(Agus Kurnia)Ok
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Developing a Genetic Manipulation System for the Antarctic Archaeon, Halorubrum Lacusprofundi: Investigating Acetamidase Gene Function
www.nature.com/scientificreports OPEN Developing a genetic manipulation system for the Antarctic archaeon, Halorubrum lacusprofundi: Received: 27 May 2016 Accepted: 16 September 2016 investigating acetamidase gene Published: 06 October 2016 function Y. Liao1, T. J. Williams1, J. C. Walsh2,3, M. Ji1, A. Poljak4, P. M. G. Curmi2, I. G. Duggin3 & R. Cavicchioli1 No systems have been reported for genetic manipulation of cold-adapted Archaea. Halorubrum lacusprofundi is an important member of Deep Lake, Antarctica (~10% of the population), and is amendable to laboratory cultivation. Here we report the development of a shuttle-vector and targeted gene-knockout system for this species. To investigate the function of acetamidase/formamidase genes, a class of genes not experimentally studied in Archaea, the acetamidase gene, amd3, was disrupted. The wild-type grew on acetamide as a sole source of carbon and nitrogen, but the mutant did not. Acetamidase/formamidase genes were found to form three distinct clades within a broad distribution of Archaea and Bacteria. Genes were present within lineages characterized by aerobic growth in low nutrient environments (e.g. haloarchaea, Starkeya) but absent from lineages containing anaerobes or facultative anaerobes (e.g. methanogens, Epsilonproteobacteria) or parasites of animals and plants (e.g. Chlamydiae). While acetamide is not a well characterized natural substrate, the build-up of plastic pollutants in the environment provides a potential source of introduced acetamide. In view of the extent and pattern of distribution of acetamidase/formamidase sequences within Archaea and Bacteria, we speculate that acetamide from plastics may promote the selection of amd/fmd genes in an increasing number of environmental microorganisms. -

Expression of Cyclomaltodextrinase Gene from Bacillus Halodurans C-125 and Characterization of Its Multisubstrate Specificity
Food Sci. Biotechnol. Vol. 18, No. 3, pp. 776 ~ 781 (2009) ⓒ The Korean Society of Food Science and Technology Expression of Cyclomaltodextrinase Gene from Bacillus halodurans C-125 and Characterization of Its Multisubstrate Specificity Hye-Jeong Kang, Chang-Ku Jeong, Myoung-Uoon Jang, Seung-Ho Choi, Min-Hong Kim1, Jun-Bae Ahn2, Sang-Hwa Lee3, Sook-Ja Jo3, and Tae-Jip Kim* Department of Food Science and Technology, Chungbuk National University, Cheongju, Chungbuk 361-763, Korea 1MH2 Biochemical Co., Ltd., Eumseong, Chungbuk 369-841, Korea 2Department of Food Service Industry, Seowon University, Cheongju, Chungbuk 361-741, Korea 3Department of Food and Nutrition, Seowon University, Cheongju, Chungbuk 361-741, Korea Abstract A putative cyclomaltodextrinase (BHCD) gene was found from the genome of Bacillus halodurans C-125, which encodes 578 amino acids with a predicted molecular mass of 67,279 Da. It shares 42-59% of amino acid sequence identity with common cyclomaltodextrinase (CDase)-family enzymes. The corresponding gene was cloned by polymerase chain reaction (PCR) and the dimeric enzyme with C-terminal 6-histidines was successfully overproduced and purified from recombinant Escherichia coli. BHCD showed the highest activity against β-CD at pH 7.0 and 50oC. Due to its versatile hydrolysis and transglycosylation activities, BHCD has been confirmed as a member of CDases. However, BHCD can be distinguished from other typical CDases on the basis of its novel multisubstrate specificity. While typical CDases have over 10 times higher activity on β-CD than starch or pullulan, the CD-hydrolyzing activity of BHCD is only 2.3 times higher than pullulan. -

Proteome Cold-Shock Response in the Extremely Acidophilic Archaeon, Cuniculiplasma Divulgatum
microorganisms Article Proteome Cold-Shock Response in the Extremely Acidophilic Archaeon, Cuniculiplasma divulgatum Rafael Bargiela 1 , Karin Lanthaler 1,2, Colin M. Potter 1,2 , Manuel Ferrer 3 , Alexander F. Yakunin 1,2, Bela Paizs 1,2, Peter N. Golyshin 1,2 and Olga V. Golyshina 1,2,* 1 School of Natural Sciences, Bangor University, Deiniol Rd, Bangor LL57 2UW, UK; [email protected] (R.B.); [email protected] (K.L.); [email protected] (C.M.P.); [email protected] (A.F.Y.); [email protected] (B.P.); [email protected] (P.N.G.) 2 Centre for Environmental Biotechnology, Bangor University, Deiniol Rd, Bangor LL57 2UW, UK 3 Systems Biotechnology Group, Department of Applied Biocatalysis, CSIC—Institute of Catalysis, Marie Curie 2, 28049 Madrid, Spain; [email protected] * Correspondence: [email protected]; Tel.: +44-1248-388607; Fax: +44-1248-382569 Received: 27 April 2020; Accepted: 15 May 2020; Published: 19 May 2020 Abstract: The archaeon Cuniculiplasma divulgatum is ubiquitous in acidic environments with low-to-moderate temperatures. However, molecular mechanisms underlying its ability to thrive at lower temperatures remain unexplored. Using mass spectrometry (MS)-based proteomics, we analysed the effect of short-term (3 h) exposure to cold. The C. divulgatum genome encodes 2016 protein-coding genes, from which 819 proteins were identified in the cells grown under optimal conditions. In line with the peptidolytic lifestyle of C. divulgatum, its intracellular proteome revealed the abundance of proteases, ABC transporters and cytochrome C oxidase. From 747 quantifiable polypeptides, the levels of 582 proteins showed no change after the cold shock, whereas 104 proteins were upregulated suggesting that they might be contributing to cold adaptation. -

Picrophilus Oshimae and Picrophilus Tomdus Fam. Nov., Gen. Nov., Sp. Nov
INTERNATIONALJOURNAL OF SYSTEMATICBACTERIOLOGY, July 1996, p. 814-816 Vol. 46, No. 3 0020-77 13/96/$04.00+0 Copyright 0 1996, International Union of Microbiological Societies Picrophilus oshimae and Picrophilus tomdus fam. nov., gen. nov., sp. nov., Two Species of Hyperacidophilic, Thermophilic, Heterotrophic, Aerobic Archaea CHRISTA SCHLEPER, GABRIELA PUHLER, HANS- PETER KLENK, AND WOLFRAM ZILLIG* Max Plank Institut fur Biochemie, 0-82152 Martinsried, Germany We describe two species of hyperacidophilic, thermophilic, heterotrophic, aerobic archaea that were isolated from solfataric hydrothermal areas in Hokkaido, Japan. These organisms, Picrophilus oshimae and Picrophilus torridus, represent a novel genus and a novel family, the Picrophilaceae, in the kingdom Euryarchaeota and the order Thermoplasmales. Both of these bacteria are more acidophilic than the genus Thermoplasma since they are able to grow at about pH 0. The moderately thermophilic, hyperacidophilic, aerobic ar- which comprises acid-loving (i.e., hyperacidophilic) organisms. chaea (archaebacteria) (7) Picrophilus oshimae and Picrophilus Separation of these taxa is justified by their phylogenetic dis- rorridus, which have been described previously (4, 5), were tance, (9.5% difference in the 16s rRNA sequences of mem- isolated from moderately hot hydrothermal areas in solfataric bers of the Picrophilaceae and T. acidophilum), by the lack of fields in Hokkaido, Japan. One of the sources of isolation was immunochemical cross-reactions in Ouchterlony immunodif- a solfataric spring which had a temperature of 53°C and a pH fusion assays between the RNA polymerases of P. oshimae and of 2.2, and the other was a rather dry hot soil which had a pH T. acidophilum, which also do not occur between members of of <OS. -

A Korarchaeal Genome Reveals Insights Into the Evolution of the Archaea
A korarchaeal genome reveals insights into the evolution of the Archaea James G. Elkinsa,b, Mircea Podarc, David E. Grahamd, Kira S. Makarovae, Yuri Wolfe, Lennart Randauf, Brian P. Hedlundg, Ce´ line Brochier-Armaneth, Victor Kunini, Iain Andersoni, Alla Lapidusi, Eugene Goltsmani, Kerrie Barryi, Eugene V. Koonine, Phil Hugenholtzi, Nikos Kyrpidesi, Gerhard Wannerj, Paul Richardsoni, Martin Kellerc, and Karl O. Stettera,k,l aLehrstuhl fu¨r Mikrobiologie und Archaeenzentrum, Universita¨t Regensburg, D-93053 Regensburg, Germany; cBiosciences Division, Oak Ridge National Laboratory, Oak Ridge, TN 37831; dDepartment of Chemistry and Biochemistry, University of Texas, Austin, TX 78712; eNational Center for Biotechnology Information, National Library of Medicine, National Institutes of Health, Bethesda, MD 20894; fDepartment of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520; gSchool of Life Sciences, University of Nevada, Las Vegas, NV 89154; hLaboratoire de Chimie Bacte´rienne, Unite´ Propre de Recherche 9043, Centre National de la Recherche Scientifique, Universite´de Provence Aix-Marseille I, 13331 Marseille Cedex 3, France; iU.S. Department of Energy Joint Genome Institute, Walnut Creek, CA 94598; jInstitute of Botany, Ludwig Maximilians University of Munich, D-80638 Munich, Germany; and kInstitute of Geophysics and Planetary Physics, University of California, Los Angeles, CA 90095 Communicated by Carl R. Woese, University of Illinois at Urbana–Champaign, Urbana, IL, April 2, 2008 (received for review January 7, 2008) The candidate division Korarchaeota comprises a group of uncul- and sediment samples from Obsidian Pool as an inoculum. The tivated microorganisms that, by their small subunit rRNA phylog- cultivation system supported the stable growth of a mixed commu- eny, may have diverged early from the major archaeal phyla nity of hyperthermophilic bacteria and archaea including an or- Crenarchaeota and Euryarchaeota. -

Archaeal Adaptation to Higher Temperatures Revealed by Genomic Sequence of Thermoplasma Volcanium
Archaeal adaptation to higher temperatures revealed by genomic sequence of Thermoplasma volcanium Tsuyoshi Kawashima*†, Naoki Amano*†‡, Hideaki Koike*†, Shin-ichi Makino†, Sadaharu Higuchi†, Yoshie Kawashima-Ohya†, Koji Watanabe§, Masaaki Yamazaki§, Keiichi Kanehori¶, Takeshi Kawamotoʈ, Tatsuo Nunoshiba**, Yoshihiro Yamamoto††, Hironori Aramaki‡‡, Kozo Makino§§, and Masashi Suzuki†¶¶ †National Institute of Bioscience and Human Technology, Core Research for Evolutional Science and Technology Centre of Structural Biology, 1-1 Higashi, Tsukuba 305-0046, Japan; ‡Doctoral Program in Medical Sciences, University of Tsukuba, 1-1-1 Tennohdai, Tsukuba 305-0006, Japan; §Bioscience Research Laboratory, Fujiya, 228 Soya, Hadano 257-0031, Japan; ¶DNA Analysis Department, Techno Research Laboratory, Hitachi Science Systems, 1-280 Higashi-Koigakubo, Kokubunji 185-8601, Japan; ʈDepartment of Biochemistry, Hiroshima University, School of Dentistry, 1-2-3 Kasumi, Minami-ku, Hiroshima 734-8553, Japan; **Department of Molecular and Cellular Biology, Biological Institute, Graduate School of Science, Tohoku University, Sendai 980-8578, Japan; ††Department of Genetics, Hyogo College of Medicine, Nishinomiya 663-8501, Japan; ‡‡Department of Molecular Biology, Daiichi College of Pharmaceutical Science, 22-1 Tamagawa-cho, Minami-ku, Fukuoka 815-8511, Japan; and §§Department of Molecular Microbiology, The Research Institute of Microbial Diseases, Osaka University, 3-1 Yamadaoka, Suita 565-0871, Japan Edited by Aaron Klug, Royal Society of London, London, United Kingdom, and approved October 16, 2000 (received for review August 18, 2000) The complete genomic sequence of the archaeon Thermoplasma contigs. The remaining gaps were bridged by DNA fragments volcanium, possessing optimum growth temperature (OGT) of constructed using the PCR. The average repetition in sequencing 60°C, is reported. By systematically comparing this genomic se- the same base positions was 13-fold. -

The Main (Glyco) Phospholipid (MPL) of Thermoplasma Acidophilum
International Journal of Molecular Sciences Review The Main (Glyco) Phospholipid (MPL) of Thermoplasma acidophilum Hans-Joachim Freisleben 1,2 1 Goethe-Universität, Gustav-Embden-Zentrum, Laboratory of Microbiological Chemistry, Theodor-Stern-Kai 7, D-60590 Frankfurt am Main, Germany; [email protected] 2 Universitas Indonesia, Medical Research Unit, Faculty of Medicine, Jalan Salemba Raya 6, Jakarta 10430, Indonesia Received: 19 September 2019; Accepted: 18 October 2019; Published: 21 October 2019 Abstract: The main phospholipid (MPL) of Thermoplasma acidophilum DSM 1728 was isolated, purified and physico-chemically characterized by differential scanning calorimetry (DSC)/differential thermal analysis (DTA) for its thermotropic behavior, alone and in mixtures with other lipids, cholesterol, hydrophobic peptides and pore-forming ionophores. Model membranes from MPL were investigated; black lipid membrane, Langmuir-Blodgett monolayer, and liposomes. Laboratory results were compared to computer simulation. MPL forms stable and resistant liposomes with highly proton-impermeable membrane and mixes at certain degree with common bilayer-forming lipids. Monomeric bacteriorhodopsin and ATP synthase from Micrococcus luteus were co-reconstituted and light-driven ATP synthesis measured. This review reports about almost four decades of research on Thermoplasma membrane and its MPL as well as transfer of this research to Thermoplasma species recently isolated from Indonesian volcanoes. Keywords: Thermoplasma acidophilum; Thermoplasma volcanium; -

Different Proteins Mediate Step-Wise Chromosome Architectures in 2 Thermoplasma Acidophilum and Pyrobaculum Calidifontis
bioRxiv preprint doi: https://doi.org/10.1101/2020.03.13.982959; this version posted May 4, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Different Proteins Mediate Step-wise Chromosome Architectures in 2 Thermoplasma acidophilum and Pyrobaculum calidifontis 3 4 5 Hugo Maruyama1†*, Eloise I. Prieto2†, Takayuki Nambu1, Chiho Mashimo1, Kosuke 6 Kashiwagi3, Toshinori Okinaga1, Haruyuki Atomi4, Kunio Takeyasu5 7 8 1 Department of Bacteriology, Osaka Dental University, Hirakata, Japan 9 2 National Institute of Molecular Biology and Biotechnology, University of the Philippines 10 Diliman, Quezon City, Philippines 11 3 Department of Fixed Prosthodontics, Osaka Dental University, Hirakata, Japan 12 4 Department of Synthetic Chemistry and Biological Chemistry, Graduate School of Engineering, 13 Kyoto University, Kyoto, Japan 14 5 Graduate School of Biostudies, Kyoto University, Kyoto, Japan 15 † These authors have contributed equally to this work 16 17 * Correspondence: 18 Hugo Maruyama 19 [email protected]; [email protected] 20 21 Keywords: archaea, higher-order chromosome structure, nucleoid, chromatin, HTa, histone, 22 transcriptional regulator, horizontal gene transfer 23 Running Title: Step-wise chromosome architecture in Archaea 24 Manuscript length: 6955 words 25 Number of Figures: 7 26 Number of Tables: 3 bioRxiv preprint doi: https://doi.org/10.1101/2020.03.13.982959; this version posted May 4, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -
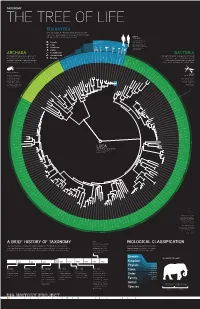
A Brief History of Taxonomy Biological Classification
TAXONOMY THE TREE OF LIFE EUKARYOTA This domain includes all of the plants, animals, and fungi, and some single-celled organisms. Eukaryotes are distinguished by their complex cells, which contain a membrane-enclosed nucleus. Humans Homo sapiens The creatures most familiar to us, Our species, primates in the animals, are members of the Animalia kingdom of the Animalia same kingdom. Eukaryota, is thought to have Fungi Mosquito Red first evolved in Africa about Pufferfish Junglefowl Roundworm Mouse 200,000 years ago. Genetically, Amoebozoa Chimpanzee our closest living relative Plantae is the chimpanzee. Archaeplastida Schizosaccharomyces pombe ARCHAEA Saccharomyces cerevisiae BACTERIA Caenorhabditis briggsae Caenorhabditis elegans Eremothecium gossypii These single-celled prokaryotic organisms often Chromalveolata Dictyostelium discoideum These single-celled prokaryotic organisms were among Cyanidioschyzon merolae live in extreme environmental conditions. Once Excavata Arabidopsis thaliana the first life forms to appear on Earth. Often spherical, Plasmodium falciparum considered to be Bacteria, these microorganisms Cryptosporidium hominis rod-like, or spiral in shape, these microorganisms Thalassiosira pseudonana Oryza sativa Anopheles gambiae Drosophila melanogaster Takifugu rubripes Danio rerio are now recognized as a separate domain of life. Gallus gallus function without a membrane-enclosed cell nucleus. Rattus norvegicus Mus musculus Methanococcus jannaschii Leishmania major Homo sapiens Pan troglodytes Methanococcus maripaludi Thermoanaerobacter -

Genome Sequence of Picrophilus Torridus and Its Implications for Life Around Ph 0
Genome sequence of Picrophilus torridus and its implications for life around pH 0 O. Fu¨ tterer*, A. Angelov*, H. Liesegang*, G. Gottschalk*, C. Schleper†, B. Schepers‡, C. Dock‡, G. Antranikian‡, and W. Liebl*§ *Institut of Microbiology and Genetics, University of Goettingen, Grisebachstrasse 8, D-37075 Goettingen, Germany; †Institut of Microbiology and Genetics, Technical University Darmstadt, Schnittspahnstrasse 10, 64287 D-Darmstadt, Germany; and ‡Technical Microbiology, Technical University Hamburg–Harburg, Kasernenstrasse 12, 21073 D-Hamburg, Germany Edited by Dieter So¨ll, Yale University, New Haven, CT, and approved April 20, 2004 (received for review February 26, 2004) The euryarchaea Picrophilus torridus and Picrophilus oshimae are (4–6). After analysis of a number of archaeal and bacterial ge- able to grow around pH 0 at up to 65°C, thus they represent the nomes, it has been argued that microorganisms that live together most thermoacidophilic organisms known. Several features that swap genes at a higher frequency (7, 8). With the genome sequence may contribute to the thermoacidophilic survival strategy of P. of P. torridus, five complete genomes of thermoacidophilic organ- torridus were deduced from analysis of its 1.55-megabase genome. isms are available, which allows a more complex investigation of the P. torridus has the smallest genome among nonparasitic aerobic evolution of organisms sharing the extreme growth conditions of a microorganisms growing on organic substrates and simulta- unique niche in the light of horizontal gene transfer. neously the highest coding density among thermoacidophiles. An exceptionally high ratio of secondary over ATP-consuming primary Methods transport systems demonstrates that the high proton concentra- Sequencing Strategy. -

Variations in the Two Last Steps of the Purine Biosynthetic Pathway in Prokaryotes
GBE Different Ways of Doing the Same: Variations in the Two Last Steps of the Purine Biosynthetic Pathway in Prokaryotes Dennifier Costa Brandao~ Cruz1, Lenon Lima Santana1, Alexandre Siqueira Guedes2, Jorge Teodoro de Souza3,*, and Phellippe Arthur Santos Marbach1,* 1CCAAB, Biological Sciences, Recoˆ ncavo da Bahia Federal University, Cruz das Almas, Bahia, Brazil 2Agronomy School, Federal University of Goias, Goiania,^ Goias, Brazil 3 Department of Phytopathology, Federal University of Lavras, Minas Gerais, Brazil Downloaded from https://academic.oup.com/gbe/article/11/4/1235/5345563 by guest on 27 September 2021 *Corresponding authors: E-mails: [email protected]fla.br; [email protected]. Accepted: February 16, 2019 Abstract The last two steps of the purine biosynthetic pathway may be catalyzed by different enzymes in prokaryotes. The genes that encode these enzymes include homologs of purH, purP, purO and those encoding the AICARFT and IMPCH domains of PurH, here named purV and purJ, respectively. In Bacteria, these reactions are mainly catalyzed by the domains AICARFT and IMPCH of PurH. In Archaea, these reactions may be carried out by PurH and also by PurP and PurO, both considered signatures of this domain and analogous to the AICARFT and IMPCH domains of PurH, respectively. These genes were searched for in 1,403 completely sequenced prokaryotic genomes publicly available. Our analyses revealed taxonomic patterns for the distribution of these genes and anticorrelations in their occurrence. The analyses of bacterial genomes revealed the existence of genes coding for PurV, PurJ, and PurO, which may no longer be considered signatures of the domain Archaea. Although highly divergent, the PurOs of Archaea and Bacteria show a high level of conservation in the amino acids of the active sites of the protein, allowing us to infer that these enzymes are analogs. -

Methanocaldococcus Jannaschii Mja Methanococci Methanococcales
Table S2 List of the archaeal species used for evaluation of physiological and metabolic potential by MAPLE system Species Species ID Class Order [Euryarchaeota: 53 species] Methanocaldococcus jannaschii mja Methanococci Methanococcales Methanotorris igneus mig Methanococci Methanococcales Methanococcus vannielii mvn Methanococci Methanococcales Methanothermococcus okinawensis mok Methanococci Methanococcales Methanosarcina barkeri mba Methanomicrobia Methanosarcinales Methanococcoides burtonii mbu Methanomicrobia Methanosarcinales Methanohalophilus mahii mmh Methanomicrobia Methanosarcinales Methanohalobium evestigatum mev Methanomicrobia Methanosarcinales Methanosalsum zhilinae mzh Methanomicrobia Methanosarcinales Methanolobus psychrophilus mpy Methanomicrobia Methanosarcinales Methanomethylovorans hollandica mhz Methanomicrobia Methanosarcinales Methanosaeta concilii mcj Methanomicrobia Methanosarcinales Methanospirillum hungatei mhu Methanomicrobia Methanomicrobiales Methanocorpusculum labreanum mla Methanomicrobia Methanomicrobiales Methanoculleus bourgensis mbg Methanomicrobia Methanomicrobiales Methanoplanus petrolearius mpi Methanomicrobia Methanomicrobiales Methanoregula boonei mbn Methanomicrobia Methanomicrobiales Candidatus Methanosphaerula palustris mpl Methanomicrobia Methanomicrobiales Methanocella paludicola mpd Methanomicrobia Methanocellales Methanomassiliicoccus sp. Mx1-Issoire mer Methanomicrobia Unclassified Methanothermobacter thermautotrophicus mth Methanobacteria Methanobacteriales Methanosphaera stadtmanae mst