Identification of Pathogenic Genes of Pterygium Based on the Gene Expression Omnibus Database
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Materials: Evaluation of Cytotoxicity and Α-Glucosidase Inhibitory Activity of Amide and Polyamino-Derivatives of Lupane Triterpenoids
Supplementary Materials: Evaluation of cytotoxicity and α-glucosidase inhibitory activity of amide and polyamino-derivatives of lupane triterpenoids Oxana B. Kazakova1*, Gul'nara V. Giniyatullina1, Akhat G. Mustafin1, Denis A. Babkov2, Elena V. Sokolova2, Alexander A. Spasov2* 1Ufa Institute of Chemistry of the Ufa Federal Research Centre of the Russian Academy of Sciences, 71, pr. Oktyabrya, 450054 Ufa, Russian Federation 2Scientific Center for Innovative Drugs, Volgograd State Medical University, Novorossiyskaya st. 39, Volgograd 400087, Russian Federation Correspondence Prof. Dr. Oxana B. Kazakova Ufa Institute of Chemistry of the Ufa Federal Research Centre of the Russian Academy of Sciences 71 Prospeсt Oktyabrya Ufa, 450054 Russian Federation E-mail: [email protected] Prof. Dr. Alexander A. Spasov Scientific Center for Innovative Drugs of the Volgograd State Medical University 39 Novorossiyskaya st. Volgograd, 400087 Russian Federation E-mail: [email protected] Figure S1. 1H and 13C of compound 2. H NH N H O H O H 2 2 Figure S2. 1H and 13C of compound 4. NH2 O H O H CH3 O O H H3C O H 4 3 Figure S3. Anticancer screening data of compound 2 at single dose assay 4 Figure S4. Anticancer screening data of compound 7 at single dose assay 5 Figure S5. Anticancer screening data of compound 8 at single dose assay 6 Figure S6. Anticancer screening data of compound 9 at single dose assay 7 Figure S7. Anticancer screening data of compound 12 at single dose assay 8 Figure S8. Anticancer screening data of compound 13 at single dose assay 9 Figure S9. Anticancer screening data of compound 14 at single dose assay 10 Figure S10. -

Active Rac1 Detection Kit 2012 09/20
Active Rac1 Detection Kit 1 Kit Orders n 877-616-CELL (2355) (30 immunoprecipitations) [email protected] Support n 877-678-TECH (8324) [email protected] Web n www.cellsignal.com rev. 09/24/20 #8815 For Research Use Only. Not For Use In Diagnostic Procedures. Species Cross-Reactivity: H, M Components Ship As: 11894S Item # Kit Quantity Storage Temp Description: The Active Rac1 Detection Kit provides all GTPgS 11521 1 X 50 µl –80°C reagents necessary for measuring activation of Rac1 GTPase in the cell. GST-PAK1-PBD fusion protein is used to bind GDP 11522 1 X 50 µl –80°C the activated form of GTP-bound Rac1, which can then be Components Ship As: 11894S Item # Kit Quantity Storage Temp immunoprecipitated with glutathione resin. Rac1 activation levels are then determined by western blot using a Rac1 GST-Human PAK1-PBD 8659 1 X 600 µg –20°C Mouse mAb. Rac1 Mouse mAb 8631 1 X 50 µl –20°C Specificity/Sensitivity: Active Rac1 Detection Kit detects Components Ship As: 11860S Item # Kit Quantity Storage Temp endogenous levels of GTP-bound (active) Rac1 as shown in Lysis/Binding/Wash Buffer 11524 1 X 100 mL 4°C Figure 1. This kit detects proteins from the indicated species, as determined through in-house testing, but may also detect Glutathione Resin 11523 1 X 3 ml 4°C homologous proteins from other species. SDS Sample Buffer 11525 1 X 1.5 ml 4°C Background: The Ras superfamily of small GTP-binding Spin Cup and Collection Tubes 11526 1 X 30 vial RT proteins (G proteins) comprise a large class of proteins (over 150 members) that can be classified into at least five families based on their sequence and functional similarities: 1 2 3 4 Figure 1. -

Transcriptomic Analysis Reveals Abnormal Expression of Prion Disease Gene Pathway in Brains from Patients with Autism Spectrum Disorders
brain sciences Article Transcriptomic Analysis Reveals Abnormal Expression of Prion Disease Gene Pathway in Brains from Patients with Autism Spectrum Disorders Salvo Danilo Lombardo 1 , Giuseppe Battaglia 2,3, Maria Cristina Petralia 4, Katia Mangano 1 , Maria Sofia Basile 1, Valeria Bruno 2,3, Paolo Fagone 1,* , Rita Bella 5, Ferdinando Nicoletti 1 and Eugenio Cavalli 1 1 Department of Biomedical and Biotechnological Sciences, University of Catania, 95123 Catania, Italy; [email protected] (S.D.L.); [email protected] (K.M.); sofi[email protected] (M.S.B.); [email protected] (F.N.); [email protected] (E.C.) 2 Department of Physiology and Pharmacology, University Sapienza, Piazzale A. Moro, 5, 00185 Roma, Italy; [email protected] (G.B.); [email protected] (V.B.) 3 IRCCS Neuromed, Località Camerelle, 86077 Pozzilli, Italy 4 Department of Educational Sciences, University of Catania, 95124 Catania, Italy; [email protected] 5 Department of Medical Sciences, Surgery and Advanced Technologies, University of Catania, 95123 Catania, Italy; [email protected] * Correspondence: [email protected] Received: 26 February 2020; Accepted: 26 March 2020; Published: 29 March 2020 Abstract: The role of infections in the pathogenesis of autism spectrum disorder (ASD) is still controversial. In this study, we aimed to evaluate markers of infections and immune activation in ASD by performing a meta-analysis of publicly available whole-genome transcriptomic datasets of brain samples from autistic patients and otherwise normal people. Among the differentially expressed genes, no significant enrichment was observed for infectious diseases previously associated with ASD, including herpes simplex virus-1 (HSV-1), cytomegalovirus and Epstein–Barr virus in brain samples, nor was it found in peripheral blood from ASD patients. -

Comparative Functional Analysis of the Rac Gtpasesprovided by Elsevier - Publisher Connector
FEBS Letters 555 (2003) 556^560 FEBS 27913 View metadata, citation and similar papers at core.ac.uk brought to you by CORE Comparative functional analysis of the Rac GTPasesprovided by Elsevier - Publisher Connector Lars Christian Haeusler, Lars Blumenstein, Patricia Stege, Radovan Dvorsky, Mohammad Reza Ahmadianà Max-Planck-Institute of Molecular Physiology, Department of Structural Biology, Otto-Hahn-Strasse 11, 44227 Dortmund, Germany Received 18 August 2003; revised 17 September 2003; accepted 5 November 2003 First published online 25 November 2003 Edited by Irmgard Sinning 3), that are encoded by di¡erent genes, share between 89 and Abstract Small GTPases of the Rho family including Rac, Rho and Cdc42 regulate di¡erent cellular processes like reorga- 93% identity in their respective amino acid sequence [3,4]. nization of the actin cytoskeleton by acting as molecular Rac1, the best-investigated isoform, regulates gene expression, switches. The three distinct mammalian Rac proteins share cell cycle progression and rearrangement of the actin cytoskel- very high sequence identity but how their speci¢city is achieved eton [5]. Rac2 is proposed to be responsible for the regulation is hitherto unknown. Here we show that Rac1 and Rac3 are of the oxidative burst in hematopoietic cells but it has been very closely related concerning their biochemical properties, shown that Rac1 contributes to this process, too [6]. Rac1 and such as e¡ector interaction, nucleotide binding and hydrolysis. Rac3 are ubiquitously expressed and therefore regulate a wide In contrast, Rac2 displays a slower nucleotide association and is variety of cellular processes, whereas Rac2 is predominantly more e⁄ciently activated by the Rac-GEF Tiam1. -

Identification of Differentially Expressed Genes in Human Bladder Cancer Through Genome-Wide Gene Expression Profiling
521-531 24/7/06 18:28 Page 521 ONCOLOGY REPORTS 16: 521-531, 2006 521 Identification of differentially expressed genes in human bladder cancer through genome-wide gene expression profiling KAZUMORI KAWAKAMI1,3, HIDEKI ENOKIDA1, TOKUSHI TACHIWADA1, TAKENARI GOTANDA1, KENGO TSUNEYOSHI1, HIROYUKI KUBO1, KENRYU NISHIYAMA1, MASAKI TAKIGUCHI2, MASAYUKI NAKAGAWA1 and NAOHIKO SEKI3 1Department of Urology, Graduate School of Medical and Dental Sciences, Kagoshima University, 8-35-1 Sakuragaoka, Kagoshima 890-8520; Departments of 2Biochemistry and Genetics, and 3Functional Genomics, Graduate School of Medicine, Chiba University, 1-8-1 Inohana, Chuo-ku, Chiba 260-8670, Japan Received February 15, 2006; Accepted April 27, 2006 Abstract. Large-scale gene expression profiling is an effective CKS2 gene not only as a potential biomarker for diagnosing, strategy for understanding the progression of bladder cancer but also for staging human BC. This is the first report (BC). The aim of this study was to identify genes that are demonstrating that CKS2 expression is strongly correlated expressed differently in the course of BC progression and to with the progression of human BC. establish new biomarkers for BC. Specimens from 21 patients with pathologically confirmed superficial (n=10) or Introduction invasive (n=11) BC and 4 normal bladder samples were studied; samples from 14 of the 21 BC samples were subjected Bladder cancer (BC) is among the 5 most common to microarray analysis. The validity of the microarray results malignancies worldwide, and the 2nd most common tumor of was verified by real-time RT-PCR. Of the 136 up-regulated the genitourinary tract and the 2nd most common cause of genes we detected, 21 were present in all 14 BCs examined death in patients with cancer of the urinary tract (1-7). -

Pig Antibodies
Pig Antibodies gene_name sku Entry_Name Protein_Names Organism Length Identity CDX‐2 ARP31476_P050 D0V4H7_PIG Caudal type homeobox 2 (Fragment) Sus scrofa (Pig) 147 100.00% CDX‐2 ARP31476_P050 A7MAE3_PIG Caudal type homeobox transcription factor 2 (Fragment) Sus scrofa (Pig) 75 100.00% Tnnt3 ARP51286_P050 Q75NH3_PIG Troponin T fast skeletal muscle type Sus scrofa (Pig) 271 85.00% Tnnt3 ARP51286_P050 Q75NH2_PIG Troponin T fast skeletal muscle type Sus scrofa (Pig) 266 85.00% Tnnt3 ARP51286_P050 Q75NH1_PIG Troponin T fast skeletal muscle type Sus scrofa (Pig) 260 85.00% Tnnt3 ARP51286_P050 Q75NH0_PIG Troponin T fast skeletal muscle type Sus scrofa (Pig) 250 85.00% Tnnt3 ARP51286_P050 Q75NG8_PIG Troponin T fast skeletal muscle type Sus scrofa (Pig) 266 85.00% Tnnt3 ARP51286_P050 Q75NG7_PIG Troponin T fast skeletal muscle type Sus scrofa (Pig) 260 85.00% Tnnt3 ARP51286_P050 Q75NG6_PIG Troponin T fast skeletal muscle type Sus scrofa (Pig) 250 85.00% Tnnt3 ARP51286_P050 TNNT3_PIG Troponin T, fast skeletal muscle (TnTf) Sus scrofa (Pig) 271 85.00% ORF Names:PANDA_000462 EMBL EFB13877.1OrganismAiluropod High mobility group protein B2 (High mobility group protein a melanoleuca (Giant panda) ARP31939_P050 HMGB2_PIG 2) (HMG‐2) Sus scrofa (Pig) 210 100.00% Agpat5 ARP47429_P050 B8XTR3_PIG 1‐acylglycerol‐3‐phosphate O‐acyltransferase 5 Sus scrofa (Pig) 365 85.00% irf9 ARP31200_P050 Q29390_PIG Transcriptional regulator ISGF3 gamma subunit (Fragment) Sus scrofa (Pig) 57 100.00% irf9 ARP31200_P050 Q0GFA1_PIG Interferon regulatory factor 9 Sus scrofa (Pig) -

Supplemental Digital Content (Sdc) Sdc, Materials
SUPPLEMENTAL DIGITAL CONTENT (SDC) SDC, MATERIALS AND METHODS Animals This study used 9-12 week old male C57BL/6 mice (Jackson Laboratory, Bar Harbor, ME). This study conformed to the National Institutes of Health guidelines and was conducted under animal protocols approved by the University of Virginia’s Institutional Animal Care and Use Committee. Murine DCD Lung Procedure Mice were anesthetized by isoflurane inhalation and euthanized by cervical dislocation followed by a 60-minute period of “no-touch” warm ischemia. Mice then underwent extended median sternotomy and midline cervical exposure followed by intubation for the initiation of mechanical ventilation at 120 strokes/minute with room air. The left atrium was vented via an atriotomy followed by infusion of the lungs with 3 mL 4°C Perfadex® solution (Vitrolife Inc., Denver, CO) supplemented with THAM Solution (Vitrolife, Kungsbacka, Sweden), estimating weight-based volume recommendations for pulmonary artery perfusion (140mL/kg) (1). The chest was then packed with ice and the trachea occluded by silk-suture tie at tidal volume (7µL/g body weight) prior to cold static preservation (CSP) for 60 minutes at 4°C. Mice were then randomized into three experimental groups: 1) CSP alone with no EVLP, 2) EVLP with Steen solution and 3) EVLP with Steen solution supplemented with the highly selective A2AR agonist, ATL1223 (30nM, Lewis and Clark Pharmaceuticals, Charlottesville, VA). Mice treated with ATL1223 during EVLP also received ATL1223 treatment (30nM) during the Perfadex flush prior to CSP whereas the EVLP group received vehicle (DMSO) during the flush. CSP lungs, which did not undergo EVLP, underwent immediate functional assessment after re-intubation as described below. -
Drosophila and Human Transcriptomic Data Mining Provides Evidence for Therapeutic
Drosophila and human transcriptomic data mining provides evidence for therapeutic mechanism of pentylenetetrazole in Down syndrome Author Abhay Sharma Institute of Genomics and Integrative Biology Council of Scientific and Industrial Research Delhi University Campus, Mall Road Delhi 110007, India Tel: +91-11-27666156, Fax: +91-11-27662407 Email: [email protected] Nature Precedings : hdl:10101/npre.2010.4330.1 Posted 5 Apr 2010 Running head: Pentylenetetrazole mechanism in Down syndrome 1 Abstract Pentylenetetrazole (PTZ) has recently been found to ameliorate cognitive impairment in rodent models of Down syndrome (DS). The mechanism underlying PTZ’s therapeutic effect is however not clear. Microarray profiling has previously reported differential expression of genes in DS. No mammalian transcriptomic data on PTZ treatment however exists. Nevertheless, a Drosophila model inspired by rodent models of PTZ induced kindling plasticity has recently been described. Microarray profiling has shown PTZ’s downregulatory effect on gene expression in fly heads. In a comparative transcriptomics approach, I have analyzed the available microarray data in order to identify potential mechanism of PTZ action in DS. I find that transcriptomic correlates of chronic PTZ in Drosophila and DS counteract each other. A significant enrichment is observed between PTZ downregulated and DS upregulated genes, and a significant depletion between PTZ downregulated and DS dowwnregulated genes. Further, the common genes in PTZ Nature Precedings : hdl:10101/npre.2010.4330.1 Posted 5 Apr 2010 downregulated and DS upregulated sets show enrichment for MAP kinase pathway. My analysis suggests that downregulation of MAP kinase pathway may mediate therapeutic effect of PTZ in DS. Existing evidence implicating MAP kinase pathway in DS supports this observation. -

The Rac3 Gtpase in Neuronal Development, Neurodevelopmental Disorders, and Cancer
cells Review The Rac3 GTPase in Neuronal Development, Neurodevelopmental Disorders, and Cancer Ivan de Curtis Cell Adhesion Unit, Division of Neuroscience, San Raffaele Scientific Institute and Vita-Salute San Raffaele University, 20132 Milano, Italy; [email protected] Received: 27 August 2019; Accepted: 8 September 2019; Published: 11 September 2019 Abstract: Rho family small guanosine triphosphatases (GTPases) are important regulators of the cytoskeleton, and are critical in many aspects of cellular and developmental biology, as well as in pathological processes such as intellectual disability and cancer. Of the three members of the family, Rac3 has a more restricted expression in normal tissues compared to the ubiquitous member of the family, Rac1. The Rac3 polypeptide is highly similar to Rac1, and orthologues of the gene for Rac3 have been found only in vertebrates, indicating the late appearance of this gene during evolution. Increasing evidence over the past few years indicates that Rac3 plays an important role in neuronal development and in tumor progression, with specificities that distinguish the functions of Rac3 from the established functions of Rac1 in these processes. Here, results highlighting the importance of Rac3 in distinct aspects of neuronal development and tumor cell biology are presented, in support of the non-redundant role of different members of the two Rac GTPases in physiological and pathological processes. Keywords: Rho family GTPases; Rac3; neuronal development; mutations; intellectual disability; cancer; invasion; metastasis 1. Introduction In vertebrates the Rac family of guanosine triphosphatases (GTPases) includes three different genes that encode three highly similar proteins: the ubiquitously expressed Rac1 with its alternative spliced variant Rac1b [1,2]; Rac2 that is expressed only in hematopoietic cells [3]; and the Rac3 protein that is co-expressed with Rac1 in developing neurons and in other cell types, with a pattern of expression more restricted compared to Rac1 [4,5]. -
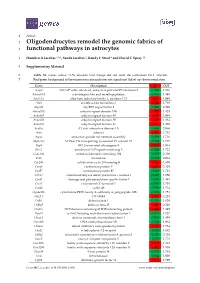
Oligodendrocytes Remodel the Genomic Fabrics of Functional Pathways in Astrocytes
1 Article 2 Oligodendrocytes remodel the genomic fabrics of 3 functional pathways in astrocytes 4 Dumitru A Iacobas 1,2,*, Sanda Iacobas 3, Randy F Stout 4 and David C Spray 2,5 5 Supplementary Material 6 Table S1. Genes whose >1.5x absolute fold-change did not meet the individual CUT criterion. 7 Red/green background of the expression ratio indicates not significant (false) up-/down-regulation. Gene Description X CUT Acap2 ArfGAP with coiled-coil, ankyrin repeat and PH domains 2 -1.540 1.816 Adamts18 a disintegrin-like and metallopeptidase -1.514 1.594 Akr1c12 aldo-keto reductase family 1, member C12 1.866 1.994 Alx3 aristaless-like homeobox 3 1.536 1.769 Alyref2 Aly/REF export factor 2 -1.880 2.208 Ankrd33b ankyrin repeat domain 33B 1.593 1.829 Ankrd45 ankyrin repeat domain 45 1.514 1.984 Ankrd50 ankyrin repeat domain 50 1.628 1.832 Ankrd61 ankyrin repeat domain 61 1.645 1.802 Arid1a AT rich interactive domain 1A -1.668 2.066 Artn artemin 1.524 1.732 Aspm abnormal spindle microtubule assembly -1.693 1.716 Atp6v1e1 ATPase, H+ transporting, lysosomal V1 subunit E1 -1.679 1.777 Bag4 BCL2-associated athanogene 4 1.723 1.914 Birc3 baculoviral IAP repeat-containing 3 -1.588 1.722 Ccdc104 coiled-coil domain containing 104 -1.819 2.130 Ccl2 chemokine -1.699 2.034 Cdc20b cell division cycle 20 homolog B 1.512 1.605 Cenpf centromere protein F 2.041 2.128 Cep97 centrosomal protein 97 -1.641 1.723 COX1 mitochondrially encoded cytochrome c oxidase I -1.607 1.650 Cpsf7 cleavage and polyadenylation specific factor 7 -1.635 1.891 Crct1 cysteine-rich -

The Neuroprotective Role of the GM1 Oligosaccharide, Ii3neu5ac-Gg4, In
Molecular Neurobiology (2019) 56:6673–6702 https://doi.org/10.1007/s12035-019-1556-8 The Neuroprotective Role of the GM1 Oligosaccharide, 3 II Neu5Ac-Gg4, in Neuroblastoma Cells Elena Chiricozzi1 & Margherita Maggioni1 & Erika di Biase1 & Giulia Lunghi1 & Maria Fazzari1 & Nicoletta Loberto 1 & Maffioli Elisa2 & Francesca Grassi Scalvini2 & Gabriella Tedeschi 2,3 & Sandro Sonnino1 Received: 10 January 2019 /Accepted: 13 March 2019 /Published online: 26 March 2019 # Springer Science+Business Media, LLC, part of Springer Nature 2019 Abstract 3 Recently, we demonstrated that the GM1 oligosaccharide, II Neu5Ac-Gg4 (OligoGM1), administered to cultured murine Neuro2a neuroblastoma cells interacts with the NGF receptor TrkA, leading to the activation of the ERK1/2 downstream pathway and to cell differentiation. To understand how the activation of the TrkA pathway is able to trigger key biochemical signaling, we performed a proteomic analysis on Neuro2a cells treated with 50 μM OligoGM1 for 24 h. Over 3000 proteins were identified. Among these, 324 proteins were exclusively expressed in OligoGM1-treated cells. Interestingly, several proteins expressed only in OligoGM1-treated cells are involved in biochemical mechanisms with a neuroprotective potential, reflecting the GM1 neuroprotective effect. In addition, we found that the exogenous administration of OligoGM1 reduced the cellular oxidative stress in Neuro2a cells and conferred protection against MPTP neurotoxicity. These results confirm and reinforce the idea that the molecular mechanisms underlying the GM1 neurotrophic and neuroprotective effects depend on its oligosaccharide chain, suggesting the activation of a positive signaling starting at plasma membrane level. Keywords GM1 ganglioside . GM1 oligosaccharide chain . TrkA neurotrophin receptor . Plasma membrane signaling . Neuroprotection . -

Translational Profiling Reveals the Transcriptome of Leptin Receptor Neurons and Its Regulation by Leptin
TRANSLATIONAL PROFILING REVEALS THE TRANSCRIPTOME OF LEPTIN RECEPTOR NEURONS AND ITS REGULATION BY LEPTIN by Margaret B. Allison A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Molecular and Integrative Physiology) In the University of Michigan 2015 Doctoral Committee: Professor Martin G. Myers Jr., Chair Associate Professor Carol F. Elias Professor Malcolm J. Low Professor Suzanne Moenter Professor Audrey Seasholtz Before you leave these portals To meet less fortunate mortals There's just one final message I would give to you: You all have learned reliance On the sacred teachings of science So I hope, through life, you never will decline In spite of philistine defiance To do what all good scientists do: Experiment! -- Cole Porter There is no cure for curiosity. -- unknown © Margaret Brewster Allison 2015 ACKNOWLEDGEMENTS If it takes a village to raise a child, it takes a research university to raise a graduate student. There are many people who have supported me over the past six years at Michigan, and it is hard to imagine pursuing my PhD without them. First and foremost among all the people I need to thank is my mentor, Martin. Nothing I might say here would ever suffice to cover the depth and breadth of my gratitude to him. Without his patience, his insight, and his at times insufferably positive outlook, I don’t know where I would be today. Martin supported my intellectual curiosity, honed my scientific inquiry, and allowed me to do some really fun research in his lab. It was a privilege and a pleasure to work for him and with him.