Manuscript Submission AAM
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A New Flagellated Dispersion Stage in Paraphysoderma Sedebokerense, a Pathogen of Haematococcus Pluvialis
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Springer - Publisher Connector J Appl Phycol (2016) 28:1553–1558 DOI 10.1007/s10811-015-0700-8 A new flagellated dispersion stage in Paraphysoderma sedebokerense, a pathogen of Haematococcus pluvialis Martina Strittmatter1 & Tiago Guerra 2 & Joana Silva2 & Claire M. M. Gachon1 Received: 31 July 2015 /Revised and accepted: 24 August 2015 /Published online: 18 October 2015 # The Author(s) 2015. This article is published with open access at Springerlink.com Abstract The blastocladialean fungus Paraphysoderma environmental conditions. They are therefore a prime target sedebokerense Boussiba, Zarka and James is a devastating for the development of disease management protocols in in- pathogen of the commercially valuable green microalga dustrial cultivation facilities, a goal which requires a detailed Haematococcus pluvialis, a natural source of the carotenoid understanding of their physiology. pigment astaxanthin. First identified in commercial Haematococcus cultivation facilities, P. sedebokerense is Keywords Pathogen . Green alga . Blastocladiales . hypothesised to have a complex life cycle that switches be- Zoospores . Amoebae . Flagellum . Life cycle tween a vegetative and a resting phase depending on favourable or unfavourable growth conditions. Rather unusu- ally for blastocladialean fungi, P. sedebokerense was de- scribed as lacking flagellated zoospores and only propagating Introduction via aplanosporic amoeboid cells. However, during repeated -

Index to Cecidology up to Vol. 31 (2016)
Index to Cecidology Up to Vol. 31 (2016) This index has been based on the contents of the papers rather than on their actual titles in order to facilitate the finding of papers on particular subjects. The figures following each entry are the year of publication, the volume and, in brackets, the number of the relevant issue. Aberbargoed Grasslands: report of 2011 field meeting 2012 27 (1) Aberrant Plantains 99 14(2) Acacia species galled by Fungi in India 2014 29(2) Acer gall mites (with illustrations) 2013 28(1) Acer galls: felt galls re-visited 2005 20(2) Acer saccharinum – possibly galled by Dasineura aceris new to Britain 2017 32(1) Acer seed midge 2009 24(1) Aceria anceps new to Ireland 2005 20 (1) Aceria geranii from North Wales 1999 14(2) Aceria heteronyx galling twigs of Norway Maple 2014 29(1) Aceria ilicis (gall mite) galling holm oak flowers in Brittany 1997 12(1) In Ireland 2010 25(1) Aceria mites on sycamore 2005 20(2) Aceria populi galling aspen in Scotland 2000 15(2) Aceria pterocaryae new to the British mite fauna 2008 23(2) Aceria rhodiolae galling roseroot 2013 28(1): 2016 31(1) Aceria rhodiolae in West Sutherland 2014 29(1) Aceria tristriata on Walnut 2007 22(2) Acericecis campestre sp. nov. on Field Maple 2004 19(2) Achillea ptarmica (sneezewort) galled by Macrosiphoniella millefolii 1993 8(2) Acorn galls on red oak 2014 29(1) Acorn stalks: peculiar elongation 2002 17(2) Aculops fuchsiae – a fuchsia-galling mite new to Britain 2008 23 (1) Aculus magnirostris new to Ireland 2005 20 (1) Acumyia acericola – the Acer seed -

Clade (Kingdom Fungi, Phylum Chytridiomycota)
TAXONOMIC STATUS OF GENERA IN THE “NOWAKOWSKIELLA” CLADE (KINGDOM FUNGI, PHYLUM CHYTRIDIOMYCOTA): PHYLOGENETIC ANALYSIS OF MOLECULAR CHARACTERS WITH A REVIEW OF DESCRIBED SPECIES by SHARON ELIZABETH MOZLEY (Under the Direction of David Porter) ABSTRACT Chytrid fungi represent the earliest group of fungi to have emerged within the Kingdom Fungi. Unfortunately despite the importance of chytrids to understanding fungal evolution, the systematics of the group is in disarray and in desperate need of revision. Funding by the NSF PEET program has provided an opportunity to revise the systematics of chytrid fungi with an initial focus on four specific clades in the order Chytridiales. The “Nowakowskiella” clade was chosen as a test group for comparing molecular methods of phylogenetic reconstruction with the more traditional morphological and developmental character system used for classification in determining generic limits for chytrid genera. Portions of the 18S and 28S nrDNA genes were sequenced for isolates identified to genus level based on morphology to seven genera in the “Nowakowskiella” clade: Allochytridium, Catenochytridium, Cladochytrium, Endochytrium, Nephrochytrium, Nowakowskiella, and Septochytrium. Bayesian, parsimony, and maximum likelihood methods of phylogenetic inference were used to produce trees based on one (18S or 28S alone) and two-gene datasets in order to see if there would be a difference depending on which optimality criterion was used and the number of genes included. In addition to the molecular analysis, taxonomic summaries of all seven genera covering all validly published species with a listing of synonyms and questionable species is provided to give a better idea of what has been described and the morphological and developmental characters used to circumscribe each genus. -

Atti Del Museo Di Storia Di Trieste
ATTI DEL MUSEOMUSEO CCIVICOIVICO DIDI STORSTORIAIA NATURALENATURALE DIDI TRTRIESTEIESTE naturale A I stor I D CO I V I C MUSEO DEL DEL I ATT TRIESTE 2014 VOVOL.L. 59 56 - -2018 2013 ATTI DEL MUSEO CIVICO DI STORIA NATURALE ATTI DI TRIESTE DEL MUSEO CIVICO DI STORIA NATURALE DI TRIESTE VOL. 59 - 2018 ISSN: 0335-1576 DIRIGENTE DIRETTORE RESPONSABILE DEL PERIODICO Laura Carlini Fanfogna COMITATO SCIENTIFICO Deborah Arbulla, paleontologia Pietro Brandmayr, entomologia Nicola Bressi, zoologia TRIESTEAndrea 2014 Colla, entomologia VOL. 56 - 2013 Guido Ferilli, botanica Pier Luigi Nimis, botanica REDAZIONE Livio Fogar con Gianni Pistrini Museo Civico di Storia Naturale via Tominz, 4 – 34139 Trieste – Italia Tel.: +39406758227/662 – Fax: +390406758230 E-mail: [email protected]; [email protected] www.retecivica.trieste.it/triestecultura/musei In copertina: Podarcis muralis ♂ Tolmezzo (foto di Gianluca Rassati) On the cover: Podarcis muralis ♂ Tolmezzo (photo of Gianluca Rassati) Finito di stampare nel mese di dicembre 2018 da Lithostampa ISSN: 0335-1576 ATTI DEL MUSEO CIVICO DI STORIA NATURALE DI TRIESTE VOL. 59 - 2018 TRIESTE 2018 Atti Mus. Civ. St. Nat. Trieste 59 2018 5/20 XII 2018 ISSN: 0335-1576 LA VISIONE DEI MINERALI DEL MUSEO CIVICO DI STORIA NATURALE DI TRIESTE ENRICO FRANGIPANI Via dei Fabbri, n. 1, 34124 Trieste – E-mail: [email protected] Abstract – The vision of the minerals of Trieste Natural History Museum. Among the many objectives of an exhi- bition, there is not only the valorization of the specimen but also the possibility of finding correlations among different disciplines and thus contribute to a dissemination of solid scientific knowledge. -

An Assessment of Invasive Species Management in Idaho
University of Idaho Archives, University of Idaho, www.invasive.org Robert L. Johnson, Cornell University, www.invasive.org Preparing to Meet the Challenge An Assessment of Eric Coombs, Oregan Department of Agriculture, www.invasive.org Invasive Species Management in Idaho Prepared for: The Idaho Invasive Species Council With the Support of: Kenneth R. Law, The Idaho Department of Agriculture USDA APHIS PPQ, www.invasive.org and The Nature Conservancy Northwest Natural Resource Group, LLC Joe Hinson, ©2003 i Table of Contents Foreword....................................................................................................................... iii Acknowledgements .........................................................................................................v Executive Summary...................................................................................................... vii Introduction.....................................................................................................................1 The Concept of ÒInvasivenessÓ ....................................................................................1 Why Should IdahoÕs Citizens Worry About Invasive Species?.....................................4 Facing Up to the Challenge..............................................................................................7 What Level of Commitment is Needed?.....................................................................14 Conclusions ...............................................................................................................18 -

Objective Plant Pathology
See discussions, stats, and author profiles for this publication at: https://www.researchgate.net/publication/305442822 Objective plant pathology Book · July 2013 CITATIONS READS 0 34,711 3 authors: Surendra Nath M. Gurivi Reddy Tamil Nadu Agricultural University Acharya N G Ranga Agricultural University 5 PUBLICATIONS 2 CITATIONS 15 PUBLICATIONS 11 CITATIONS SEE PROFILE SEE PROFILE Prabhukarthikeyan S. R ICAR - National Rice Research Institute, Cuttack 48 PUBLICATIONS 108 CITATIONS SEE PROFILE Some of the authors of this publication are also working on these related projects: Management of rice diseases View project Identification and characterization of phytoplasma View project All content following this page was uploaded by Surendra Nath on 20 July 2016. The user has requested enhancement of the downloaded file. Objective Plant Pathology (A competitive examination guide)- As per Indian examination pattern M. Gurivi Reddy, M.Sc. (Plant Pathology), TNAU, Coimbatore S.R. Prabhukarthikeyan, M.Sc (Plant Pathology), TNAU, Coimbatore R. Surendranath, M. Sc (Horticulture), TNAU, Coimbatore INDIA A.E. Publications No. 10. Sundaram Street-1, P.N.Pudur, Coimbatore-641003 2013 First Edition: 2013 © Reserved with authors, 2013 ISBN: 978-81972-22-9 Price: Rs. 120/- PREFACE The so called book Objective Plant Pathology is compiled by collecting and digesting the pertinent information published in various books and review papers to assist graduate and postgraduate students for various competitive examinations like JRF, NET, ARS conducted by ICAR. It is mainly helpful for students for getting an in-depth knowledge in plant pathology. The book combines the basic concepts and terminology in Mycology, Bacteriology, Virology and other applied aspects. -

Zjgbrlwms.Pdf
目录 前言:史前真菌 Part 1:各路“英雄” “二战”英雄 产黄青霉 器官移植技术大跃进 多孔木霉 石磨水洗牛仔裤 里氏霉菌 生物农药的先驱 球孢白僵菌 农夫小帮手 绿色木霉 Part 2:农业“杀手” 美国的灾难 禾生炭疽病菌 不再随风摇曳的麦田 禾谷镰刀菌、禾生球腔菌、禾柄锈菌与小麦德氏霉 香蕉王国的没落 尖孢镰刀菌与香蕉黑条叶斑病菌 消失的咖啡帝国 咖啡驼孢锈菌 鬼火燃烧的稻田 稻瘟病菌 最后一颗巧克力 可可丛枝病菌、可可链疫孢荚腐病菌 饥饿爱尔兰 致病疫霉 Part 3:迷幻“嬉皮” 超级玛丽的能量 毒蝇伞 欧洲历史急转弯 毒鹅膏 巫师的黑暗咒语 麦角菌 与神明共舞 光盖伞 Part 4:真菌简史 众神的食物 洋菇 菇中鲍鱼 金针菇、平菇与杏鲍菇 南北货三兄弟 香菇、木耳与竹荪 父不传子的秘密 松茸 厨房里的钻石 松露 美人腿 菰黑穗菌 天神的肠子 冬虫夏草 中国草药之王 灵芝 狄俄尼索斯的魔法 酿酒酵母菌 酒中之王,王者之酒 灰霉菌 日本国菌 米曲霉 红色的滋味 红曲霉 饮料工业的二次革命 黑曲霉与解脂假丝酵母 臭的艺术 腐乳毛霉与米黑毛霉菌 奶酪之王 洛克福特青霉 新时代健康卫士 汉逊德巴利酵母 Part 5:生态“黑客” 寂静的春天 蛙壶菌 蝙蝠杀手 地霉锈腐菌 消失的蜜蜂群 东方蜜蜂微孢子虫 栗树浩劫 木层孔褐根腐病菌与寄生隐丛赤壳菌 后记:真菌告诉我们的事 真菌Q&A Beckoning the Wind , Stories of Mushroom Summoning the Rain. 一缕菌丝 搅动6 000年人类文明史 从餐桌、工厂、实验室、战场到农田 那些人类迷恋、依赖或惧怕的真菌 与它们的秘密生活 前言:史前真菌 现在,蘑菇已经是非常常见的食材,不过真菌的考古信息非常贫 乏,原因是它们没有较硬的结构,而且很容易腐烂,因此很难产生化 石,在考古遗址中也就很少被发现。即使在考古遗址中发现与真菌有关 的证据,也多是在文献和壁画中,或是在生活器具上发现的真菌图案。 最古老的陆生生物 剑桥大学的科学家,在英国的内赫布里底群岛(inner Hebridean island)及瑞典的哥得兰岛(Gotland island)上,发现了一种比人类头 发丝还细的神秘化石。科学家推断,在这一化石存在的年代,几乎所有 生物都还在大海里,即使陆地上存在生物,构造也不会比苔藓来得更复 杂,而且在那时,就连地衣都还没有经过演化出现在陆地上。这是马丁 ·史密斯(Martin Smith)博士在 2016年发表的论文中谈到的,是迄今发 现的最古老的陆生生物的证据。 由史密斯博士发现的化石,之后被确认为一种真菌,被命名为“古 怪管状真菌”(tortotubus)。这个研究也说明了,真菌可能是地球上第 一个从海里登上陆地的复杂有机体。这种微小的“古怪管状真菌”出现在 距今约4.4亿年,是迄今为止发现的最古老的陆生生物化石。这也进一 步说明,真菌的率先登陆为后来的生物提供了富饶的土壤,让其他植物 得以上岸生长,并以此吸引动物从海中迁移到陆地。 最大的生物 4.2亿年前到3.5亿年前,陆生植物从海洋登上陆地不久,最高的陆 地植物还不到1米。那时候,却有一种生物可以长到8米高,1米宽。这 种1843年由加拿大科学家发现、出土于阿拉伯的生物化石,最初被认为 是古老的大树化石。因此,它被命名为“原杉藻属”(Prototaxites)。即 使我们现在已经知道它不是树,但依照命名法则,这个名称还是要必须 使用,不能更改。后来经过漫长的争论,有一派科学家认为这个“巨无 霸”是真菌,另一派则认为它是早期的蕨类。虽然 2007 年经过同位素的 测定,几乎已经可以确定它是真菌,但至今依然没有得出令所有人满意 -
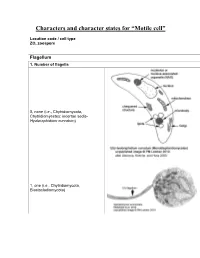
Characters and Character States for “Motile Cell”
Characters and character states for “Motile cell” Location code / cell type ZO, zoospore Flagellum 1. Number of flagella 0, none (i.e., Chytridiomycota, Chytridiomycetes: incertae sedis- Hyaloraphidium curvatum) 1, one (i.e., Chytridiomycota, Blastocladiomycota) 2, multiple (i.e., Neocallimastigomycota) 2. Electron-opaque plug in axoneme core and between axoneme and flagellar membrane 0, absent 1, present (i.e., Chytridiomycota: Chytridiomycetes: Chytridiales, Lobulomycetales, Cladochytriales, incertae sedis: Synchytrium endobioticum, Polychytrium clade; Monoblepharidomycetes) 3. Flagellum coating 0, absent (all taxa except Polyphagus euglenae) 1, present (Polyphagus euglenae) Kinetosome (the term “basal body” is synonymous; Andersen et al. 1991) 4. Electron-opaque core in kinetosome 0, absent (all taxa except Kappamyces) 1, present (Chytridiomycota: Chytridiomycetes: Rhizophydiales- Kappamyces) 5. Scalloped ring within kinetosome, extensions of the A, B, or C microtubule 0, absent 1, present (Lacustromyces hiemalis) Kinetosome-associated structures 6. Kinetosome support 0, absent (Thalassochytrium gracilaripsidis) 1, kinetosome props 2, broken kinetosome props (Olpidium radicale) 3, skirt-like structure surrounding kinetosome (Neocallimastigomycota) 7. Kinetosome-associated plates 0, absent 1, present (Chytridiomycota: Chytridiomycetes: Chytridiales- Group I- type zoospore [Barr 1980]) 8. Kinetosome-associated saddle 0, absent 1, present (Chytridiomycota: Chytridiomycetes: Chytridiales- Group II-type zoospore [Barr 1980]- Chytridium -

Synchytrium Shuteriae and Other Doubtful Species
©Verlag Ferdinand Berger & Söhne Ges.m.b.H., Horn, Austria, download unter www.biologiezentrum.at Synchytrium shuteriae and other doubtful Species. •./,-f mfii.-'t, i. • • By John S. Karling, . ,;.:. j„:<:'i, ,,,^^\ Department of Biological Sciences, Purdue University, Lafayette, f; Indiana, USA. ;• "'•• •••'-• '•-" ; With 9 Textfig. • • ' ' ' f This study has been supported by a grant from the National Science Foundation* i < ••••• •••--'•_ Synchytrium shuteriae was described by II e n n i n g* (1895) as a parasite of Shuteria africana which G. V o 1 k e n s collected near Kilimandscharo, in German East Africa, and he diagnosed it as follows: "maculis pallidis rotundatis, tuberculis amphigenis sparsis vel confluentibus, subhemisphaericis applanatis, dein irregulariter rugosis, flavescenti-viridulis dein subfuscescentibus, 0.3—1 mm dia- metro, plerumque e pluribus cellulis perdurantibus formatis; sporis globosis vel subellipsoideis, saepe acutangulis, flavosubfuscescen- tibus 10—15 # 9—13 ^, episporio levi, 1—1.5 |x crasso, subhyalino." In 1912 T o b 1 e r studied specimens of this species in the Berlin herbarium and came to the conclusion that it may not be a valid species of Synchyirium, or that more than one fungus is present on the host. Recently, through the generosity of Dr. Sten A h 1 n e r, the author received material of this species from the Riksmuseet in Stock- holm, Sweden, which was collected by G. V o 1 k e n s near Kilimand- scharo, June 13, 1894, and a study of fixed and stained sections of this material has shown that S. shuteriae is a true Synchytrium species which is very similar to members of the subgenus Woro- ninella. -

Aquatic Fungi and Straminipilous Organisms on Decomposing Fragments of Wetland Plants
MYCOLOGIA BALCANICA 4: 31–44 (2007) 31 Aquatic fungi and straminipilous organisms on decomposing fragments of wetland plants Bazyli Czeczuga *, Elżbieta Muszyńska, Anna Godlewska & Bożenna Mazalska Department of General Biology, Medical University, Kilińskiego 1, 15-089 Białystok, Poland Received 2 January 2007 / Accepted 16 April 2007 Abstract. Straminipilous organisms and fungus species on dead fragments of 25 taxa of wetland plants from three water bodies were investigated. A total of 219 fungal taxa, including 85 straminipilous organisms and 134 fungus species were recorded. Th e largest number of straminipilous organisms and fungus species occurred on fragments of Lathyrus palustre and Rorippa amphibia (45 and 44 species respectively), the fewest on Calla palustris (20). Th e most common species were Chytridium xylophilum (on 23 plants), Pythium rostratum (on 15), Pithomyces obscuriseptatus (on all plants), Trinacrium subtile (on 17) and Titaea (Tetracladium) maxilliformis (on 16). Th e larger numbers of species were recorded in running water (Jaroszówka Spring, Supraśl River, 124 and 122 respectively), with fewest (106) in stagnant water at Dojlidy Pond. Th irty-one taxa were recorded for the fi rst time from Poland. Key words: aquatic fungi, hydrochemistry, Poland, straminipilous organisms, wetland plants Introduction many years (Czeczuga et al. 2002, 2003a, b, 2004; Czeczuga 2004). Th erefore, we decided to establish the species diversity Th e border between land and water reservoirs of diff erent types of straminipilous and aquatic fungi which participate in the is frequently overgrown with herbaceous plants which mostly mineralization of some wetland plants in diff erent freshwater die in autumn. Th is relatively large mass of organic matter is ecosystems. -

This Article Was Published in an Elsevier Journal. the Attached Copy Is Furnished to the Author for Non-Commercial Research
This article was published in an Elsevier journal. The attached copy is furnished to the author for non-commercial research and education use, including for instruction at the author’s institution, sharing with colleagues and providing to institution administration. Other uses, including reproduction and distribution, or selling or licensing copies, or posting to personal, institutional or third party websites are prohibited. In most cases authors are permitted to post their version of the article (e.g. in Word or Tex form) to their personal website or institutional repository. Authors requiring further information regarding Elsevier’s archiving and manuscript policies are encouraged to visit: http://www.elsevier.com/copyright Author's personal copy mycological research 112 (2008) 70–81 journal homepage: www.elsevier.com/locate/mycres Isolation and characterization of a novel chytrid species (phylum Blastocladiomycota), parasitic on the green alga Haematococcus Yoram HOFFMANa, Claude AFLALOa,*, Aliza ZARKAa, Jenia GUTMANa, Timothy Y. JAMESb,y, Sammy BOUSSIBAa aMicroalgal Biotechnology Laboratory, the Jacob Blaustein Institutes for Desert Research, Ben Gurion University of the Negev, Sede-Boker Campus 84990, Israel bDepartment of Biology, Duke University, Durham, NC 27708, USA article info abstract Article history: A parasite was found in cultures of the green microalga Haematococcus pluvialis that grew Received 21 February 2007 epibiotically on algal cells and caused epidemics resulting in damage to the host cultures. Received in revised -
Checklist of Microfungi on Grasses in Thailand (Excluding Bambusicolous Fungi)
Asian Journal of Mycology 1(1): 88–105 (2018) ISSN 2651-1339 www.asianjournalofmycology.org Article Doi 10.5943/ajom/1/1/7 Checklist of microfungi on grasses in Thailand (excluding bambusicolous fungi) Goonasekara ID1,2,3, Jayawardene RS1,2, Saichana N3, Hyde KD1,2,3,4 1 Center of Excellence in Fungal Research, Mae Fah Luang University, Chiang Rai 57100, Thailand 2 School of Science, Mae Fah Luang University, Chiang Rai 57100, Thailand 3 Key Laboratory for Plant Biodiversity and Biogeography of East Asia (KLPB), Kunming Institute of Botany, Chinese Academy of Science, Kunming 650201, Yunnan, China 4 World Agroforestry Centre, East and Central Asia, 132 Lanhei Road, Kunming 650201, Yunnan, China Goonasekara ID, Jayawardene RS, Saichana N, Hyde KD 2018 – Checklist of microfungi on grasses in Thailand (excluding bambusicolous fungi). Asian Journal of Mycology 1(1), 88–105, Doi 10.5943/ajom/1/1/7 Abstract An updated checklist of microfungi, excluding bambusicolous fungi, recorded on grasses from Thailand is provided. The host plant(s) from which the fungi were recorded in Thailand is given. Those species for which molecular data is available is indicated. In total, 172 species and 35 unidentified taxa have been recorded. They belong to the main taxonomic groups Ascomycota: 98 species and 28 unidentified, in 15 orders, 37 families and 68 genera; Basidiomycota: 73 species and 7 unidentified, in 8 orders, 8 families and 18 genera; and Chytridiomycota: one identified species in Physodermatales, Physodermataceae. Key words – Ascomycota – Basidiomycota – Chytridiomycota – Poaceae – molecular data Introduction Grasses constitute the plant family Poaceae (formerly Gramineae), which includes over 10,000 species of herbaceous annuals, biennials or perennial flowering plants commonly known as true grains, pasture grasses, sugar cane and bamboo (Watson 1990, Kellogg 2001, Sharp & Simon 2002, Encyclopedia of Life 2018).