A Multilevel Assessment of Sediment Bioremediation with Bioturbating Macrofauna
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Metaproteogenomic Insights Beyond Bacterial Response to Naphthalene
ORIGINAL ARTICLE ISME Journal – Original article Metaproteogenomic insights beyond bacterial response to 5 naphthalene exposure and bio-stimulation María-Eugenia Guazzaroni, Florian-Alexander Herbst, Iván Lores, Javier Tamames, Ana Isabel Peláez, Nieves López-Cortés, María Alcaide, Mercedes V. del Pozo, José María Vieites, Martin von Bergen, José Luis R. Gallego, Rafael Bargiela, Arantxa López-López, Dietmar H. Pieper, Ramón Rosselló-Móra, Jesús Sánchez, Jana Seifert and Manuel Ferrer 10 Supporting Online Material includes Text (Supporting Materials and Methods) Tables S1 to S9 Figures S1 to S7 1 SUPPORTING TEXT Supporting Materials and Methods Soil characterisation Soil pH was measured in a suspension of soil and water (1:2.5) with a glass electrode, and 5 electrical conductivity was measured in the same extract (diluted 1:5). Primary soil characteristics were determined using standard techniques, such as dichromate oxidation (organic matter content), the Kjeldahl method (nitrogen content), the Olsen method (phosphorus content) and a Bernard calcimeter (carbonate content). The Bouyoucos Densimetry method was used to establish textural data. Exchangeable cations (Ca, Mg, K and 10 Na) extracted with 1 M NH 4Cl and exchangeable aluminium extracted with 1 M KCl were determined using atomic absorption/emission spectrophotometry with an AA200 PerkinElmer analyser. The effective cation exchange capacity (ECEC) was calculated as the sum of the values of the last two measurements (sum of the exchangeable cations and the exchangeable Al). Analyses were performed immediately after sampling. 15 Hydrocarbon analysis Extraction (5 g of sample N and Nbs) was performed with dichloromethane:acetone (1:1) using a Soxtherm extraction apparatus (Gerhardt GmbH & Co. -

Microbiome Exploration of Deep-Sea Carnivorous Cladorhizidae Sponges
Microbiome exploration of deep-sea carnivorous Cladorhizidae sponges by Joost Theo Petra Verhoeven A Thesis submitted to the School of Graduate Studies in partial fulfillment of the requirements for the degree of Doctor of Philosophy Department of Biology Memorial University of Newfoundland March 2019 St. John’s, Newfoundland and Labrador ABSTRACT Members of the sponge family Cladorhizidae are unique in having replaced the typical filter-feeding strategy of sponges by a predatory lifestyle, capturing and digesting small prey. These carnivorous sponges are found in many different environments, but are particularly abundant in deep waters, where they constitute a substantial component of the benthos. Sponges are known to host a wide range of microbial associates (microbiome) important for host health, but the extent of the microbiome in carnivorous sponges has never been extensively investigated and their importance is poorly understood. In this thesis, the microbiome of two deep-sea carnivorous sponge species (Chondrocladia grandis and Cladorhiza oxeata) is investigated for the first time, leveraging recent advances in high-throughput sequencing and through custom developed bioinformatic and molecular methods. Microbiome analyses showed that the carnivorous sponges co-occur with microorganisms and large differences in the composition and type of associations were observed between sponge species. Tissues of C. grandis hosted diverse bacterial communities, similar in composition between individuals, in stark contrast to C. oxeata where low microbial diversity was found with a high host-to-host variability. In C. grandis the microbiome was not homogeneous throughout the host tissue, and significant shifts occured within community members across anatomical regions, with the enrichment of specific bacterial taxa in particular anatomical niches, indicating a potential symbiotic role of such taxa within processes like prey digestion and chemolithoautotrophy. -
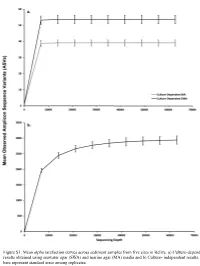
Figure S1. Mean Alpha Rarefaction Curves Across Sediment Samples from Five Sites in Belize
Figure S1. Mean alpha rarefaction curves across sediment samples from five sites in Belize. a) Culture-dependent results obtained using seawater agar (SWA) and marine agar (MA) media and b) Culture- independent results. Error bars represent standard error among replicates. a. 200 150 100 Faith’s PD Faith’s 50 0 Culturedependent MA Culturedependent SWA Cultureindependent Method b. 0.8 0.6 Pielou’s Evenness 0.4 Culturedependent MA Culturedependent SWA Cultureindependent Method Figure S2. Alpha diversity boxplots of marine sediment microbial communities from Carrie Bow Cay, Belize in culture-dependent and culture-independent samples determined using a) Faith’s Phylogenetic Diversity Index and b) Pielou’s Evenness. Culture-dependent methods include the use of marine agar medium (MA) and seawater agar medium (SWA. Data points are overlayed on the boxplot to show variation. a. 100% Proteobacteria Nitrospinae Bacteroidetes Fibrobacteres Planctomycetes Entotheonellaeota 90% Cyanobacteria Marinimicrobia (SAR406 clade) Firmicutes Deinococcus-Thermus Chloroflexi Margulisbacteria [A] Thaumarchaeota Unassigned 80% Acidobacteria Archaea UA Verrucomicrobia Acetothermia Actinobacteria Elusimicrobia 70% [A] Nanoarchaeaeota Tenericutes Kiritimatiellaeota TA06 Latescibacteria WS2 60% Spirochaetes Dependentiae Patescibacteria LCP-89 Gemmatimonadetes FCPU426 Omnitrophicaeota WPS-2 50% Fusobacteria CK-2C2-2 Bacteria UA Armatimonadetes [A] Euryarchaeota [A] Altiarchaeota Relative Percent 40% Calditrichaeota Poribacteria Lentisphaerae Cloacimonetes [A] Crenarchaeota -

Acetobacteroides Hydrogenigenes Gen. Nov., Sp. Nov., an Anaerobic Hydrogen-Producing Bacterium in the Family Rikenellaceae Isolated from a Reed Swamp
%paper no. ije063917 charlesworth ref: ije063917& New Taxa - Bacteroidetes International Journal of Systematic and Evolutionary Microbiology (2014), 64, 000–000 DOI 10.1099/ijs.0.063917-0 Acetobacteroides hydrogenigenes gen. nov., sp. nov., an anaerobic hydrogen-producing bacterium in the family Rikenellaceae isolated from a reed swamp Xiao-Li Su,1,2 Qi Tian,1,3 Jie Zhang,1,2 Xian-Zheng Yuan,1 Xiao-Shuang Shi,1 Rong-Bo Guo1 and Yan-Ling Qiu1 Correspondence 1Key Laboratory of Biofuels, Qingdao Institute of Bioenergy and Bioprocess Technology, Yan-Ling Qiu Chinese Academy of Sciences, Qingdao, Shandong Province 266101, PR China [email protected] 2University of Chinese Academy of Sciences, Beijing 100049, PR China 3Ocean University of China, Qingdao, 266101, PR China A strictly anaerobic, mesophilic, carbohydrate-fermenting, hydrogen-producing bacterium, designated strain RL-CT, was isolated from a reed swamp in China. Cells were Gram-stain- negative, catalase-negative, non-spore-forming, non-motile rods measuring 0.7–1.0 mm in width and 3.0–8.0 mm in length. The optimum temperature for growth of strain RL-CT was 37 6C (range 25–40 6C) and pH 7.0–7.5 (range pH 5.7–8.0). The strain could grow fermentatively on yeast extract, tryptone, arabinose, glucose, galactose, mannose, maltose, lactose, glycogen, pectin and starch. The main end products of glucose fermentation were acetate, H2 and CO2. Organic acids, alcohols and amino acids were not utilized for growth. Yeast extract was not required for growth; however, it stimulated growth slightly. Nitrate, sulfate, sulfite, thiosulfate, elemental sulfur and Fe(III) nitrilotriacetate were not reduced as terminal electron acceptors. -

Carboxylicivirga Flava Sp. Nov., Isolated from Marine Surface Sediment Hui Wang,1 Cancan Qi,1 Weiwei Chen,1 Wenwen Dong,1 Haitian Tang2 and Xiaoke Hu1
International Journal of Systematic and Evolutionary Microbiology (2016), 66, 5412–5416 DOI 10.1099/ijsem.0.001533 Carboxylicivirga flava sp. nov., isolated from marine surface sediment Hui Wang,1 Cancan Qi,1 Weiwei Chen,1 Wenwen Dong,1 Haitian Tang2 and Xiaoke Hu1 Correspondence 1Key Laboratory of Coastal Biology and Bioresource Utilization, Yantai Institute of Costal Zone Xiaoke Hu Research, Chinese Academy of Sciences, Yantai 264003, PR China [email protected] 2Yantai Marine Environment Monitoring Central Station, State Oceanic Administration, Yantai 264006, PR China A novel bacterial strain, designated Q15T, was isolated from sediments obtained from the Bohai Sea in China and subjected to a polyphasic taxonomic study. Cells of strain Q15T were Gram- stain-negative, strictly aerobic rods that produced circular, flat, orange colonies. Phylogenetic analysis based on 16S rRNA gene sequences revealed that Q15T was affiliated with the genus Carboxylicivirga in the family Marinilabiliaceae of the phylum Bacteroidetes. Strain Q15T differed genotypically from the type strains of the three recognized species of this genus (Carboxylicivirga taeanensis MEBiC 08903T, Carboxylicivirga mesophila MEBiC 07026T and Carboxylicivirga linearis FB218T) and shared 94.0–95.2 % 16S rRNA gene sequence similarity with them. The DNA G+C content of strain Q15T was 44.7 mol%. The predominant cellular fatty acids were iso-C15 : 0, anteiso-C15 : 0 and iso-C17 : 0 3-OH, and menaquinone MK-7 was the main respiratory quinone. Polar lipids contained phosphatidylethanolamine, an unidentified aminolipid, an unidentified phospholipid and other unknown lipids. Based on the data from the current polyphasic analysis, a novel species, Carboxylicivirga flava sp. -

Supplementary Information
Supplementary Information Comparative Microbiome and Metabolome Analyses of the Marine Tunicate Ciona intestinalis from Native and Invaded Habitats Caroline Utermann 1, Martina Blümel 1, Kathrin Busch 2, Larissa Buedenbender 1, Yaping Lin 3,4, Bradley A. Haltli 5, Russell G. Kerr 5, Elizabeta Briski 3, Ute Hentschel 2,6, Deniz Tasdemir 1,6* 1 GEOMAR Centre for Marine Biotechnology (GEOMAR-Biotech), Research Unit Marine Natural Products Chemistry, GEOMAR Helmholtz Centre for Ocean Research Kiel, Am Kiel-Kanal 44, 24106 Kiel, Germany 2 Research Unit Marine Symbioses, GEOMAR Helmholtz Centre for Ocean Research Kiel, Duesternbrooker Weg 20, 24105 Kiel, Germany 3 Research Group Invasion Ecology, Research Unit Experimental Ecology, GEOMAR Helmholtz Centre for Ocean Research Kiel, Duesternbrooker Weg 20, 24105 Kiel, Germany 4 Chinese Academy of Sciences, Research Center for Eco-Environmental Sciences, 18 Shuangqing Rd., Haidian District, Beijing, 100085, China 5 Department of Chemistry, University of Prince Edward Island, 550 University Avenue, Charlottetown, PE C1A 4P3, Canada 6 Faculty of Mathematics and Natural Sciences, Kiel University, Christian-Albrechts-Platz 4, Kiel 24118, Germany * Corresponding author: Deniz Tasdemir ([email protected]) This document includes: Supplementary Figures S1-S11 Figure S1. Genotyping of C. intestinalis with the mitochondrial marker gene COX3-ND1. Figure S2. Influence of the quality filtering steps on the total number of observed read pairs from amplicon sequencing. Figure S3. Rarefaction curves of OTU abundances for C. intestinalis and seawater samples. Figure S4. Multivariate ordination plots of the bacterial community associated with C. intestinalis. Figure S5. Across sample type and geographic origin comparison of the C. intestinalis associated microbiome. -

Flavobacterial Response to Organic Pollution
Vol. 51: 31–43, 2008 AQUATIC MICROBIAL ECOLOGY Published April 24 doi: 10.3354/ame01174 Aquat Microb Ecol Flavobacterial response to organic pollution Andrew Bissett1, 2, 3,*, John P. Bowman2, Chris M. Burke1 1School of Aquaculture, Tasmanian Aquaculture and Fisheries Institute, University of Tasmania and Aquafin CRC, Launceston, Tasmania 7250, Australia 2School of Agricultural Science, University of Tasmania, Hobart, Tasmania 7001, Australia 3Present address: Max Planck Institute for Marine Microbiology, Celsiusstrasse 1, 28359 Bremen, Germany ABSTRACT: Bacteria of the Cytophaga-Flavobacterium-Bacteroides (CFB) group (phylum Bac- teroidetes), in particular members of the class Flavobacteria, are among the most prominent hetero- trophic organisms in marine pelagic systems. They have also previously been found to be important in the initial biopolymer degradation of sedimentary organic matter. The Flavobacteria community was analysed in inshore, marine sediments subject to regular inputs of highly labile organic carbon in order to understand the importance of this group in carbon degradation. We used denaturing gra- dient gel electrophoresis and real-time PCR in a statistically robust manner, over 2 consecutive years, to demonstrate that the number of Flavobacteria in the sediment increased and community composi- tion shifted with organic loading. Further community shifts occurred after cessation of organic load- ing, and population numbers also decreased. Flavobacteria appear to be important in the initial responses of the sediment microbial community to organic loading, regardless of sediment type, but flavobacterial composition was not predictable. The highly dynamic nature and large diversity (func- tional redundancy) of the Flavobacteria in these sediments may contribute to this unpredictable response. KEY WORDS: Flavobacteria . -

Compile.Xlsx
Silva OTU GS1A % PS1B % Taxonomy_Silva_132 otu0001 0 0 2 0.05 Bacteria;Acidobacteria;Acidobacteria_un;Acidobacteria_un;Acidobacteria_un;Acidobacteria_un; otu0002 0 0 1 0.02 Bacteria;Acidobacteria;Acidobacteriia;Solibacterales;Solibacteraceae_(Subgroup_3);PAUC26f; otu0003 49 0.82 5 0.12 Bacteria;Acidobacteria;Aminicenantia;Aminicenantales;Aminicenantales_fa;Aminicenantales_ge; otu0004 1 0.02 7 0.17 Bacteria;Acidobacteria;AT-s3-28;AT-s3-28_or;AT-s3-28_fa;AT-s3-28_ge; otu0005 1 0.02 0 0 Bacteria;Acidobacteria;Blastocatellia_(Subgroup_4);Blastocatellales;Blastocatellaceae;Blastocatella; otu0006 0 0 2 0.05 Bacteria;Acidobacteria;Holophagae;Subgroup_7;Subgroup_7_fa;Subgroup_7_ge; otu0007 1 0.02 0 0 Bacteria;Acidobacteria;ODP1230B23.02;ODP1230B23.02_or;ODP1230B23.02_fa;ODP1230B23.02_ge; otu0008 1 0.02 15 0.36 Bacteria;Acidobacteria;Subgroup_17;Subgroup_17_or;Subgroup_17_fa;Subgroup_17_ge; otu0009 9 0.15 41 0.99 Bacteria;Acidobacteria;Subgroup_21;Subgroup_21_or;Subgroup_21_fa;Subgroup_21_ge; otu0010 5 0.08 50 1.21 Bacteria;Acidobacteria;Subgroup_22;Subgroup_22_or;Subgroup_22_fa;Subgroup_22_ge; otu0011 2 0.03 11 0.27 Bacteria;Acidobacteria;Subgroup_26;Subgroup_26_or;Subgroup_26_fa;Subgroup_26_ge; otu0012 0 0 1 0.02 Bacteria;Acidobacteria;Subgroup_5;Subgroup_5_or;Subgroup_5_fa;Subgroup_5_ge; otu0013 1 0.02 13 0.32 Bacteria;Acidobacteria;Subgroup_6;Subgroup_6_or;Subgroup_6_fa;Subgroup_6_ge; otu0014 0 0 1 0.02 Bacteria;Acidobacteria;Subgroup_6;Subgroup_6_un;Subgroup_6_un;Subgroup_6_un; otu0015 8 0.13 30 0.73 Bacteria;Acidobacteria;Subgroup_9;Subgroup_9_or;Subgroup_9_fa;Subgroup_9_ge; -

Insights Into Xylan Degradation and Haloalkaline
G C A T T A C G G C A T genes Article Insights into Xylan Degradation and Haloalkaline Adaptation through Whole-Genome Analysis of Alkalitalea saponilacus, an Anaerobic Haloalkaliphilic Bacterium Capable of Secreting Novel Halostable Xylanase Ziya Liao 1, Mark Holtzapple 2, Yanchun Yan 1, Haisheng Wang 1, Jun Li 3 and Baisuo Zhao 1,3,* 1 Graduate School, Chinese Academy of Agricultural Sciences, Beijing 100081, China; [email protected] (Z.L.); [email protected] (Y.Y.); [email protected] (H.W.) 2 Department of Chemical Engineering, Texas A&M University, College Station, TX 77843, USA; [email protected] 3 Institute of Agricultural Resources and Regional Planning, Chinese Academy of Agricultural Sciences, Beijing 100081, China; [email protected] * Correspondence: [email protected] Received: 30 October 2018; Accepted: 13 December 2018; Published: 20 December 2018 Abstract: The obligately anaerobic haloalkaliphilic bacterium Alkalitalea saponilacus can use xylan as the sole carbon source and produce propionate as the main fermentation product. Using mixed carbon sources of 0.4% (w/v) sucrose and 0.1% (w/v) birch xylan, xylanase production from A. saponilacus was 3.2-fold greater than that of individual carbon sources of 0.5% (w/v) sucrose or 0.5% (w/v) birch xylan. The xylanse is halostable and exhibits optimal activity over a broad salt concentration (2–6% NaCl). Its activity increased approximately 1.16-fold by adding 0.2% (v/v) Tween 20. To understand the potential genetic mechanisms of xylan degradation and molecular adaptation to saline-alkali extremes, the complete genome sequence of A. -

Abstract Tracing Hydrocarbon
ABSTRACT TRACING HYDROCARBON CONTAMINATION THROUGH HYPERALKALINE ENVIRONMENTS IN THE CALUMET REGION OF SOUTHEASTERN CHICAGO Kathryn Quesnell, MS Department of Geology and Environmental Geosciences Northern Illinois University, 2016 Melissa Lenczewski, Director The Calumet region of Southeastern Chicago was once known for industrialization, which left pollution as its legacy. Disposal of slag and other industrial wastes occurred in nearby wetlands in attempt to create areas suitable for future development. The waste creates an unpredictable, heterogeneous geology and a unique hyperalkaline environment. Upgradient to the field site is a former coking facility, where coke, creosote, and coal weather openly on the ground. Hydrocarbons weather into characteristic polycyclic aromatic hydrocarbons (PAHs), which can be used to create a fingerprint and correlate them to their original parent compound. This investigation identified PAHs present in the nearby surface and groundwaters through use of gas chromatography/mass spectrometry (GC/MS), as well as investigated the relationship between the alkaline environment and the organic contamination. PAH ratio analysis suggests that the organic contamination is not mobile in the groundwater, and instead originated from the air. 16S rDNA profiling suggests that some microbial communities are influenced more by pH, and some are influenced more by the hydrocarbon pollution. BIOLOG Ecoplates revealed that most communities have the ability to metabolize ring structures similar to the shape of PAHs. Analysis with bioinformatics using PICRUSt demonstrates that each community has microbes thought to be capable of hydrocarbon utilization. The field site, as well as nearby areas, are targets for habitat remediation and recreational development. In order for these remediation efforts to be successful, it is vital to understand the geochemistry, weathering, microbiology, and distribution of known contaminants. -

16S Rrna Gene Metabarcoding Indicates Species-Characteristic
16S rRNA Gene Metabarcoding Indicates Species-Characteristic Microbiomes in Deep-Sea Benthic Foraminifera Iines Salonen, Panagiota-Myrsini Chronopoulou, Hidetaka Nomaki, Dewi Langlet, Masashi Tsuchiya, Karoliina Koho To cite this version: Iines Salonen, Panagiota-Myrsini Chronopoulou, Hidetaka Nomaki, Dewi Langlet, Masashi Tsuchiya, et al.. 16S rRNA Gene Metabarcoding Indicates Species-Characteristic Microbiomes in Deep- Sea Benthic Foraminifera. Frontiers in Microbiology, Frontiers Media, 2021, 12, pp.694406. 10.3389/fmicb.2021.694406. hal-03306215 HAL Id: hal-03306215 https://hal.archives-ouvertes.fr/hal-03306215 Submitted on 29 Jul 2021 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. ORIGINAL RESEARCH published: 27 July 2021 doi: 10.3389/fmicb.2021.694406 16S rRNA Gene Metabarcoding Indicates Species-Characteristic Microbiomes in Deep-Sea Benthic Foraminifera Iines S. Salonen 1,2*, Panagiota-Myrsini Chronopoulou 1, Hidetaka Nomaki 2, Dewi Langlet 2,3,4, Masashi Tsuchiya 5 and Karoliina A. Koho 1 1 Ecosystems and Environment Research Program, University -

Extracellular Electron Shuttle Mediated
EXTRACELLULAR ELECTRON SHUTTLE MEDIATED BIODEGRADATION OF HEXAHYDRO-1,3,5-TRINITRO-1,3,5-TRIAZINE BY MAN JAE KWON B.S., Korea University, 1999 M.S., Korea University, 2001 DISSERTATION Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Environmental Engineering in Civil Engineering in the Graduate College of the University of Illinois at Urbana-Champaign, 2009 Urbana, Illinois Doctoral Committee: Assistant Professor Kevin T. Finneran, Chair Professor Charles J. Werth Professor Joseph J. Stucki Assistant Professor Timothy J. Strathmann ABSTRACT EXTRACELLULAR ELECTRON SHUTTLE MEDIATED BIODEGRADATION OF HEXAHYDRO-1,3,5-TRINITRO-1,3,5-TRIAZINE Man Jae Kwon, Ph.D. Department of Civil and Environmental Engineering University of Illinois at Urbana-Champaign, 2009 Kevin T. Finneran, Advisor Bioremediation is one suggested technology for hexahydro-1,3,5-trinitro- 1,3,5-triazine (RDX), as well as alternate explosive contaminants. The potential use of extracellular electron shuttling compounds to mediate electron transfer between iron (hydr)oxides and Fe(III)-reducing microorganisms was investigated for decontaminating RDX. The ultimate goal of this study is to develop an optimal strategy for the biodegradation of nitramine explosive compounds. This study demonstrated that Geobacter metallireducens , a model Fe(III)- and electron shuttle-reducing microorganism, utilize electron shuttles to stimulate cyclic nitramine reduction at rates significantly faster than those previously reported. This work was the first to demonstrate electron shuttle-mediated nitramine biodegradation with environmentally relevant pure cultures. Electron shuttle-mediated explosives biodegradation was also investigated among four additional species of Fe(III)- and quinone-reducing bacteria. All species showed the fastest RDX degradation when electron shuttling compounds were present although biotransformation kinetics were different among four Bacterial species.