microtubule nucleation in the CMT array Key words: Noncentrosomal microtubules, γ-tubulin, Arabidopsis thaliana, Branching microtubules, Computer simulations, Interphase Corresponding Author: Ram Dixit Biology Department Washington University in St. Louis One Brookings Drive, CB 1137 St. Louis, MO 63130. Phone: (314) 935-8823 Fax: (314) 935-4432 Email: [email protected] 2 Abstract The interphase cortical microtubules (CMTs) of plant cells form strikingly ordered arrays in the absence of a dedicated microtubule-organizing center. Considerable effort has focused on activities such as bundling and severing that occur after CMT nucleation and are thought to be important for generating and maintaining ordered arrays. -

Microtubule Nucleation by Γ-Tubulin Complexes
REVIEWS Microtubule nucleation by γ‑tubulin complexes Justin M. Kollman*, Andreas Merdes‡, Lionel Mourey§ and David A. Agard* Abstract | Microtubule nucleation is regulated by the γ‑tubulin ring complex (γTuRC) and related γ‑tubulin complexes, providing spatial and temporal control over the initiation of microtubule growth. Recent structural work has shed light on the mechanism of γTuRC-based microtubule nucleation, confirming the long-standing hypothesis that the γTuRC functions as a microtubule template. The first crystallographic analysis of a non-γ‑tubulin γTuRC component (γ‑tubulin complex protein 4 (GCP4)) has resulted in a new appreciation of the relationships among all γTuRC proteins, leading to a refined model of their organization and function. The structures have also suggested an unexpected mechanism for regulating γTuRC activity via conformational modulation of the complex component GCP3. New experiments on γTuRC localization extend these insights, suggesting a direct link between its attachment at specific cellular sites and its activation. 4 Microtubule catastrophe The microtubule cytoskeleton is critically important in vitro from purified tubulin . In vivo, though, almost The rapid depolymerization of for the spatial and temporal organization of eukaryotic all microtubules have 13 protofilaments5–7, suggesting microtubules that occurs when cells, playing a central part in functions as diverse as that one level of cellular control involves defining unique GTP has been hydrolysed in all intracellular transport, organelle positioning, motil‑ microtubule geometry. The 13‑fold symmetry is probably tubulin subunits up to the growing tip. ity, signalling and cell division. The ability to play this preferred because it is the only geometry in which proto‑ range of parts requires microtubules to be arranged in filaments run straight along the microtubule length, as complex arrays that are capable of rapid reorganization. -

Specialized Cilia in Mammalian Sensory Systems
Cells 2015, 4, 500-519; doi:10.3390/cells4030500 OPEN ACCESS cells ISSN 2073-4409 www.mdpi.com/journal/cells Review Specialized Cilia in Mammalian Sensory Systems Nathalie Falk, Marlene Lösl, Nadja Schröder and Andreas Gießl * Department of Biology, Animal Physiology, University of Erlangen-Nuremberg, 91058 Erlangen, Germany; E-Mails: [email protected] (N.F.); [email protected] (M.L.); [email protected] (A.G.) * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +49-9131-85-28055; Fax: +49-9131-85-28060. Academic Editors: Gang Dong and William Tsang Received: 18 May 2015 / Accepted: 9 September 2015 / Published: 11 September 2015 Abstract: Cilia and flagella are highly conserved and important microtubule-based organelles that project from the surface of eukaryotic cells and act as antennae to sense extracellular signals. Moreover, cilia have emerged as key players in numerous physiological, developmental, and sensory processes such as hearing, olfaction, and photoreception. Genetic defects in ciliary proteins responsible for cilia formation, maintenance, or function underlie a wide array of human diseases like deafness, anosmia, and retinal degeneration in sensory systems. Impairment of more than one sensory organ results in numerous syndromic ciliary disorders like the autosomal recessive genetic diseases Bardet-Biedl and Usher syndrome. Here we describe the structure and distinct functional roles of cilia in sensory organs like the inner ear, the olfactory epithelium, and the retina of the mouse. The spectrum of ciliary function in fundamental cellular processes highlights the importance of elucidating ciliopathy-related proteins in order to find novel potential therapies. -

In This Table Protein Name, Uniprot Code, Gene Name P-Value
Supplementary Table S1: In this table protein name, uniprot code, gene name p-value and Fold change (FC) for each comparison are shown, for 299 of the 301 significantly regulated proteins found in both comparisons (p-value<0.01, fold change (FC) >+/-0.37) ALS versus control and FTLD-U versus control. Two uncharacterized proteins have been excluded from this list Protein name Uniprot Gene name p value FC FTLD-U p value FC ALS FTLD-U ALS Cytochrome b-c1 complex P14927 UQCRB 1.534E-03 -1.591E+00 6.005E-04 -1.639E+00 subunit 7 NADH dehydrogenase O95182 NDUFA7 4.127E-04 -9.471E-01 3.467E-05 -1.643E+00 [ubiquinone] 1 alpha subcomplex subunit 7 NADH dehydrogenase O43678 NDUFA2 3.230E-04 -9.145E-01 2.113E-04 -1.450E+00 [ubiquinone] 1 alpha subcomplex subunit 2 NADH dehydrogenase O43920 NDUFS5 1.769E-04 -8.829E-01 3.235E-05 -1.007E+00 [ubiquinone] iron-sulfur protein 5 ARF GTPase-activating A0A0C4DGN6 GIT1 1.306E-03 -8.810E-01 1.115E-03 -7.228E-01 protein GIT1 Methylglutaconyl-CoA Q13825 AUH 6.097E-04 -7.666E-01 5.619E-06 -1.178E+00 hydratase, mitochondrial ADP/ATP translocase 1 P12235 SLC25A4 6.068E-03 -6.095E-01 3.595E-04 -1.011E+00 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 MIC J3QTA6 CHCHD6 1.090E-04 -5.913E-01 2.124E-03 -5.948E-01 Protein kinase C and casein Q9BY11 PACSIN1 3.837E-03 -5.863E-01 3.680E-06 -1.824E+00 kinase substrate in neurons protein 1 Tubulin polymerization- O94811 TPPP 6.466E-03 -5.755E-01 6.943E-06 -1.169E+00 promoting protein MIC C9JRZ6 CHCHD3 2.912E-02 -6.187E-01 2.195E-03 -9.781E-01 Mitochondrial 2- -

Mutational Mechanisms That Activate Wnt Signaling and Predict Outcomes in Colorectal Cancer Patients William Hankey1, Michael A
Published OnlineFirst December 6, 2017; DOI: 10.1158/0008-5472.CAN-17-1357 Cancer Genome and Epigenome Research Mutational Mechanisms That Activate Wnt Signaling and Predict Outcomes in Colorectal Cancer Patients William Hankey1, Michael A. McIlhatton1, Kenechi Ebede2, Brian Kennedy3, Baris Hancioglu3, Jie Zhang4, Guy N. Brock3, Kun Huang4, and Joanna Groden1 Abstract APC biallelic loss-of-function mutations are the most prevalent also exhibiting unique changes in pathways related to prolifera- genetic changes in colorectal tumors, but it is unknown whether tion, cytoskeletal organization, and apoptosis. Apc-mutant ade- these mutations phenocopy gain-of-function mutations in the nomas were characterized by increased expression of the glial CTNNB1 gene encoding b-catenin that also activate canonical nexin Serpine2, the human ortholog, which was increased in WNT signaling. Here we demonstrate that these two mutational advanced human colorectal tumors. Our results support the mechanisms are not equivalent. Furthermore, we show how hypothesis that APC-mutant colorectal tumors are transcription- differences in gene expression produced by these different ally distinct from APC-wild-type colorectal tumors with canonical mechanisms can stratify outcomes in more advanced human WNT signaling activated by other mechanisms, with possible colorectal cancers. Gene expression profiling in Apc-mutant and implications for stratification and prognosis. Ctnnb1-mutant mouse colon adenomas identified candidate Significance: These findings suggest that colon adenomas genes for subsequent evaluation of human TCGA (The Cancer driven by APC mutations are distinct from those driven by WNT Genome Atlas) data for colorectal cancer outcomes. Transcrip- gain-of-function mutations, with implications for identifying tional patterns exhibited evidence of activated canonical Wnt at-risk patients with advanced disease based on gene expression signaling in both types of adenomas, with Apc-mutant adenomas patterns. -

"The Genecards Suite: from Gene Data Mining to Disease Genome Sequence Analyses". In: Current Protocols in Bioinformat
The GeneCards Suite: From Gene Data UNIT 1.30 Mining to Disease Genome Sequence Analyses Gil Stelzer,1,5 Naomi Rosen,1,5 Inbar Plaschkes,1,2 Shahar Zimmerman,1 Michal Twik,1 Simon Fishilevich,1 Tsippi Iny Stein,1 Ron Nudel,1 Iris Lieder,2 Yaron Mazor,2 Sergey Kaplan,2 Dvir Dahary,2,4 David Warshawsky,3 Yaron Guan-Golan,3 Asher Kohn,3 Noa Rappaport,1 Marilyn Safran,1 and Doron Lancet1,6 1Department of Molecular Genetics, Weizmann Institute of Science, Rehovot, Israel 2LifeMap Sciences Ltd., Tel Aviv, Israel 3LifeMap Sciences Inc., Marshfield, Massachusetts 4Toldot Genetics Ltd., Hod Hasharon, Israel 5These authors contributed equally to the paper 6Corresponding author GeneCards, the human gene compendium, enables researchers to effectively navigate and inter-relate the wide universe of human genes, diseases, variants, proteins, cells, and biological pathways. Our recently launched Version 4 has a revamped infrastructure facilitating faster data updates, better-targeted data queries, and friendlier user experience. It also provides a stronger foundation for the GeneCards suite of companion databases and analysis tools. Improved data unification includes gene-disease links via MalaCards and merged biological pathways via PathCards, as well as drug information and proteome expression. VarElect, another suite member, is a phenotype prioritizer for next-generation sequencing, leveraging the GeneCards and MalaCards knowledgebase. It au- tomatically infers direct and indirect scored associations between hundreds or even thousands of variant-containing genes and disease phenotype terms. Var- Elect’s capabilities, either independently or within TGex, our comprehensive variant analysis pipeline, help prepare for the challenge of clinical projects that involve thousands of exome/genome NGS analyses. -

Dynein Activators and Adaptors at a Glance Mara A
© 2019. Published by The Company of Biologists Ltd | Journal of Cell Science (2019) 132, jcs227132. doi:10.1242/jcs.227132 CELL SCIENCE AT A GLANCE Dynein activators and adaptors at a glance Mara A. Olenick and Erika L. F. Holzbaur* ABSTRACT ribonucleoprotein particles for BICD2, and signaling endosomes for Cytoplasmic dynein-1 (hereafter dynein) is an essential cellular motor Hook1. In this Cell Science at a Glance article and accompanying that drives the movement of diverse cargos along the microtubule poster, we highlight the conserved structural features found in dynein cytoskeleton, including organelles, vesicles and RNAs. A long- activators, the effects of these activators on biophysical parameters, standing question is how a single form of dynein can be adapted to a such as motor velocity and stall force, and the specific intracellular wide range of cellular functions in both interphase and mitosis. functions they mediate. – Recent progress has provided new insights dynein interacts with a KEY WORDS: BICD2, Cytoplasmic dynein, Dynactin, Hook1, group of activating adaptors that provide cargo-specific and/or Microtubule motors, Trafficking function-specific regulation of the motor complex. Activating adaptors such as BICD2 and Hook1 enhance the stability of the Introduction complex that dynein forms with its required activator dynactin, leading Microtubule-based transport is vital to cellular development and to highly processive motility toward the microtubule minus end. survival. Microtubules provide a polarized highway to facilitate Furthermore, activating adaptors mediate specific interactions of the active transport by the molecular motors dynein and kinesin. While motor complex with cargos such as Rab6-positive vesicles or many types of kinesins drive transport toward microtubule plus- ends, there is only one major form of dynein, cytoplasmic dynein-1, University of Pennsylvania Perelman School of Medicine, Philadelphia, PA 19104, which drives the trafficking of a wide array of minus-end-directed USA. -

A Stable Sub-Complex Between GCP4, GCP5 and GCP6 Promotes The
© 2020. Published by The Company of Biologists Ltd | Journal of Cell Science (2020) 133, jcs244368. doi:10.1242/jcs.244368 RESEARCH ARTICLE A stable sub-complex between GCP4, GCP5 and GCP6 promotes the assembly of γ-tubulin ring complexes Laurence Haren, Dorian Farache*, Laurent Emorine and Andreas Merdes‡ ABSTRACT grip1 and grip2 motifs, corresponding to the N-terminal and γ-Tubulin is the main protein involved in the nucleation of C-terminal halves of GCP4, the smallest GCP (Gunawardane et al., microtubules in all eukaryotes. It forms two different complexes with 2000; Guillet et al., 2011). The crystallographic structure of GCP4 α proteins of the GCP family (γ-tubulin complex proteins): γ-tubulin shows that these domains correspond to bundles of -helices. The small complexes (γTuSCs) that contain γ-tubulin, and GCPs 2 and 3; other GCPs contain additional specific sequences, mainly at the and γ-tubulin ring complexes (γTuRCs) that contain multiple γTuSCs extreme N-terminus or in the region that links the grip1 and grip2 in addition to GCPs 4, 5 and 6. Whereas the structure and assembly motifs, as in GCPs 5 and 6 (Guillet et al., 2011; Farache et al., 2016). properties of γTuSCs have been intensively studied, little is known Depletion of GCP2 or GCP3 leads to severe spindle about the assembly of γTuRCs and the specific roles of GCPs 4, 5 abnormalities, and depleted cells are not viable. Depletion of and 6. Here, we demonstrate that two copies of GCP4 and one copy GCP4, 5 or 6 can be tolerated in fission yeast or in somatic cells of Drosophila each of GCP5 and GCP6 form a salt (KCl)-resistant sub-complex but not in vertebrates, where removal of either of these γ within the γTuRC that assembles independently of the presence of GCPs prevents the formation of the TuRC and provokes spindle γTuSCs. -
Inter-Chromosomal Variation in the Pattern of Human Population Genetic Structure Tesfaye M
PRIMARY RESEARCH Inter-chromosomal variation in the pattern of human population genetic structure Tesfaye M. Baye* Cincinnati Children’s Hospital Medical Center, Division of Asthma Research, Department of Pediatrics, University of Cincinnati, 3333 Burnet Avenue, Cincinnati, OH 45229, USA *Correspondence to: Tel: þ1 513 803 2766; Fax: þ1 513 636 1657; E-mail: [email protected] Date received (in revised form): 1st March 2011 Abstract Emerging technologies now make it possible to genotype hundreds of thousands of genetic variations in individuals, across the genome. The study of loci at finer scales will facilitate the understanding of genetic variation at genomic and geographic levels. We examined global and chromosomal variations across HapMap populations using 3.7 million single nucleotide polymorphisms to search for the most stratified genomic regions of human populations and linked these regions to ontological annotation and functional network analysis. To achieve this, we used five complementary statistical and genetic network procedures: principal component (PC), cluster, discriminant, fix- ation index (FST) and network/pathway analyses. At the global level, the first two PC scores were sufficient to account for major population structure; however, chromosomal level analysis detected subtle forms of population structure within continental populations, and as many as 31 PCs were required to classify individuals into homo- geneous groups. Using recommended population ancestry differentiation measures, a total of 126 regions of the genome were catalogued. Gene ontology and networks analyses revealed that these regions included the genes encoding oculocutaneous albinism II (OCA2), hect domain and RLD 2 (HERC2), ectodysplasin A receptor (EDAR) and solute carrier family 45, member 2 (SLC45A2).