GNGT2 (NM 031498) Human Mass Spec Standard Product Data
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

G Protein-Coupled Receptors
G PROTEIN-COUPLED RECEPTORS Overview:- The completion of the Human Genome Project allowed the identification of a large family of proteins with a common motif of seven groups of 20-24 hydrophobic amino acids arranged as α-helices. Approximately 800 of these seven transmembrane (7TM) receptors have been identified of which over 300 are non-olfactory receptors (see Frederikson et al., 2003; Lagerstrom and Schioth, 2008). Subdivision on the basis of sequence homology allows the definition of rhodopsin, secretin, adhesion, glutamate and Frizzled receptor families. NC-IUPHAR recognizes Classes A, B, and C, which equate to the rhodopsin, secretin, and glutamate receptor families. The nomenclature of 7TM receptors is commonly used interchangeably with G protein-coupled receptors (GPCR), although the former nomenclature recognises signalling of 7TM receptors through pathways not involving G proteins. For example, adiponectin and membrane progestin receptors have some sequence homology to 7TM receptors but signal independently of G-proteins and appear to reside in membranes in an inverted fashion compared to conventional GPCR. Additionally, the NPR-C natriuretic peptide receptor has a single transmembrane domain structure, but appears to couple to G proteins to generate cellular responses. The 300+ non-olfactory GPCR are the targets for the majority of drugs in clinical usage (Overington et al., 2006), although only a minority of these receptors are exploited therapeutically. Signalling through GPCR is enacted by the activation of heterotrimeric GTP-binding proteins (G proteins), made up of α, β and γ subunits, where the α and βγ subunits are responsible for signalling. The α subunit (tabulated below) allows definition of one series of signalling cascades and allows grouping of GPCRs to suggest common cellular, tissue and behavioural responses. -

Multi-Functionality of Proteins Involved in GPCR and G Protein Signaling: Making Sense of Structure–Function Continuum with In
Cellular and Molecular Life Sciences (2019) 76:4461–4492 https://doi.org/10.1007/s00018-019-03276-1 Cellular andMolecular Life Sciences REVIEW Multi‑functionality of proteins involved in GPCR and G protein signaling: making sense of structure–function continuum with intrinsic disorder‑based proteoforms Alexander V. Fonin1 · April L. Darling2 · Irina M. Kuznetsova1 · Konstantin K. Turoverov1,3 · Vladimir N. Uversky2,4 Received: 5 August 2019 / Revised: 5 August 2019 / Accepted: 12 August 2019 / Published online: 19 August 2019 © Springer Nature Switzerland AG 2019 Abstract GPCR–G protein signaling system recognizes a multitude of extracellular ligands and triggers a variety of intracellular signal- ing cascades in response. In humans, this system includes more than 800 various GPCRs and a large set of heterotrimeric G proteins. Complexity of this system goes far beyond a multitude of pair-wise ligand–GPCR and GPCR–G protein interactions. In fact, one GPCR can recognize more than one extracellular signal and interact with more than one G protein. Furthermore, one ligand can activate more than one GPCR, and multiple GPCRs can couple to the same G protein. This defnes an intricate multifunctionality of this important signaling system. Here, we show that the multifunctionality of GPCR–G protein system represents an illustrative example of the protein structure–function continuum, where structures of the involved proteins represent a complex mosaic of diferently folded regions (foldons, non-foldons, unfoldons, semi-foldons, and inducible foldons). The functionality of resulting highly dynamic conformational ensembles is fne-tuned by various post-translational modifcations and alternative splicing, and such ensembles can undergo dramatic changes at interaction with their specifc partners. -

A System-Level, Molecular Evolutionary Analysis of Mam- Malian Phototransduction (Supplementary Material)
A system-level, molecular evolutionary analysis of mam- malian phototransduction (supplementary material) Brandon M Invergo1 , Ludovica Montanucci∗1 , Hafid Laayouni1 and Jaume Bertranpetit1 1IBE-Institute of Evolutionary Biology (UPF-CSIC), CEXS-UPF-PRBB, Barcelona, Catalonia, Spain Email: Brandon Invergo - [email protected]; Ludovica Montanucci∗- [email protected]; Hafid Laayouni - hafi[email protected]; Jaume Bertranpetit - [email protected]; ∗Corresponding author Table S1 - Classifications of the genes Genes were assigned classifications according to their photoreceptor cell-type specificity, the process in which the encoded protein is primarily active, and the general function of the encoded protein. (Note: here "enzyme" specifically refers to enzymes involved in retinoid recycling.) 1 gene cell type process function ABCA4 shared retinoid cycle enzyme AIPL1 shared phototransduction other ARR3 cone phototransduction signal regulator ASCL1 rod development transcription regulation CNGA1 rod phototransduction ion channel CNGA3 cone phototransduction ion channel CNGB1 rod phototransduction ion channel CNGB3 cone phototransduction ion channel CRX shared development transcription regulation GNAT1 rod phototransduction G protein GNAT2 cone phototransduction G protein GNB1 rod phototransduction G protein GNB3 cone phototransduction G protein GNB5 shared phototransduction G protein GNGT1 rod phototransduction G protein GNGT2 cone phototransduction G protein GPSM2 shared phototransduction other GRK1 shared phototransduction -

Network Pharmacology of JAK Inhibitors
Network pharmacology of JAK inhibitors Devapregasan Moodleya, Hideyuki Yoshidaa, Sara Mostafavib,c, Natasha Asinovskia, Adriana Ortiz-Lopeza, Peter Symanowiczd, Jean-Baptiste Telliezd, Martin Hegend, James D. Clarkd, Diane Mathisa,1, and Christophe Benoista,1 aDivision of Immunology, Department of Microbiology and Immunobiology, Harvard Medical School, Boston, MA 02115; bDepartment of Statistics, University of British Columbia, Vancouver, BC, Canada V6T 1Z4; cDepartment of Medical Genetics, University of British Columbia, Vancouver, BC, Canada V6T 1Z4; and dInflammation and Immunology, Pfizer, Cambridge, MA 02139 Contributed by Christophe Benoist, June 24, 2016 (sent for review April 27, 2016; reviewed by Tadatsugu Taniguchi and Arthur Weiss) Small-molecule inhibitors of the Janus kinase family (JAKis) are side effects that likely reflect cytokine blockade, such as bacterial clinically efficacious in multiple autoimmune diseases, albeit with and fungal infections, in particular, (re)activation of the varicella increased risk of certain infections. Their precise mechanism of action zoster virus, and at high doses, anemia and thrombocytopenia (2, 3). is unclear, with JAKs being signaling hubs for several cytokines. We A new JAKi generation targets single JAK isoforms, which might assessed the in vivo impact of pan- and isoform-specific JAKi in mice improve adverse events by restricting the range of activity. Efficacy by immunologic and genomic profiling. Effects were broad across has been observed with JAK1-selective compounds (7) and com- the immunogenomic network, with overlap between inhibitors. Nat- pounds of reported specificity for JAK3 (ref. 8, but see ref. 2). ural killer (NK) cell and macrophage homeostasis were most imme- However, the premise of substantially improved in vivo specificity diately perturbed, with network-level analysis revealing a rewiring remains unproven, because the impact of JAKi compounds on the of coregulated modules of NK cell transcripts. -

Anti-GNGT2 Antibody (ARG58880)
Product datasheet [email protected] ARG58880 Package: 100 μl anti-GNGT2 antibody Store at: -20°C Summary Product Description Rabbit Polyclonal antibody recognizes GNGT2 Tested Reactivity Ms Tested Application WB Host Rabbit Clonality Polyclonal Isotype IgG Target Name GNGT2 Antigen Species Human Immunogen Recombinant fusion protein corresponding to aa. 1-69 of Human GNGT2 (NP_113686.1). Conjugation Un-conjugated Alternate Names GNG8; GNG9; GNGT8; G-GAMMA-8; G-GAMMA-C; Guanine nucleotide-binding protein G(I)/G(S)/G(O) subunit gamma-T2; G gamma-C; G-gamma-8; G-gamma-9; Guanine nucleotide binding protein gamma transducing activity polypeptide 2 Application Instructions Application table Application Dilution WB 1:500 - 1:2000 Application Note * The dilutions indicate recommended starting dilutions and the optimal dilutions or concentrations should be determined by the scientist. Calculated Mw 8 kDa Properties Form Liquid Purification Affinity purified. Buffer PBS (pH 7.3), 0.02% Sodium azide and 50% Glycerol. Preservative 0.02% Sodium azide Stabilizer 50% Glycerol Storage instruction For continuous use, store undiluted antibody at 2-8°C for up to a week. For long-term storage, aliquot and store at -20°C. Storage in frost free freezers is not recommended. Avoid repeated freeze/thaw cycles. Suggest spin the vial prior to opening. The antibody solution should be gently mixed before use. Note For laboratory research only, not for drug, diagnostic or other use. www.arigobio.com 1/2 Bioinformation Gene Symbol GNGT2 Gene Full Name guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 2 Background Phototransduction in rod and cone photoreceptors is regulated by groups of signaling proteins. -

And Diabetes-Associated Pancreatic Cancer: a GWAS Data Analysis
Published OnlineFirst October 17, 2013; DOI: 10.1158/1055-9965.EPI-13-0437-T Cancer Epidemiology, Research Article Biomarkers & Prevention Genes–Environment Interactions in Obesity- and Diabetes-Associated Pancreatic Cancer: A GWAS Data Analysis Hongwei Tang1, Peng Wei3, Eric J. Duell4, Harvey A. Risch5, Sara H. Olson6, H. Bas Bueno-de-Mesquita7, Steven Gallinger9, Elizabeth A. Holly10, Gloria M. Petersen11, Paige M. Bracci10, Robert R. McWilliams11, Mazda Jenab12, Elio Riboli16, Anne Tjønneland18, Marie Christine Boutron-Ruault13,14,15, Rudolf Kaaks19, Dimitrios Trichopoulos20,21,22, Salvatore Panico23, Malin Sund24, Petra H.M. Peeters8,16, Kay-Tee Khaw17, Christopher I. Amos2, and Donghui Li1 Abstract Background: Obesity and diabetes are potentially alterable risk factors for pancreatic cancer. Genetic factors that modify the associations of obesity and diabetes with pancreatic cancer have previously not been examined at the genome-wide level. Methods: Using genome-wide association studies (GWAS) genotype and risk factor data from the Pancreatic Cancer Case Control Consortium, we conducted a discovery study of 2,028 cases and 2,109 controls to examine gene–obesity and gene–diabetes interactions in relation to pancreatic cancer risk by using the likelihood-ratio test nested in logistic regression models and Ingenuity Pathway Analysis (IPA). Results: After adjusting for multiple comparisons, a significant interaction of the chemokine signaling À pathway with obesity (P ¼ 3.29 Â 10 6) and a near significant interaction of calcium signaling pathway with À diabetes (P ¼ 1.57 Â 10 4) in modifying the risk of pancreatic cancer were observed. These findings were supported by results from IPA analysis of the top genes with nominal interactions. -

Pulmonary Hypertension Remodels the Genomic Fabrics of Major Functional Pathways
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 21 December 2019 doi:10.20944/preprints201912.0280.v1 Peer-reviewed version available at Genes 2020, 11, 126; doi:10.3390/genes11020126 1 Article 2 Pulmonary Hypertension Remodels the Genomic 3 Fabrics of Major Functional Pathways 4 Rajamma Mathew 1,2, Jing Huang 1, Sanda Iacobas 3 and Dumitru A Iacobas 4,* 5 1 Department of Pediatrics, New York Medical College, Valhalla, NY 10595, U.S.A. 6 2 Department of Physiology, New York Medical College, Valhalla, NY 10595, U.S.A. 7 3 Department of Pathology, New York Medical College, Valhalla, NY 10595, U.S.A. 8 4 Personalized Genomics Laboratory, Center for Computational Systems Biology, Roy G Perry College of 9 Engineering, Prairie View A&M University, Prairie View, TX77446, U.S.A. 10 * Correspondence: [email protected]; Tel.: +1(936)261-9926 11 Abstract: Pulmonary hypertension (PH) is a serious disorder with high morbidity and mortality 12 rate. We analyzed the right ventricular systolic pressure (RVSP), right ventricular hypertrophy 13 (RVH), lung histology and transcriptomes of six weeks old male rats with PH induced by: 1) hypoxia 14 (HO), 2) administration of monocrotaline (CM) or 3) administration of monocrotaline and exposure 15 to hypoxia (HM). The results in PH rats were compared to those in control rats (CO). After four 16 weeks exposure, increased RVSP and RVH, pulmonary arterial wall thickening, and alteration of 17 the lung transcriptome were observed in all PH groups. The HM group exhibited the largest 18 alterations and also neointimal lesions and obliteration of lumen in small arteries. -

Canine Cone Transducin-Y Gene and Cone Degeneration in the Cd Dog
Canine Cone Transducin-y Gene and Cone Degeneration in the cd Dog Novrouz B. Akhmedov,1 Natik I. Piriev,1 Sue Pearce-Kelling,5 Gregory M. Acland,5 Gustavo D. Aguirre,5 and Debora B. Farber1'2 PURPOSE. TO characterize the cDNA and the organization of the gene encoding the cone-specific y subunit of transducin (Tyc) and to examine this gene as a candidate for the recessively inherited cone photoreceptor degeneration in the cd dog. METHODS. Canine Tyc cDNA was cloned and sequenced. Polymerase chain reaction (PCR) was used to define the Tyc gene structure, northern blot analysis to examine the level of expression of Tyc mRNA in control and cd-affected retinas, and immunocytochemistry to determine the presence and localization of Tyc in normal and cd retinas. RESULTS. Immunocytochemical results showed Tyc localized to cone photoreceptor outer segments in the normal retina, whereas no Tyc immunoreactivity was observed in the cd retinas. However, the level of transcription and the primary structure of the cloned cDNA coding for the 69 -amino acid protein were identical in retinas from wild-type and affected dogs. CONCLUSIONS. Although Tyc immunoreactivity was specifically absent in the cd dog retina, no differences were detected between normal and cd retinas in the nucleotide sequence of Tyc mRNA or in its synthesis. These results indicate that a mutation in the Tyc gene may not be causally associated with the cd dog disease. These findings suggest that possible abnormalities in posttrans- lational modification of Tyc or defective assembly of the transducin a/3y complex could lead to rapid degradation of Tyc. -
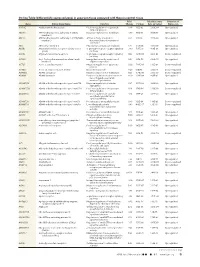
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

SUPPLEMENTARY APPENDIX Exome Sequencing Reveals Heterogeneous Clonal Dynamics in Donor Cell Myeloid Neoplasms After Stem Cell Transplantation
SUPPLEMENTARY APPENDIX Exome sequencing reveals heterogeneous clonal dynamics in donor cell myeloid neoplasms after stem cell transplantation Julia Suárez-González, 1,2 Juan Carlos Triviño, 3 Guiomar Bautista, 4 José Antonio García-Marco, 4 Ángela Figuera, 5 Antonio Balas, 6 José Luis Vicario, 6 Francisco José Ortuño, 7 Raúl Teruel, 7 José María Álamo, 8 Diego Carbonell, 2,9 Cristina Andrés-Zayas, 1,2 Nieves Dorado, 2,9 Gabriela Rodríguez-Macías, 9 Mi Kwon, 2,9 José Luis Díez-Martín, 2,9,10 Carolina Martínez-Laperche 2,9* and Ismael Buño 1,2,9,11* on behalf of the Spanish Group for Hematopoietic Transplantation (GETH) 1Genomics Unit, Gregorio Marañón General University Hospital, Gregorio Marañón Health Research Institute (IiSGM), Madrid; 2Gregorio Marañón Health Research Institute (IiSGM), Madrid; 3Sistemas Genómicos, Valencia; 4Department of Hematology, Puerta de Hierro General University Hospital, Madrid; 5Department of Hematology, La Princesa University Hospital, Madrid; 6Department of Histocompatibility, Madrid Blood Centre, Madrid; 7Department of Hematology and Medical Oncology Unit, IMIB-Arrixaca, Morales Meseguer General University Hospital, Murcia; 8Centro Inmunológico de Alicante - CIALAB, Alicante; 9Department of Hematology, Gregorio Marañón General University Hospital, Madrid; 10 Department of Medicine, School of Medicine, Com - plutense University of Madrid, Madrid and 11 Department of Cell Biology, School of Medicine, Complutense University of Madrid, Madrid, Spain *CM-L and IB contributed equally as co-senior authors. Correspondence: -

Gene Expressions for Signal Transduction Under Acidic Conditions
Genes 2013, 4, 65-85; doi:10.3390/genes4010065 OPEN ACCESS genes ISSN 2073-4425 www.mdpi.com/journal/genes Article Gene Expressions for Signal Transduction under Acidic Conditions Toshihiko Fukamachi 1, Syunsuke Ikeda 1, Xin Wang 1, Hiromi Saito 1, Masatoshi Tagawa 2 and Hiroshi Kobayashi 1,* 1 Graduate School of Pharmaceutical Sciences, Chiba University, 1-8-1, Inohana, Chuo-ku, Chiba 260-8675, Japan; E-Mails: [email protected] (T.F.); [email protected] (S.I.); [email protected] (X.W.); [email protected] (H.S.) 2 Division of Pathology and Cell Therapy, Chiba Cancer Center Research Institute, 666-2, Nitona, Chuo-ku, Chiba 260-8717, Japan; E-Mail: [email protected] * Author to whom correspondence should be addressed; E-Mail: [email protected]. Received: 10 January 2013; in revised form: 18 February 2013 / Accepted: 27 February 2013 / Published: 8 March 2013 Abstract: Although it is now well known that some diseased areas, such as cancer nests, inflammation loci, and infarction areas, are acidified, little is known about cellular signal transduction, gene expression, and cellular functions under acidic conditions. Our group showed that different signal proteins were activated under acidic conditions compared with those observed in a typical medium of around pH 7.4 that has been used until now. Investigations of gene expression under acidic conditions may be crucial to our understanding of signal transduction in acidic diseased areas. In this study, we investigated gene expression in mesothelioma cells cultured at an acidic pH using a DNA microarray technique.