G Protein-Coupled Receptors
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Original Article Upregulation of HOXA13 As a Potential Tumorigenesis and Progression Promoter of LUSC Based on Qrt-PCR and Bioinformatics
Int J Clin Exp Pathol 2017;10(10):10650-10665 www.ijcep.com /ISSN:1936-2625/IJCEP0065149 Original Article Upregulation of HOXA13 as a potential tumorigenesis and progression promoter of LUSC based on qRT-PCR and bioinformatics Rui Zhang1*, Yun Deng1*, Yu Zhang1, Gao-Qiang Zhai1, Rong-Quan He2, Xiao-Hua Hu2, Dan-Ming Wei1, Zhen-Bo Feng1, Gang Chen1 Departments of 1Pathology, 2Medical Oncology, First Affiliated Hospital of Guangxi Medical University, Nanning, Guangxi Zhuang Autonomous Region, China. *Equal contributors. Received September 7, 2017; Accepted September 29, 2017; Epub October 1, 2017; Published October 15, 2017 Abstract: In this study, we investigated the levels of homeobox A13 (HOXA13) and the mechanisms underlying the co-expressed genes of HOXA13 in lung squamous cancer (LUSC), the signaling pathways in which the co-ex- pressed genes of HOXA13 are involved and their functional roles in LUSC. The clinical significance of 23 paired LUSC tissues and adjacent non-tumor tissues were gathered. HOXA13 levels in LUSC were detected by quantita- tive real-time polymerase chain reaction (qRT-PCR). HOXA13 levels in LUSC from The Cancer Genome Atlas (TCGA) and Oncomine were analyzed. We performed receiver operator characteristic (ROC) curves of various clinicopath- ological features of LUSC. Co-expressed of HOXA13 were collected from MEM, cBioPortal and GEPIA. The func- tions and pathways of the most reliable overlapped genes were achieved from the Gene Otology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) databases, respectively. The protein-protein interaction (PPI) net- works were mapped using STRING. HOXA13 in LUSC were markedly upregulated compared with those in the non- cancerous controls as demonstrated by qRT-PCR (LUSC: 0.330±0.360; CONTROLS: 0.155±0.142; P=0.021). -

Identi®Cation and Role of Adenylyl Cyclase in Auxin Signalling in Higher
letters to nature + + + P.P. thank the Academy of Finland and the Deutsche Forschungsgemeinschaft, respectively, for ®nancial CO , 53), 77 (C6H5 , 60), 73 (TMSi , 84); 6-methyl-4-hydroxy-2-pyrone: RRt 0.35, 198 (M+, 18), 183 ([M-Me]+, 16), 170 ([M-CO]+, 54), 155 ([M-CO-Me]+, support. + + Correspondence and requests for materials should be addressed to J.S. (e-mail: [email protected] 15), 139 ([M-Me-CO2] , 10), 127 ([M-Me-2CO] , 13), 99 (12), 84 (13), 73 + + freiburg.de). (TMSi , 100), 43 (CH3CO , 55). The numbers show m/z values, and the key fragments and their relative intensities are indicated in parentheses. Received 4 August; accepted 14 October 1998. erratum 1. Helariutta, Y. et al. Chalcone synthase-like genes active during corolla development are differentially expressed and encode enzymes with different catalytic properties in Gerbera hybrida (Asteraceae). Plant Mol. Biol. 28, 47±60 (1995). 2. Helariutta, Y. et al. Duplication and functional divergence in the chalcone synthase gene family of 8 Asteraceae: evolution with substrate change and catalytic simpli®cation. Proc. Natl Acad. Sci. USA 93, Crystal structure of the complex 9033±9038 (1996). 3. Thaisrivongs, S. et al. Structure-based design of HIV protease inhibitors: 5,6-dihydro-4-hydroxy-2- of the cyclin D-dependent pyrones as effective, nonpeptidic inhibitors. J. Med. Chem. 39, 4630±4642 (1996). 4. Hagen, S. E. et al. Synthesis of 5,6-dihydro-4-hydroxy-2-pyrones as HIV-1 protease inhibitors: the kinase Cdk6 bound to the profound effect of polarity on antiviral activity. J. Med. Chem. -
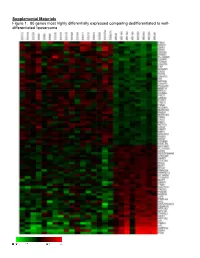
Supplemental Materials Figure 1. 80 Genes Most Highly Differentially Expressed Comparing Dedifferentiated to Well- Differentiated Liposarcoma
Supplemental Materials Figure 1. 80 genes most highly differentially expressed comparing dedifferentiated to well- differentiated liposarcoma Figure 2. Validation of microa CDC2, RACGAP1, FGFR-2, MAD2 and CITED1 200 CDK4 by Liposarcoma Subtype 16 0 12 0 80 Expression40 Level 0 Nl Fat rray data by quantitative RT-P Well Diff 16 0 p16 by Liposarcoma Subtype Dediff 12 0 80 Myxoid Expression40 Level Round Cell 0 12 0 MDM2 by Liposarcoma Subtype 10 0 Pleomorphic Nl Fat 80 60 40 Expression Level 20 RACGAP1 by LiposarcomaWell Diff Subtype 0 70 CR for genes CDK4, MDM2, p16, 60 Dediff 50 Nl Fat 40 30 Myxoid Expression20 Level Well Diff 10 50 0 Round Cell CDC2 by Liposarcoma Subtype Dediff 40 Nl Fat Pleomorphic 30 Myxoid 20 Expression Level Well Diff 10 MAD2L1 by Liposarcoma Subtype Round Cell 60 0 50 Dediff Pleomorphic 40 Nl Fat 30 Myxoid 20 Expression Level Well Diff 10 FGFR-2 by Liposarcoma Subtype 0 Round Cell 200 16 0 Dediff Nl Fat Pleomorphic 12 0 Myxoid 80 Expression Level Well Diff 40 Round Cell 0 Dediff Pleomorphic Nl Fat Myxoid Well Diff CITED1 by Liposarcoma Subtype Round Cell 10 0 80 Dediff Pleomorphic 60 40 Myxoid Expression Level 20 0 Round Cell Nl Fat Pleomorphic Well Diff Dediff Myxoid Round Cell Pleomorphic Table 1. 142 Gene classifier for liposarcoma subtype The genes used for each pair wise subtype comparison are grouped together. The flag column indicates which genes are unique to each subtype comparison. The values show the mean expression levels (actually the mean of the log expression levels was computed and than transformed back to absolute expression level). -

Supplemental Material
Supplemental Table B ARGs in alphabetical order Symbol Title 3 months 6 months 9 months 12 months 23 months ANOVA Direction Category 38597 septin 2 1557 ± 44 1555 ± 44 1579 ± 56 1655 ± 26 1691 ± 31 0.05219 up Intermediate 0610031j06rik kidney predominant protein NCU-G1 491 ± 6 504 ± 14 503 ± 11 527 ± 13 534 ± 12 0.04747 up Early Adult 1G5 vesicle-associated calmodulin-binding protein 662 ± 23 675 ± 17 629 ± 16 617 ± 20 583 ± 26 0.03129 down Intermediate A2m alpha-2-macroglobulin 262 ± 7 272 ± 8 244 ± 6 290 ± 7 353 ± 16 0.00000 up Midlife Aadat aminoadipate aminotransferase (synonym Kat2) 180 ± 5 201 ± 12 223 ± 7 244 ± 14 275 ± 7 0.00000 up Early Adult Abca2 ATP-binding cassette, sub-family A (ABC1), member 2 958 ± 28 1052 ± 58 1086 ± 36 1071 ± 44 1141 ± 41 0.05371 up Early Adult Abcb1a ATP-binding cassette, sub-family B (MDR/TAP), member 1A 136 ± 8 147 ± 6 147 ± 13 155 ± 9 185 ± 13 0.01272 up Midlife Acadl acetyl-Coenzyme A dehydrogenase, long-chain 423 ± 7 456 ± 11 478 ± 14 486 ± 13 512 ± 11 0.00003 up Early Adult Acadvl acyl-Coenzyme A dehydrogenase, very long chain 426 ± 14 414 ± 10 404 ± 13 411 ± 15 461 ± 10 0.01017 up Late Accn1 amiloride-sensitive cation channel 1, neuronal (degenerin) 242 ± 10 250 ± 9 237 ± 11 247 ± 14 212 ± 8 0.04972 down Late Actb actin, beta 12965 ± 310 13382 ± 170 13145 ± 273 13739 ± 303 14187 ± 269 0.01195 up Midlife Acvrinp1 activin receptor interacting protein 1 304 ± 18 285 ± 21 274 ± 13 297 ± 21 341 ± 14 0.03610 up Late Adk adenosine kinase 1828 ± 43 1920 ± 38 1922 ± 22 2048 ± 30 1949 ± 44 0.00797 up Early -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid
The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid Leukemia Cells Supported Only by Exogenous Cytokines Is Associated with a Patient Subset with Adverse Outcome Annette K. Brenner, Elise Aasebø, Maria Hernandez-Valladares, Frode Selheim, Frode Berven, Ida-Sofie Grønningsæter, Sushma Bartaula-Brevik and Øystein Bruserud Supplementary Material S2 of S31 Table S1. Detailed information about the 68 AML patients included in the study. # of blasts Viability Proliferation Cytokine Viable cells Change in ID Gender Age Etiology FAB Cytogenetics Mutations CD34 Colonies (109/L) (%) 48 h (cpm) secretion (106) 5 weeks phenotype 1 M 42 de novo 241 M2 normal Flt3 pos 31.0 3848 low 0.24 7 yes 2 M 82 MF 12.4 M2 t(9;22) wt pos 81.6 74,686 low 1.43 969 yes 3 F 49 CML/relapse 149 M2 complex n.d. pos 26.2 3472 low 0.08 n.d. no 4 M 33 de novo 62.0 M2 normal wt pos 67.5 6206 low 0.08 6.5 no 5 M 71 relapse 91.0 M4 normal NPM1 pos 63.5 21,331 low 0.17 n.d. yes 6 M 83 de novo 109 M1 n.d. wt pos 19.1 8764 low 1.65 693 no 7 F 77 MDS 26.4 M1 normal wt pos 89.4 53,799 high 3.43 2746 no 8 M 46 de novo 26.9 M1 normal NPM1 n.d. n.d. 3472 low 1.56 n.d. no 9 M 68 MF 50.8 M4 normal D835 pos 69.4 1640 low 0.08 n.d. -

Predicting Coupling Probabilities of G-Protein Coupled Receptors Gurdeep Singh1,2,†, Asuka Inoue3,*,†, J
Published online 30 May 2019 Nucleic Acids Research, 2019, Vol. 47, Web Server issue W395–W401 doi: 10.1093/nar/gkz392 PRECOG: PREdicting COupling probabilities of G-protein coupled receptors Gurdeep Singh1,2,†, Asuka Inoue3,*,†, J. Silvio Gutkind4, Robert B. Russell1,2,* and Francesco Raimondi1,2,* 1CellNetworks, Bioquant, Heidelberg University, Im Neuenheimer Feld 267, 69120 Heidelberg, Germany, 2Biochemie Zentrum Heidelberg (BZH), Heidelberg University, Im Neuenheimer Feld 328, 69120 Heidelberg, Germany, 3Graduate School of Pharmaceutical Sciences, Tohoku University, Sendai, Miyagi 980-8578, Japan and 4Department of Pharmacology and Moores Cancer Center, University of California, San Diego, La Jolla, CA 92093, USA Received February 10, 2019; Revised April 13, 2019; Editorial Decision April 24, 2019; Accepted May 01, 2019 ABSTRACT great use in tinkering with signalling pathways in living sys- tems (5). G-protein coupled receptors (GPCRs) control multi- Ligand binding to GPCRs induces conformational ple physiological states by transducing a multitude changes that lead to binding and activation of G-proteins of extracellular stimuli into the cell via coupling to situated on the inner cell membrane. Most of mammalian intra-cellular heterotrimeric G-proteins. Deciphering GPCRs couple with more than one G-protein giving each which G-proteins couple to each of the hundreds receptor a distinct coupling profile (6) and thus specific of GPCRs present in a typical eukaryotic organism downstream cellular responses. Determining these coupling is therefore critical to understand signalling. Here, profiles is critical to understand GPCR biology and phar- we present PRECOG (precog.russelllab.org): a web- macology. Despite decades of research and hundreds of ob- server for predicting GPCR coupling, which allows served interactions, coupling information is still missing for users to: (i) predict coupling probabilities for GPCRs many receptors and sequence determinants of coupling- specificity are still largely unknown. -

Quantification of Cardiovascular Disease Biomarkers in Human
proteomes Article Quantification of Cardiovascular Disease Biomarkers in Human Platelets by Targeted Mass Spectrometry Sebastian Malchow ID , Christina Loosse, Albert Sickmann and Christin Lorenz * Leibniz-Institut für Analytische Wissenschaften-ISAS-e.V., 44139 Dortmund, Germany; [email protected] (S.M.); [email protected] (C.L.); [email protected] (A.S.) * Correspondence: [email protected]; Tel.: +49-231-1392-289 Received: 3 October 2017; Accepted: 13 November 2017; Published: 15 November 2017 Abstract: Platelets are known to be key players in thrombosis and hemostasis, contributing to the genesis and progression of cardiovascular diseases. Due to their pivotal role in human physiology and pathology, platelet function is regulated tightly by numerous factors which have either stimulatory or inhibitory effects. A variety of factors, e.g., collagen, fibrinogen, ADP, vWF, thrombin, and thromboxane promote platelet adhesion and aggregation by utilizing multiple intracellular signal cascades. To quantify platelet proteins for this work, a targeted proteomics workflow was applied. In detail, platelets are isolated and lyzed, followed by a tryptic protein digest. Subsequently, a mix of stable isotope-labeled peptides of interesting biomarker proteins in concentrations ranging from 0.1 to 100 fmol is added to 3 µg digest. These peptides are used as an internal calibration curve to accurately quantify endogenous peptides and corresponding proteins in a pooled platelet reference sample by nanoLC-MS/MS with parallel reaction monitoring. In order to assure a valid quantification, limit of detection (LOD) and limit of quantification (LOQ), as well as linear range, were determined. This quantification of platelet activation and proteins by targeted mass spectrometry may enable novel diagnostic strategies in the detection and prevention of cardiovascular diseases. -

Supplementary Information Method CLEAR-CLIP. Mouse Keratinocytes
Supplementary Information Method CLEAR-CLIP. Mouse keratinocytes of the designated genotype were maintained in E-low calcium medium. Inducible cells were treated with 3 ug/ml final concentration doxycycline for 24 hours before performing CLEAR-CLIP. One 15cm dish of confluent cells was used per sample. Cells were washed once with cold PBS. 10mls of cold PBS was then added and cells were irradiated with 300mJ/cm2 UVC (254nM wavelength). Cells were then scraped from the plates in cold PBS and pelleted by centrifugation at 1,000g for 2 minutes. Pellets were frozen at -80oC until needed. Cells were then lysed on ice with occasional vortexing in 1ml of lysis buffer (50mM Tris-HCl pH 7.4, 100mM NaCl, 1mM MgCl2, 0.1 mM CaCl2, 1% NP-40, 0.5% Sodium Deoxycholate, 0.1% SDS) containing 1X protease inhibitors (Roche #88665) and RNaseOUT (Invitrogen #10777019) at 4ul/ml final concentration. Next, TurboDNase (Invitrogen #AM2238, 10U), RNase A (0.13ug) and RNase T1 (0.13U) were added and samples were incubated at 37oC for 5 minutes with occasional mixing. Samples were immediately placed on ice and then centrifuged at 16,160g at 4oC for 20 minutes to clear lysate. 25ul of Protein-G Dynabeads (Invitrogen #10004D) were used per IP. Dynabeads were pre-washed with lysis buffer and pre- incubated with 3ul of Wako Anti-Mouse-Ago2 (2D4) antibody. The dynabead/antibody mixture was added to the lysate and rocked for 2 hours at 4oC. All steps after the IP were done on bead until samples were loaded into the polyacrylamide gel. -

G-Protein ␥-Complex Is Crucial for Efficient Signal Amplification in Vision
The Journal of Neuroscience, June 1, 2011 • 31(22):8067–8077 • 8067 Cellular/Molecular G-Protein ␥-Complex Is Crucial for Efficient Signal Amplification in Vision Alexander V. Kolesnikov,1 Loryn Rikimaru,2 Anne K. Hennig,1 Peter D. Lukasiewicz,1 Steven J. Fliesler,4,5,6,7 Victor I. Govardovskii,8 Vladimir J. Kefalov,1 and Oleg G. Kisselev2,3 1Department of Ophthalmology and Visual Sciences, Washington University School of Medicine, St. Louis, Missouri 63110, Departments of 2Ophthalmology and 3Biochemistry and Molecular Biology, Saint Louis University School of Medicine, Saint Louis, Missouri 63104, 4Research Service, Veterans Administration Western New York Healthcare System, and Departments of 5Ophthalmology (Ross Eye Institute) and 6Biochemistry, University at Buffalo/The State University of New York (SUNY), and 7SUNY Eye Institute, Buffalo, New York 14215, and 8Sechenov Institute for Evolutionary Physiology and Biochemistry, Russian Academy of Sciences, Saint Petersburg 194223, Russia A fundamental question of cell signaling biology is how faint external signals produce robust physiological responses. One universal mechanism relies on signal amplification via intracellular cascades mediated by heterotrimeric G-proteins. This high amplification system allows retinal rod photoreceptors to detect single photons of light. Although much is now known about the role of the ␣-subunit of the rod-specific G-protein transducin in phototransduction, the physiological function of the auxiliary ␥-complex in this process remains a mystery. Here, we show that elimination of the transducin ␥-subunit drastically reduces signal amplification in intact mouse rods. The consequence is a striking decline in rod visual sensitivity and severe impairment of nocturnal vision. Our findings demonstrate that transducin ␥-complex controls signal amplification of the rod phototransduction cascade and is critical for the ability of rod photoreceptors to function in low light conditions. -

G Protein-Coupled Receptors
www.aladdin-e.com Address:800 S Wineville Avenue, Ontario, CA 91761,USA Website:www.aladdin-e.com Email USA: [email protected] Email EU: [email protected] Email Asia Pacific: [email protected] G PROTEIN-COUPLED RECEPTORS Overview: The completion of the Human Genome Project allowed the identification of a large family of proteins with a common motif of seven groups of 20–24 hydrophobic amino acids arranged as a-helices. Approximately 800 of these seven transmembrane (7TM) receptors have been identified of which over 300 are non-olfactory receptors (see Fredriksson et al., 2003; Lagerstrom and Schioth, 2008). Subdivision on the basis of sequence homology allows the definition of rhodopsin, secretin, adhesion, glutamate and Frizzled receptor families. NC-IUPHAR recognizes Classes A, B, and C, which equate to the rhodopsin, secretin, and glutamate receptor families. The nomenclature of 7TM receptors is commonly used interchangeably with G protein-coupled receptors (GPCR), although the former nomenclature recognises signalling of 7TM receptors through pathways not involving G proteins. For example, adiponectin and membrane progestin receptors have some sequence homology to 7TM receptors but signal independently of G proteins and appear to reside in membranes in an inverted fashion compared to conventional GPCR. Additionally, the NPR-C natriuretic peptide receptor (see Page S195) has a single transmembrane domain structure, but appears to couple to G proteins to generate cellular responses. The 300+ non-olfactory GPCR are the targets for the majority of drugs in clinical usage (Overington et al., 2006), although only a minority of these receptors are exploited therapeutically. -

Comparative Analysis of Human Chromosome 7Q21 and Mouse
Downloaded from genome.cshlp.org on October 2, 2021 - Published by Cold Spring Harbor Laboratory Press Letter Comparative analysis of human chromosome 7q21 and mouse proximal chromosome 6 reveals a placental-specific imprinted gene, TFPI2/Tfpi2, which requires EHMT2 and EED for allelic-silencing David Monk,1,6 Alexandre Wagschal,2 Philippe Arnaud,2 Pari-Sima Mu¨ller,3 Layla Parker-Katiraee,4 Déborah Bourc’his,5 Stephen W. Scherer,4 Robert Feil,2 Philip Stanier,1 and Gudrun E. Moore1 1Institute of Child Health, London WC1N 1EH, United Kingdom; 2Institute of Molecular Genetics, CNRS UMR-5535 and University of Montpellier-II, 34293 Montpellier, France; 3Sir William Dunn School of Pathology, University of Oxford, Oxford OX1 3RE, United Kingdom; 4Center for Applied Genomics, The Hospital for Sick Children, Toronto M5G 1L7, Canada; 5Inserm U741, F-75251 Paris Cedex 05, France Genomic imprinting is a developmentally important mechanism that involves both differential DNA methylation and allelic histone modifications. Through detailed comparative characterization, a large imprinted domain mapping to chromosome 7q21 in humans and proximal chromosome 6 in mice was redefined. This domain is organized around a maternally methylated CpG island comprising the promoters of the adjacent PEG10 and SGCE imprinted genes. Examination of Dnmt3l−/+ conceptuses shows that imprinted expression for all genes of the cluster depends upon the germline methylation at this putative “imprinting control region” (ICR). Similarly as for other ICRs, we find its DNA-methylated allele to be associated with trimethylation of lysine 9 on histone H3 (H3K9me3) and trimethylation of lysine 20 on histone H4 (H4K20me3), whereas the transcriptionally active paternal allele is enriched in H3K4me2 and H3K9 acetylation.