Evidence for Non-Methanogenic Metabolisms in Globally Distributed Archaeal Clades Basal
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Developing a Genetic Manipulation System for the Antarctic Archaeon, Halorubrum Lacusprofundi: Investigating Acetamidase Gene Function
www.nature.com/scientificreports OPEN Developing a genetic manipulation system for the Antarctic archaeon, Halorubrum lacusprofundi: Received: 27 May 2016 Accepted: 16 September 2016 investigating acetamidase gene Published: 06 October 2016 function Y. Liao1, T. J. Williams1, J. C. Walsh2,3, M. Ji1, A. Poljak4, P. M. G. Curmi2, I. G. Duggin3 & R. Cavicchioli1 No systems have been reported for genetic manipulation of cold-adapted Archaea. Halorubrum lacusprofundi is an important member of Deep Lake, Antarctica (~10% of the population), and is amendable to laboratory cultivation. Here we report the development of a shuttle-vector and targeted gene-knockout system for this species. To investigate the function of acetamidase/formamidase genes, a class of genes not experimentally studied in Archaea, the acetamidase gene, amd3, was disrupted. The wild-type grew on acetamide as a sole source of carbon and nitrogen, but the mutant did not. Acetamidase/formamidase genes were found to form three distinct clades within a broad distribution of Archaea and Bacteria. Genes were present within lineages characterized by aerobic growth in low nutrient environments (e.g. haloarchaea, Starkeya) but absent from lineages containing anaerobes or facultative anaerobes (e.g. methanogens, Epsilonproteobacteria) or parasites of animals and plants (e.g. Chlamydiae). While acetamide is not a well characterized natural substrate, the build-up of plastic pollutants in the environment provides a potential source of introduced acetamide. In view of the extent and pattern of distribution of acetamidase/formamidase sequences within Archaea and Bacteria, we speculate that acetamide from plastics may promote the selection of amd/fmd genes in an increasing number of environmental microorganisms. -

Growth of Sedimentary Bathyarchaeota on Lignin As an Energy Source
Growth of sedimentary Bathyarchaeota on lignin as an energy source Tiantian Yua,b,1, Weichao Wuc,d,1, Wenyue Lianga,b, Mark Alexander Levere, Kai-Uwe Hinrichsc,d, and Fengping Wanga,b,2 aState Key Laboratory of Microbial Metabolism, School of Life Sciences and Biotechnology, Shanghai Jiao Tong University, 200240 Shanghai, China; bState Key Laboratory of Ocean Engineering, Shanghai Jiao Tong University, 200240 Shanghai, China; cOrganic Geochemistry Group, MARUM-Center for Marine Environmental Sciences, University of Bremen, 28359 Bremen, Germany; dDepartment of Geosciences, University of Bremen, 28359 Bremen, Germany; and eInstitute of Biogeochemistry and Pollutant Dynamics, Department of Environmental Systems Science, Swiss Federal Institute of Technology Zurich, 8092 Zurich, Switzerland Edited by Edward F. DeLong, University of Hawaii at Manoa, Honolulu, HI, and approved April 16, 2018 (received for review October 30, 2017) Members of the archaeal phylum Bathyarchaeota are among the other hand, acetogenesis from H2/CO2 has been proposed for most abundant microorganisms on Earth. Although versatile met- some lineages of Bathyarchaeota (3). The reductive acetyl-CoA abolic capabilities such as acetogenesis, methanogenesis, and fer- pathway for carbon fixation has been identified in most of the mentation have been suggested for bathyarchaeotal members, no obtained bathyarchaeotal genomes (3, 12, 14). Among these, direct confirmation of these metabolic functions has been achieved members of the bathyarchaeotal subgroups Bathy-3 and Bathy- through growth of Bathyarchaeota in the laboratory. Here we dem- 8 have been suggested to be capable of methanogenesis (12). All onstrate, on the basis of gene-copy numbers and probing of ar- the above studies indicate great versatility in the metabolic po- chaeal lipids, the growth of Bathyarchaeota subgroup Bathy-8 in tentials of Bathyarchaeota, with some lineages possibly being enrichments of estuarine sediments with the biopolymer lignin. -

Phylogenetics of Archaeal Lipids Amy Kelly 9/27/2006 Outline
Phylogenetics of Archaeal Lipids Amy Kelly 9/27/2006 Outline • Phlogenetics of Archaea • Phlogenetics of archaeal lipids • Papers Phyla • Two? main phyla – Euryarchaeota • Methanogens • Extreme halophiles • Extreme thermophiles • Sulfate-reducing – Crenarchaeota • Extreme thermophiles – Korarchaeota? • Hyperthermophiles • indicated only by environmental DNA sequences – Nanoarchaeum? • N. equitans a fast evolving euryarchaeal lineage, not novel, early diverging archaeal phylum – Ancient archael group? • In deepest brances of Crenarchaea? Euryarchaea? Archaeal Lipids • Methanogens – Di- and tetra-ethers of glycerol and isoprenoid alcohols – Core mostly archaeol or caldarchaeol – Core sometimes sn-2- or Images removed due to sn-3-hydroxyarchaeol or copyright considerations. macrocyclic archaeol –PMI • Halophiles – Similar to methanogens – Exclusively synthesize bacterioruberin • Marine Crenarchaea Depositional Archaeal Lipids Biological Origin Environment Crocetane methanotrophs? methane seeps? methanogens, PMI (2,6,10,15,19-pentamethylicosane) methanotrophs hypersaline, anoxic Squalane hypersaline? C31-C40 head-to-head isoprenoids Smit & Mushegian • “Lost” enzymes of MVA pathway must exist – Phosphomevalonate kinase (PMK) – Diphosphomevalonate decarboxylase – Isopentenyl diphosphate isomerase (IPPI) Kaneda et al. 2001 Rohdich et al. 2001 Boucher et al. • Isoprenoid biosynthesis of archaea evolved through a combination of processes – Co-option of ancestral enzymes – Modification of enzymatic specificity – Orthologous and non-orthologous gene -

Characteristics and Metabolic Patterns of Soil Methanogenic Archaea Communities in the High Latitude Natural Wetlands of China
Characteristics and Metabolic Patterns of Soil Methanogenic Archaea Communities in the High Latitude Natural Wetlands of China Di Wu Northeast Forestry University Caihong Zhao Northeast Forestry University Hui Bai Forestry Science Research Institute of Heilongjiang Province Fujuan Feng Northeast Forestry University Xin Sui Heilongjiang University Guangyu Sun ( [email protected] ) Northeast Forestry University Research article Keywords: Wetlands, Methanogens, Community diversity, Indicator species, Methanogenic metabolic patterns Posted Date: August 12th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-54821/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/18 Abstract Background: Soil methanogenic microorganisms are one of the primary methane-producing microbes in wetlands. However, we still poorly understand the community characteristic and metabolic patterns of these microorganisms according to vegetation type and seasonal changes. Therefore, to better elucidate the effects of the vegetation type and seasonal factors on the methanogenic community structure and metabolic patterns, we detected the characteristics of the soil methanogenic mcrA gene from three types of natural wetlands in different seasons in the Xiaoxing'an Mountain region, China. Result: The results indicated that the distribution of Methanobacteriaceae (hydrogenotrophic methanogens) was higher in winter, while Methanosarcinaceae and Methanosaetaceae accounted for a higher proportion in summer. Hydrogenotrophic methanogenesis was the dominant trophic pattern in each wetland. The results of principal coordinate analysis and cluster analysis showed that the vegetation type considerably inuenced the methanogenic community composition. The methanogenic community structure in the Betula platyphylla – Larix gmelinii wetland was relatively different from the structure of the other two wetland types. -

Proteome Cold-Shock Response in the Extremely Acidophilic Archaeon, Cuniculiplasma Divulgatum
microorganisms Article Proteome Cold-Shock Response in the Extremely Acidophilic Archaeon, Cuniculiplasma divulgatum Rafael Bargiela 1 , Karin Lanthaler 1,2, Colin M. Potter 1,2 , Manuel Ferrer 3 , Alexander F. Yakunin 1,2, Bela Paizs 1,2, Peter N. Golyshin 1,2 and Olga V. Golyshina 1,2,* 1 School of Natural Sciences, Bangor University, Deiniol Rd, Bangor LL57 2UW, UK; [email protected] (R.B.); [email protected] (K.L.); [email protected] (C.M.P.); [email protected] (A.F.Y.); [email protected] (B.P.); [email protected] (P.N.G.) 2 Centre for Environmental Biotechnology, Bangor University, Deiniol Rd, Bangor LL57 2UW, UK 3 Systems Biotechnology Group, Department of Applied Biocatalysis, CSIC—Institute of Catalysis, Marie Curie 2, 28049 Madrid, Spain; [email protected] * Correspondence: [email protected]; Tel.: +44-1248-388607; Fax: +44-1248-382569 Received: 27 April 2020; Accepted: 15 May 2020; Published: 19 May 2020 Abstract: The archaeon Cuniculiplasma divulgatum is ubiquitous in acidic environments with low-to-moderate temperatures. However, molecular mechanisms underlying its ability to thrive at lower temperatures remain unexplored. Using mass spectrometry (MS)-based proteomics, we analysed the effect of short-term (3 h) exposure to cold. The C. divulgatum genome encodes 2016 protein-coding genes, from which 819 proteins were identified in the cells grown under optimal conditions. In line with the peptidolytic lifestyle of C. divulgatum, its intracellular proteome revealed the abundance of proteases, ABC transporters and cytochrome C oxidase. From 747 quantifiable polypeptides, the levels of 582 proteins showed no change after the cold shock, whereas 104 proteins were upregulated suggesting that they might be contributing to cold adaptation. -

Picrophilus Oshimae and Picrophilus Tomdus Fam. Nov., Gen. Nov., Sp. Nov
INTERNATIONALJOURNAL OF SYSTEMATICBACTERIOLOGY, July 1996, p. 814-816 Vol. 46, No. 3 0020-77 13/96/$04.00+0 Copyright 0 1996, International Union of Microbiological Societies Picrophilus oshimae and Picrophilus tomdus fam. nov., gen. nov., sp. nov., Two Species of Hyperacidophilic, Thermophilic, Heterotrophic, Aerobic Archaea CHRISTA SCHLEPER, GABRIELA PUHLER, HANS- PETER KLENK, AND WOLFRAM ZILLIG* Max Plank Institut fur Biochemie, 0-82152 Martinsried, Germany We describe two species of hyperacidophilic, thermophilic, heterotrophic, aerobic archaea that were isolated from solfataric hydrothermal areas in Hokkaido, Japan. These organisms, Picrophilus oshimae and Picrophilus torridus, represent a novel genus and a novel family, the Picrophilaceae, in the kingdom Euryarchaeota and the order Thermoplasmales. Both of these bacteria are more acidophilic than the genus Thermoplasma since they are able to grow at about pH 0. The moderately thermophilic, hyperacidophilic, aerobic ar- which comprises acid-loving (i.e., hyperacidophilic) organisms. chaea (archaebacteria) (7) Picrophilus oshimae and Picrophilus Separation of these taxa is justified by their phylogenetic dis- rorridus, which have been described previously (4, 5), were tance, (9.5% difference in the 16s rRNA sequences of mem- isolated from moderately hot hydrothermal areas in solfataric bers of the Picrophilaceae and T. acidophilum), by the lack of fields in Hokkaido, Japan. One of the sources of isolation was immunochemical cross-reactions in Ouchterlony immunodif- a solfataric spring which had a temperature of 53°C and a pH fusion assays between the RNA polymerases of P. oshimae and of 2.2, and the other was a rather dry hot soil which had a pH T. acidophilum, which also do not occur between members of of <OS. -
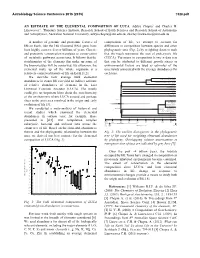
An Estimate of the Elemental Composition of Luca
Astrobiology Science Conference 2015 (2015) 7328.pdf AN ESTIMATE OF THE ELEMENTAL COMPOSITION OF LUCA. Aditya Chopra1 and Charles H. Lineweaver1, 1Planetary Science Institute, Research School of Earth Sciences and Research School of Astronomy and Astrophysics, Australian National University, [email protected], [email protected] A number of genomic and proteomic features of composition of life, we attempt to account for life on Earth, like the 16S ribosomal RNA gene, have differences in composition between species and other been highly conserved over billions of years. Genetic phylogenetic taxa (Fig. 2) by weighting datasets such and proteomic conservation translates to conservation that the result represents the root of prokaryotic life of metabolic pathways across taxa. It follows that the (LUCA). Variations in composition between data sets stoichiometry of the elements that make up some of that can be attributed to different growth stages or the biomolecules will be conserved. By extension, the environmental factors are used as estimates of the elemental make up of the whole organism is a uncertainty associated with the average abundances for relatively conserved feature of life on Earth [1,2]. each taxa. We describe how average bulk elemental Euryarchaeota 1a Methanococci, Methanobacteria, Methanopyri 7 Euryarchaeota 1b abundances in extant life can yield an indirect estimate 6 Thermoplasmata, Methanomicrobia, Halobacteria, Archaeoglobi Euryarchaeota 2 4 Thermococci of relative abundances of elements in the Last Crenarchaeota Sulfolobus, Thermoproteus ? Thaumarchaeota Universal Common Ancestor (LUCA). The results Cenarchaeum ? Korarchaeota ARMAN could give us important hints about the stoichiometry ? 2 Archaeal Richmond Mine Acidophilic Nanoorganisms Nanoarchaeota of the environment where LUCA existed and perhaps Archaea Terrabacteria clues to the processes involved in the origin and early Actinobacteria, Deinococcus-Thermus, 1 Cyanobacteria, Life Chloroflexi, 9 evolution of life [3]. -

Archaeal Adaptation to Higher Temperatures Revealed by Genomic Sequence of Thermoplasma Volcanium
Archaeal adaptation to higher temperatures revealed by genomic sequence of Thermoplasma volcanium Tsuyoshi Kawashima*†, Naoki Amano*†‡, Hideaki Koike*†, Shin-ichi Makino†, Sadaharu Higuchi†, Yoshie Kawashima-Ohya†, Koji Watanabe§, Masaaki Yamazaki§, Keiichi Kanehori¶, Takeshi Kawamotoʈ, Tatsuo Nunoshiba**, Yoshihiro Yamamoto††, Hironori Aramaki‡‡, Kozo Makino§§, and Masashi Suzuki†¶¶ †National Institute of Bioscience and Human Technology, Core Research for Evolutional Science and Technology Centre of Structural Biology, 1-1 Higashi, Tsukuba 305-0046, Japan; ‡Doctoral Program in Medical Sciences, University of Tsukuba, 1-1-1 Tennohdai, Tsukuba 305-0006, Japan; §Bioscience Research Laboratory, Fujiya, 228 Soya, Hadano 257-0031, Japan; ¶DNA Analysis Department, Techno Research Laboratory, Hitachi Science Systems, 1-280 Higashi-Koigakubo, Kokubunji 185-8601, Japan; ʈDepartment of Biochemistry, Hiroshima University, School of Dentistry, 1-2-3 Kasumi, Minami-ku, Hiroshima 734-8553, Japan; **Department of Molecular and Cellular Biology, Biological Institute, Graduate School of Science, Tohoku University, Sendai 980-8578, Japan; ††Department of Genetics, Hyogo College of Medicine, Nishinomiya 663-8501, Japan; ‡‡Department of Molecular Biology, Daiichi College of Pharmaceutical Science, 22-1 Tamagawa-cho, Minami-ku, Fukuoka 815-8511, Japan; and §§Department of Molecular Microbiology, The Research Institute of Microbial Diseases, Osaka University, 3-1 Yamadaoka, Suita 565-0871, Japan Edited by Aaron Klug, Royal Society of London, London, United Kingdom, and approved October 16, 2000 (received for review August 18, 2000) The complete genomic sequence of the archaeon Thermoplasma contigs. The remaining gaps were bridged by DNA fragments volcanium, possessing optimum growth temperature (OGT) of constructed using the PCR. The average repetition in sequencing 60°C, is reported. By systematically comparing this genomic se- the same base positions was 13-fold. -

The Main (Glyco) Phospholipid (MPL) of Thermoplasma Acidophilum
International Journal of Molecular Sciences Review The Main (Glyco) Phospholipid (MPL) of Thermoplasma acidophilum Hans-Joachim Freisleben 1,2 1 Goethe-Universität, Gustav-Embden-Zentrum, Laboratory of Microbiological Chemistry, Theodor-Stern-Kai 7, D-60590 Frankfurt am Main, Germany; [email protected] 2 Universitas Indonesia, Medical Research Unit, Faculty of Medicine, Jalan Salemba Raya 6, Jakarta 10430, Indonesia Received: 19 September 2019; Accepted: 18 October 2019; Published: 21 October 2019 Abstract: The main phospholipid (MPL) of Thermoplasma acidophilum DSM 1728 was isolated, purified and physico-chemically characterized by differential scanning calorimetry (DSC)/differential thermal analysis (DTA) for its thermotropic behavior, alone and in mixtures with other lipids, cholesterol, hydrophobic peptides and pore-forming ionophores. Model membranes from MPL were investigated; black lipid membrane, Langmuir-Blodgett monolayer, and liposomes. Laboratory results were compared to computer simulation. MPL forms stable and resistant liposomes with highly proton-impermeable membrane and mixes at certain degree with common bilayer-forming lipids. Monomeric bacteriorhodopsin and ATP synthase from Micrococcus luteus were co-reconstituted and light-driven ATP synthesis measured. This review reports about almost four decades of research on Thermoplasma membrane and its MPL as well as transfer of this research to Thermoplasma species recently isolated from Indonesian volcanoes. Keywords: Thermoplasma acidophilum; Thermoplasma volcanium; -

Different Proteins Mediate Step-Wise Chromosome Architectures in 2 Thermoplasma Acidophilum and Pyrobaculum Calidifontis
bioRxiv preprint doi: https://doi.org/10.1101/2020.03.13.982959; this version posted May 4, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Different Proteins Mediate Step-wise Chromosome Architectures in 2 Thermoplasma acidophilum and Pyrobaculum calidifontis 3 4 5 Hugo Maruyama1†*, Eloise I. Prieto2†, Takayuki Nambu1, Chiho Mashimo1, Kosuke 6 Kashiwagi3, Toshinori Okinaga1, Haruyuki Atomi4, Kunio Takeyasu5 7 8 1 Department of Bacteriology, Osaka Dental University, Hirakata, Japan 9 2 National Institute of Molecular Biology and Biotechnology, University of the Philippines 10 Diliman, Quezon City, Philippines 11 3 Department of Fixed Prosthodontics, Osaka Dental University, Hirakata, Japan 12 4 Department of Synthetic Chemistry and Biological Chemistry, Graduate School of Engineering, 13 Kyoto University, Kyoto, Japan 14 5 Graduate School of Biostudies, Kyoto University, Kyoto, Japan 15 † These authors have contributed equally to this work 16 17 * Correspondence: 18 Hugo Maruyama 19 [email protected]; [email protected] 20 21 Keywords: archaea, higher-order chromosome structure, nucleoid, chromatin, HTa, histone, 22 transcriptional regulator, horizontal gene transfer 23 Running Title: Step-wise chromosome architecture in Archaea 24 Manuscript length: 6955 words 25 Number of Figures: 7 26 Number of Tables: 3 bioRxiv preprint doi: https://doi.org/10.1101/2020.03.13.982959; this version posted May 4, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

(Agus Kurnia)Ok
HAYATI Journal of Biosciences September 2012 Available online at: Vol. 19 No. 3, p 150-154 http://journal.ipb.ac.id/index.php/hayati EISSN: 2086-4094 DOI: 10.4308/hjb.19.3.150 SHORT COMMUNICATION Archaeal Life on Tangkuban Perahu- Sampling and Culture Growth in Indonesian Laboratories SRI HANDAYANI1, IMAN SANTOSO1, HANS-JOACHIM FREISLEBEN2∗, HARALD HUBER3, ANDI1, FERY ARDIANSYAH1, CENMI MULYANTO1, ZESSINDA LUTHFA1, ROSARI SALEH1, SERUNI KUSUMA UDYANINGSIH FREISLEBEN1, SEPTELIA INAWATI WANANDI2, MICHAEL THOMM3 1Faculty of Mathematics and Natural Sciences, Universitas Indonesia, Jakarta-Depok, Jakarta 10430, Indonesia 2Faculty of Medicine, Universitas Indonesia, Jalan Salemba Raya No. 6, Jakarta 10430, Indonesia 3Department of Microbiology, Archaea Centre, University of Regensburg, Germany Received May 9, 2012/Accepted September 21, 2012 The aim of the expedition to Tangkuban Perahu, West Java was to obtain archaeal samples from the solfatara fields located in Domas crater. This was one of the places, where scientists from the University of Regensburg Germany had formerly isolated Indonesian archaea, especially Thermoplasma and Sulfolobus species but not fully characterized. We collected five samples from mud holes with temperatures from 57 to 88 oC and pH of 1.5-2. A portion of each sample was grown at the University of Regensburg in modified Allen’s medium at 80 oC. From four out of five samples enrichment cultures were obtained, autotrophically on elemental sulphur and heterotrophically on sulfur and yeast extract; electron micrographs are presented. In the laboratories of Universitas Indonesia the isolates were cultured at 55-60 oC in order to grow tetraetherlipid synthesizing archaea, both Thermoplasmatales and Sulfolobales. Here, we succeeded to culture the same type of archaeal cells, which had been cultured in Regensburg, probably a Sulfolobus species and in Freundt’s medium, Thermoplasma species. -

Pan-Genome Analysis and Ancestral State Reconstruction Of
www.nature.com/scientificreports OPEN Pan‑genome analysis and ancestral state reconstruction of class halobacteria: probability of a new super‑order Sonam Gaba1,2, Abha Kumari2, Marnix Medema 3 & Rajeev Kaushik1* Halobacteria, a class of Euryarchaeota are extremely halophilic archaea that can adapt to a wide range of salt concentration generally from 10% NaCl to saturated salt concentration of 32% NaCl. It consists of the orders: Halobacteriales, Haloferaciales and Natriabales. Pan‑genome analysis of class Halobacteria was done to explore the core (300) and variable components (Softcore: 998, Cloud:36531, Shell:11784). The core component revealed genes of replication, transcription, translation and repair, whereas the variable component had a major portion of environmental information processing. The pan‑gene matrix was mapped onto the core‑gene tree to fnd the ancestral (44.8%) and derived genes (55.1%) of the Last Common Ancestor of Halobacteria. A High percentage of derived genes along with presence of transformation and conjugation genes indicate the occurrence of horizontal gene transfer during the evolution of Halobacteria. A Core and pan‑gene tree were also constructed to infer a phylogeny which implicated on the new super‑order comprising of Natrialbales and Halobacteriales. Halobacteria1,2 is a class of phylum Euryarchaeota3 consisting of extremely halophilic archaea found till date and contains three orders namely Halobacteriales4,5 Haloferacales5 and Natrialbales5. Tese microorganisms are able to dwell at wide range of salt concentration generally from 10% NaCl to saturated salt concentration of 32% NaCl6. Halobacteria, as the name suggests were once considered a part of a domain "Bacteria" but with the discovery of the third domain "Archaea" by Carl Woese et al.7, it became part of Archaea.